Team:BU Wellesley Software/Notebook/CaseyNotebook
From 2011.igem.org
This summer the Wellesley HCI Lab worked on re-implementing an old system, G-nome Surfer 2.0, new and improved and optimized for prokaryotic genomes. To make an application practical for researching prokaryotic genomes we had to create a number of new interaction styles for dealing with navigation of a circular chromosome. Working on the system I worked on the new chromosome wheel, the publications system, the new primer designer and the associated BLAST.
Implementation
G-Nome Surfer Pro is written in C# using the Microsoft SDK for use on the Microsoft Surface. After much trial and error we re-implemented many parts of the system using data binding which allows a complete separation of state and logic. (more information about data binding can be found on MSDN [http://msdn.microsoft.com/en-us/library/ms752347.aspx here]) By using data binding we have managed to create remarkably stable code and haven't seen any crashes at all in usability testing and user studies. However, the structure and time required to set up the system is difficult to manage and some components outside the core (like primer designer) are implemented without. Fortunately, the underlying structure allows for editing of individual components and collaboration through the svn repository.
Navigation
The central chromosome wheel went through several iterations before arriving at the final simple rotation model we currently use. An early implementation allowed for zooming through use of a pair of arms (like in the early paper prototype.) Like many of the early user controls the arms involved a fair amount of hacking through the presentation logic and item templates through the .xaml files.
Publications
Primer Design
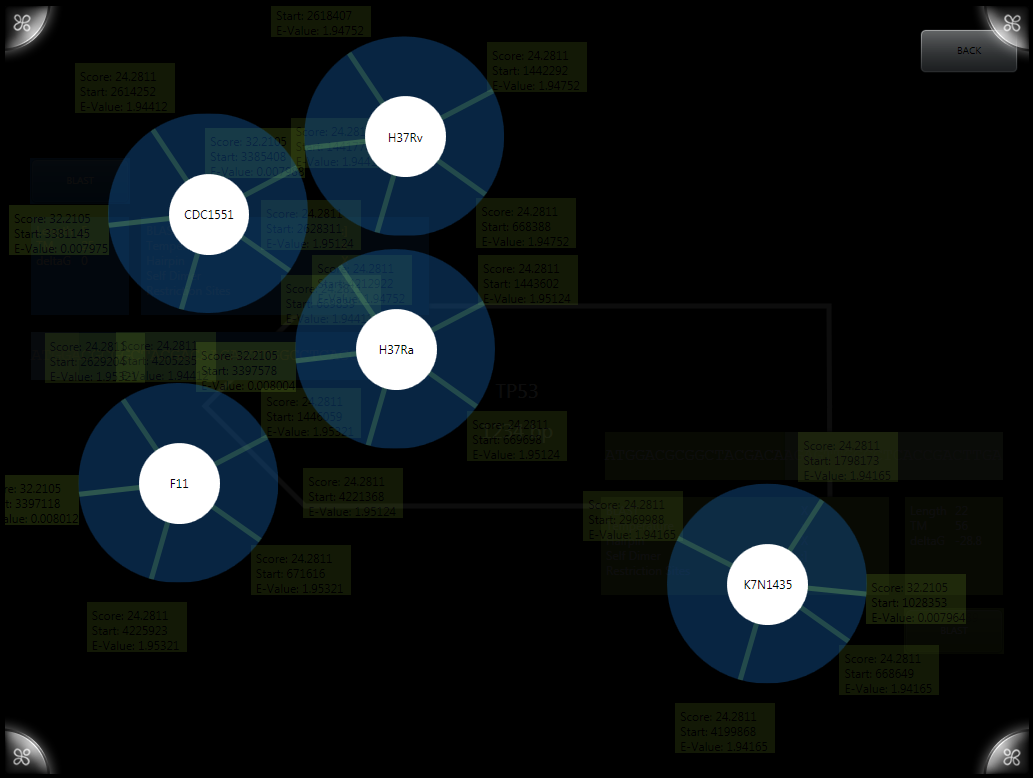
BLASTing
 "
"