Team:Freiburg/Notebook/25 July
From 2011.igem.org
Contents |
Commons
PCR
| Name: Sophie
| Date: 25.07.11 |
| Continue from Experiment (Date)
(Name): Commons | |
| Project Name: more linearized backbones (4 different vectors) | |
PCR-Mixture for one Reaction:
For a 50 µl reaction use
| 32,5µl | H20 | Name |
| 10µl | 5x Phusion Buffer | of Primer |
| 2.5µl | Primer fw | SB-prep-3P-1 |
| 2.5µl | Primer dw | SB-prep-2Ea |
| 1µl | dNTPs | of Template DNA |
| 1µl | DNA-Template | PSB 1 A3
PSB 1 C3 PSB 1 K3 PSB 1 T3 |
| 0.5 µl | Phusion (add in the end) |
What program do you use?
- 2Min 94°C
- 30s 94°C
- 30s 55°C
- 3min 72°C
- 10min 72°C
step 2,3 and 4 in 35 cycles
To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel.
How did you label the PCR-Product, where is it stored and what do you do next?
Labeled PSB 1 A3, PSB 1 C3, PSB 1 K3 and PSB 1 T3 stored in -20°C in last drawer
green light receptor
Testdigest Ligation of CcaS into pSB1K3
Investigators:JULIA
Testdigest
| Continue from Experiment (Date 22.07) Ligation of CcaS into pSB1K3
|
| Project Name: Green light receptor |
For one reaction you need For Mastermix: 35 samples+2extra
| 4μl | H2O | 178μl | H2O |
| 1μl | Buffer, NEB4 | 37μl | Buffer, NEB4 |
| 1μl | BSA (10x) | 37μl | BSA (10x) |
| 0,5 μl | Enzym 1 | 7μl | |
| 0,5 μl | Enzym 2 | ||
| 3 μl | DNA |
10 μl total volume
Give 3 μl of DNA in an eppi and add 7μl of the mastermix.
Incubate for about 1h at 37°C.
Add 1 μl Loading dye buffer and load the gel.
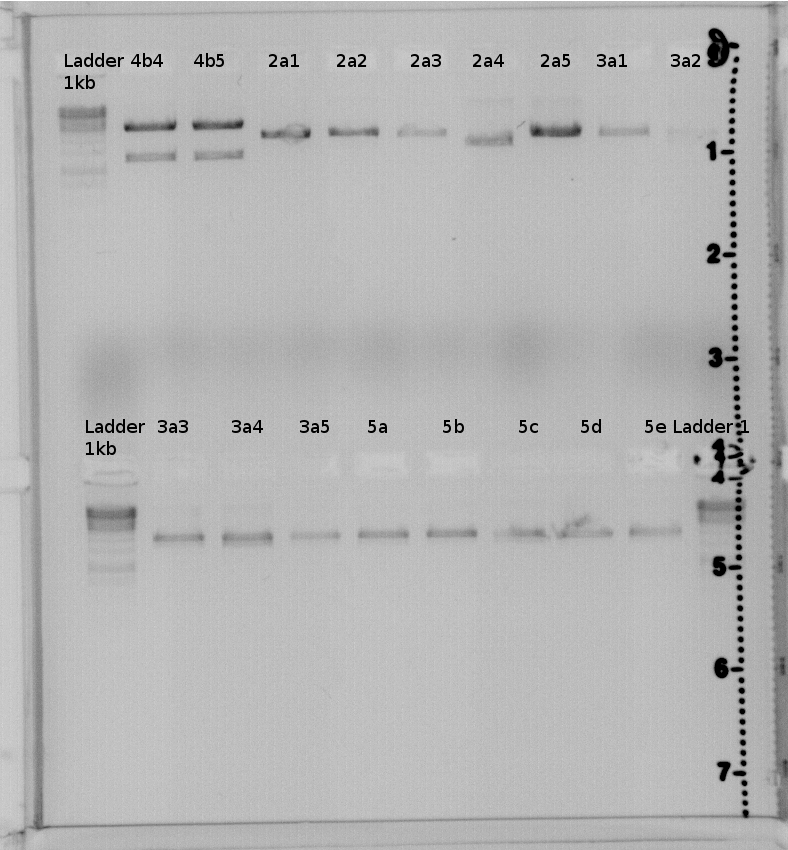
result:4a2,4b2,4b3,4b4,4b5 should have the right insert.
blue light receptor
NAME OF YOUR EXPERIMENT
- Gibson NOT-Gate
Investigators:
- Sophie
Transformation
| Name: Sophie | Date: 25.07.11 |
| Continue from Date Name
Experiment | |
| Project Name: Blue light (NOT-Gate) | |
Procedure
- take cells from -80°C freezer and put them on ice! (every eppi contains about 400 μl cells)
- thaw cells on ice 20 minutes
- pipette 50 μl cells and 2 μl DNA into eppi still on ice!
- Incubate for 30 minutes on ice
- Heat at 42°C for 60 sec
- Incubate on ice for 5 minutes
- Add 200 μl LB Broth
- Incubate for 2 hours at 37°C (cells with lysis cassette at 30°C!!)
- Plate 50 μl and 200μl on two different LB/Agar plates with appropriate antibiotic resistance
Documentation:
Why are you doing this experiment? Name of the sample? Where are they stored? Name the vector with inserts, antibiotika resistance etc.
| We need a NOT Gate for our Blue light system. In this experiment I transformed E.coli with the Part Bba_Q04400 from the iGEM distribution kit. The vector is pSBK3 with Kanamycin resistance. The strain ist named S 45 and will be plated out. The plates will be stored in the incubator. |
PCR
| Name: Sophie
| Date: 25.07.11 |
| Continue from Experiment: new experiment. We need NOT-Gate and LovTAP with Gibson-overhangs, so 2 PCRs are made | |
| Project Name: Blue Light | |
PCR-Mixture for one Reaction:
For a 50 µl reaction use
| 32,5µl | H20 | Name |
| 10µl | 5x Phusion Buffer | of Primer: |
| 2.5µl | Primer fw | NOT_G_up
LOV_G_up |
| 2.5µl | Primer dw | NOT_G_dw
LOV_G_dw |
| 1µl | dNTPs | of Template DNA |
| 1µl | DNA-Template | S35 (BBa_K322999)
S45 (BBa_Q04400) |
| 0.5 µl | Phusion (add in the end) |
What program do you use?
"57°C auf 70°C" (first annealing temperature:55°c, after 10 cycles 65°c)
To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel.
How did you label the PCR-Product, where is it stored and what do you do next?
I labelled it with a heart and a star and stored it in the minipreps 3 box.
I will do Gibson assemblz of the two parts next.
DNA-concentration measured with nanodrop:
| sample | DNA-concentration (ng/μl) |
| S35 | 75.3 |
| S45 | 27.2 |
 We will repeat the PCr, because of the bad result of S35. We will do a PCR with higher temperature.
We will repeat the PCr, because of the bad result of S35. We will do a PCR with higher temperature.
red light receptor
NAME OF YOUR EXPERIMENT
Investigators:NAME
Lysis cassette
NAME OF YOUR EXPERIMENT
Investigators:NAME
Precipitator
NAME OF YOUR EXPERIMENT
Investigators: NAME
 "
"