Team:Cambridge/Experiments/Initial Exercise Group Alpha
From 2011.igem.org
Fusion of SrfA promoter with GFP
Contents |
Aim
Our week 1 experimental challenge was to generate a temporal and or spatial pattern of GFP expression in Bacillus subtilis, using Gibson Assembly to join a sequence of interest to a GFP coding plasmid. We chose to visualise sporulation via a fusion of SrfA promoter with GFP.
Role of SrfA
- SrfA codes for surfactin production, a protein that is produced by B. subtilis prior to sporulation.
- SrfA is regulated by ComA, a protein which itself responds to an intercellular population-density-signalling molecule, CSF, in a concentration dependent manner.
- ComA has low activity at low CSF concentration(low population density), highly active as CSF levels increase, and less active at high CSF concentrations.
Construct Design
We wish to:
- keep the original promotor for SrfA and comA boxes
- remove GFP promotor sequence
- maintain ComGA stabiliser sequence
- do not include restriction sites
Predicted results
- Bacillus colonies exhibit a range of colony patterns and superimposed on the colony shape we expect a temporal pattern of green fluorescence marking sporulation activity.
- intially zero or low GFP detection, higher GFP activity on activation of SrfA prior to sporulation.
- Flourescence will mark spatially sporulation sites.
Method
We designed 20bp primers with 20bp tails with which to perform gibson assembly to fuse the SrfA promoter with a plasmid containing GFP.
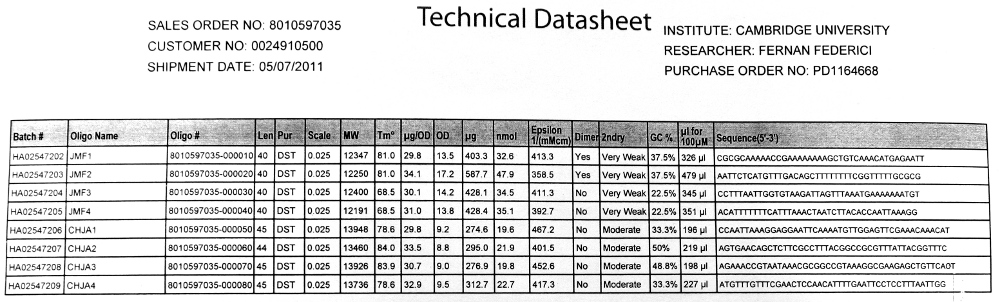
The primers arrived as dry pellets with datasheets that detailed the quantity of water required to make these dry pellets into stock solutions of 100uM concentration. We then made a four fold dilution to make 25uM 'working stock' solutions. We were advised to take extra care to make the inital stock amounts perfect as these affect all future dilutions and that making a working stock is useful as it means you have the capacity to repeat your experiment if anything goes awry.
We were provided with genomic DNA from B. subtilis and the vector plasmid containing GFP in two fragments. In addition we were given two primers which would combine with our designed primers to seal the two fragments.
Our three PCR tubes contained:
- 1.
- 1μl primer JMF2
- 1μl primer JMF3
- 1μl B. subtilis genomic DNA
- 2.
- 1μl primer JMF1
- 1μl primer A Forward (provided)
- 1μl vector DNA
- 3
- 1μl primer JMF4
- 1μl primer B reverse (provided)
- 1μl vector DNA
to each tube we also added:
- 9.5μl of water
- 12.5μl Master mix (SyBR Green and Rox, Hotstart Taq polymerase, dNTPs and dyes)
for a total volume 25μl
The complete tubes were run in a real-time PCR machine for 30 cycles with a 2 minute extension time and a primer annealing temperature of 50ºc.
Technical Data
We used Finnzymes melting temperature calculator to work out the melting temperature of the primer part (not the tail) of our Gibson Assembly oligos (this was the 3' 20bp of each oligo) these values in ºC are shown below
JMF1 - 56.91 JMF2 - 68.8 JMF3 - 50.48 JMF4 - 53.09
Results
References
Sporulation occurs late in the life cycle of B. subtilis when the colony reaches a high population density and we hope that GFP could be visualised following overnight growth. This paper[http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1178101/?page=2] details some sporulation inducing culture conditions.
 "
"