Team:Potsdam Bioware/Project/Details Selection
From 2011.igem.org
Contents |
In Vivo Selection
Introduction
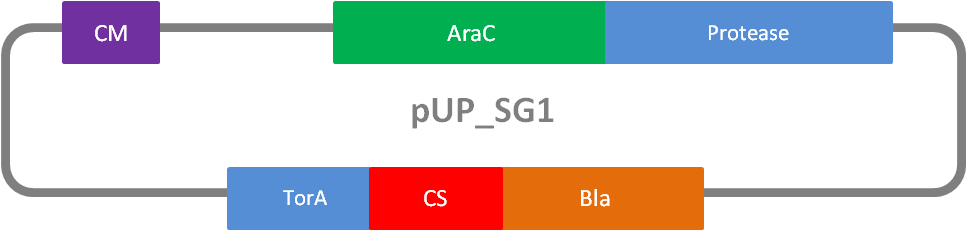
In addition to the phage display we developed a novel selection system. The design aimed for a cheap and time-saving alternative in contrast to an in vitro screen of protease inhibition kinetics. The assay allows us to select effective inhibitors for any protease, among the billions of randomly generated mutants of the Microviridin. For this purpose we designed a plasmid containing two devices, first a protease activity detector and second a protease generator. We used the BioBrick <partinfo>I757010</partinfo> (β-lactamase) and <partinfo>K208005</partinfo> (ssTorA) and fused them together via a linker peptide to create our first device. The protease generator is an Arabinose inducible protease in iGEM standard.
The system works in a combined manner of the two devices. In order to confer β-lactam antibiotic resistance to cells the β-lactamase has to be exported in the periplasm. This transport is mediated by the TorA export sequence via the Twin-Arginine Translocation (TAT) system (DeLisa, 2008). The linker peptide between the TorA export sequence and the β-lactamase displays the corresponding protease cleavage site. In addition the linker peptide is chosen as short as possible to imitate folding because the TAT pathway only allows transport of correctly folded substrates. The linker peptide as well as the protease are designed as exchangeable parts.
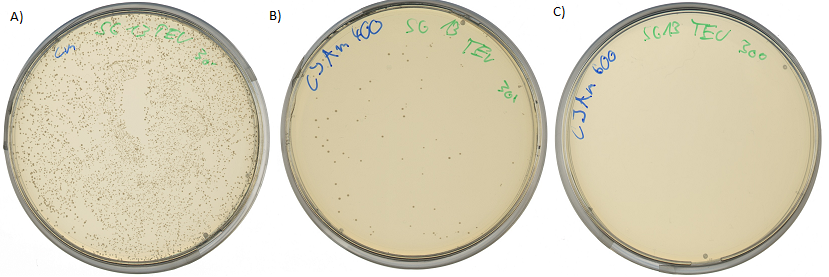
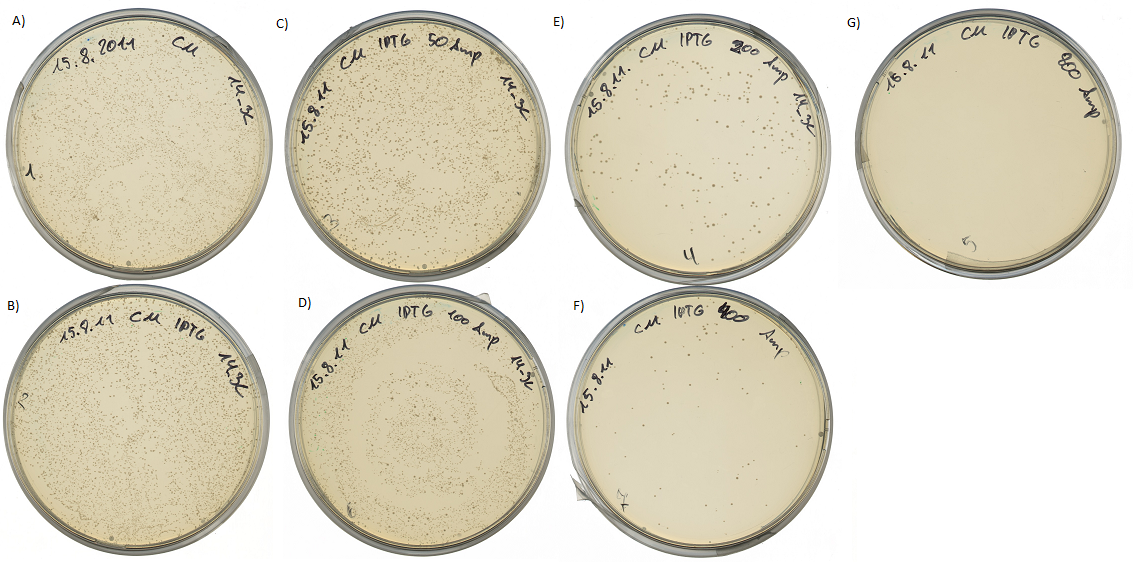
If the protease device is functional, it will cleave the linker peptide between TorA and β-lactamase construct which leaves the cell without any antibiotic resistance. The construct was tested with increasing Ampicillin concentrations. The number of surviving colonies was depended on the export rate of the β-lactamase into the periplasm. A high cleavage rate of the linker peptide leads to a reduced Ampicillin resistance. With expressed protease a dramatically drop of the number of surviving colonies could be observed.
The survival assay was carried out with and without expressed protease. By increasing of the Ampicillin concentration we could detect a cutoff ampicillin concentration. The colony forming units (CFU) were counted and compared.
Site-directed mutagenesis of TEV and 14_3C proteases in iGEM standard
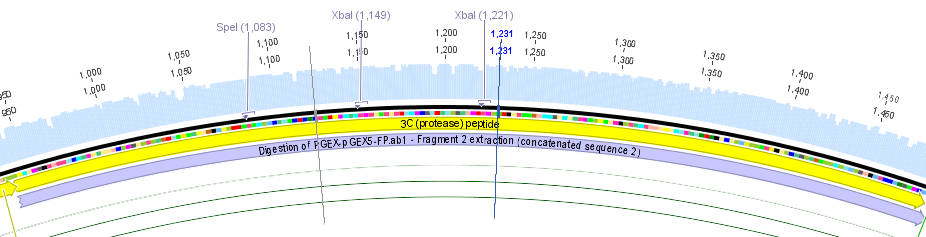
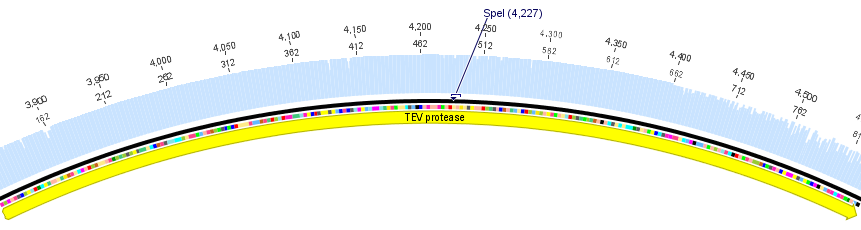
We chose two model proteases, the Tobacco Etch Virus protease, further along called TEV, and the 14_3C protease from the human rhinovirus, also known as PreScission. Both proteases contained iGEM restriction sites, one in case of the TEV and three for 14 3C, respectively.
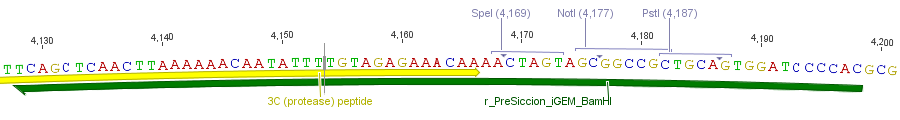
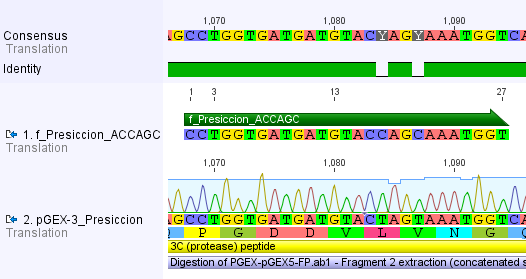
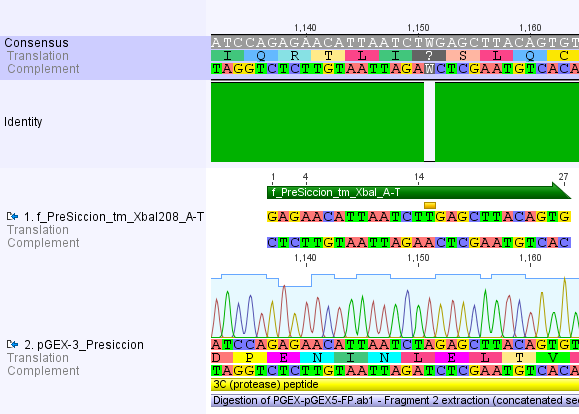
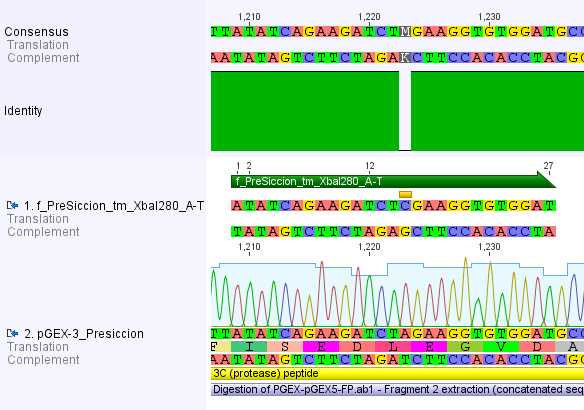
Thus, site-directed mutagenesis was applied to mutate each restriction site in E. coli. For every mutation a forward and a reverse primer had to be designed. The following pictures show the forward primers for the side directed mutagenesis, the reverse primers are the reverse complement sequence of the forward primers.
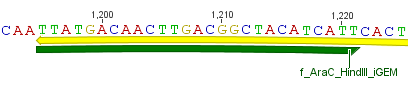
For the 14_3C protease:
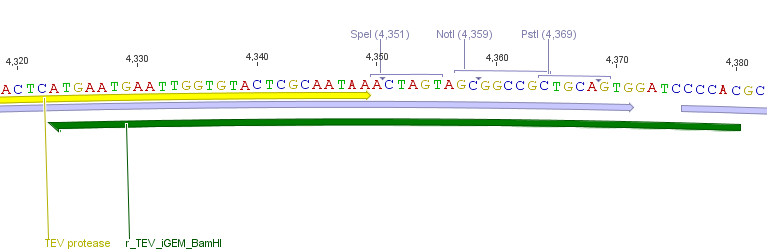
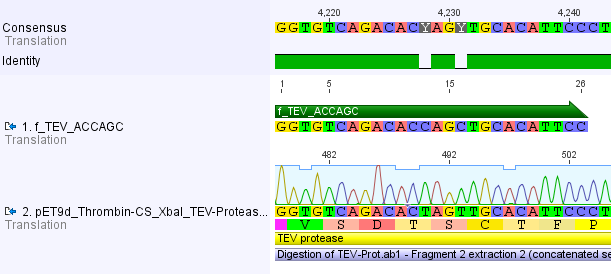
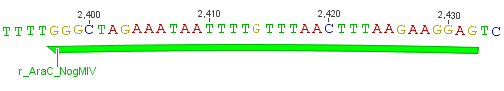
For the TEV protease:
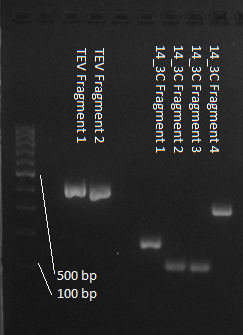
We used assembly PCR to gain the final product. In case of PreScission two assembly PCRs had to be done. The expected size of the fragments are shown in the gel picture. In the following picture all fragments contain the correct sizes.
Cloning of arabinose inducible induction system for fusion with both proteases, TEV and 14_3C
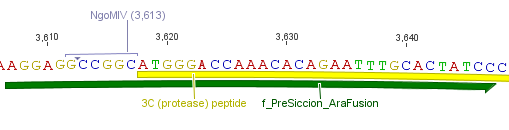
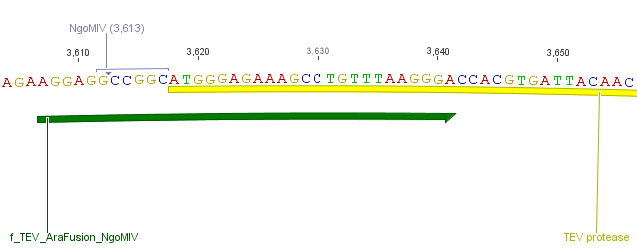
The need of time independent induction of the ssTorA_CS-Protease_blaFL device and the protease, we amplified the arabinose inducible induction system out of the pBAD_iGEMexpress vector. The fusion of both, protease and induction system was performed via an NgoMIV restriction site we introduced with our amplification primers.
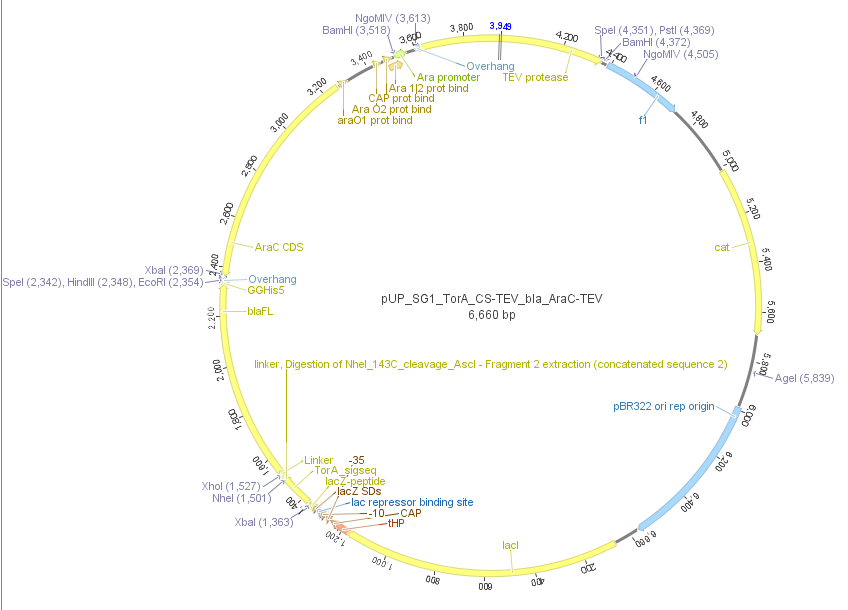
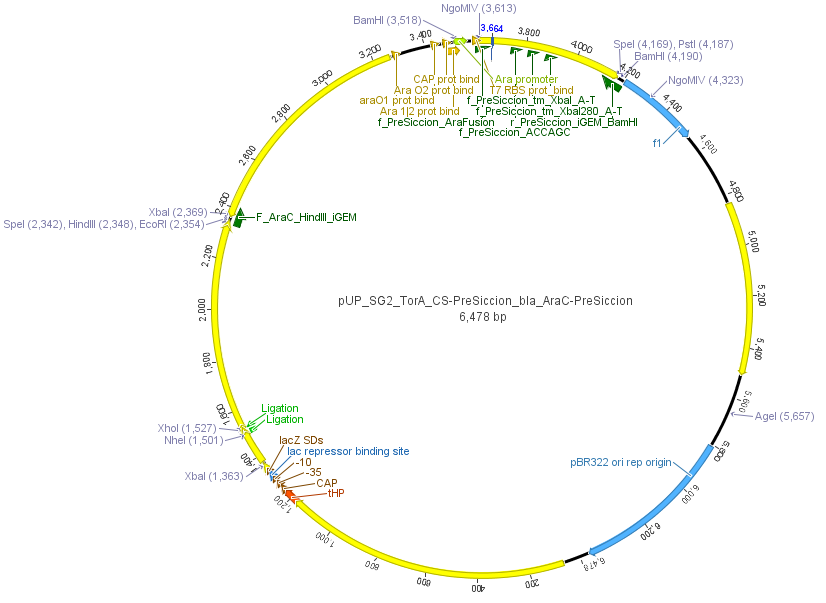
We used the RFC 23 cloning strategy to get a fusion protein of our desired protease and the induction system, which allows us to induce the protease. Both genes were amplified with PCR using two special primers generating a NgoMIV restrictioin site at the end of the induction system and infront of the protease. The amplified protease were digested with BamHI and NgoMIV, the arabinose inducible induction system with NgoMIV and HindIII. The vector, which contains the ssTorA_CS-Protease_blaFL device was digested with HindIII and BamHI. A triple ligation yielded the final construct with both devices: the protease detector and the protease generator. The two pictures below show the final constructed vector, which where used for the survival screening.
Testing the protease detector activity
Both pictures below show the surviving rate of transformed cells with constructed vector:
Background
Twin Arginine Translocon (TAT)
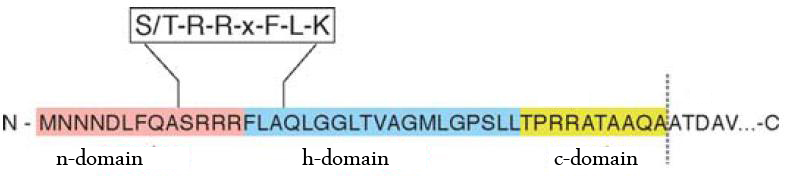
The TAT pathway is responsible for the transport of folded proteins across energy-transducing membranes and it is able of discriminating of unfolded proteins. This transport pathway is common for bacteria, archea and plants and translocates circa 6% of E.coli produced secreted proteins. The translocated proteins must have a signal peptide for targeting of the TAT-transporter, such proteins are e.g. hydrogenases, dehydrogenases and reductases. The signal sequence itself is composed of an N-terminal positive charged domain, a hydrophobic domain and a C-terminal domain.
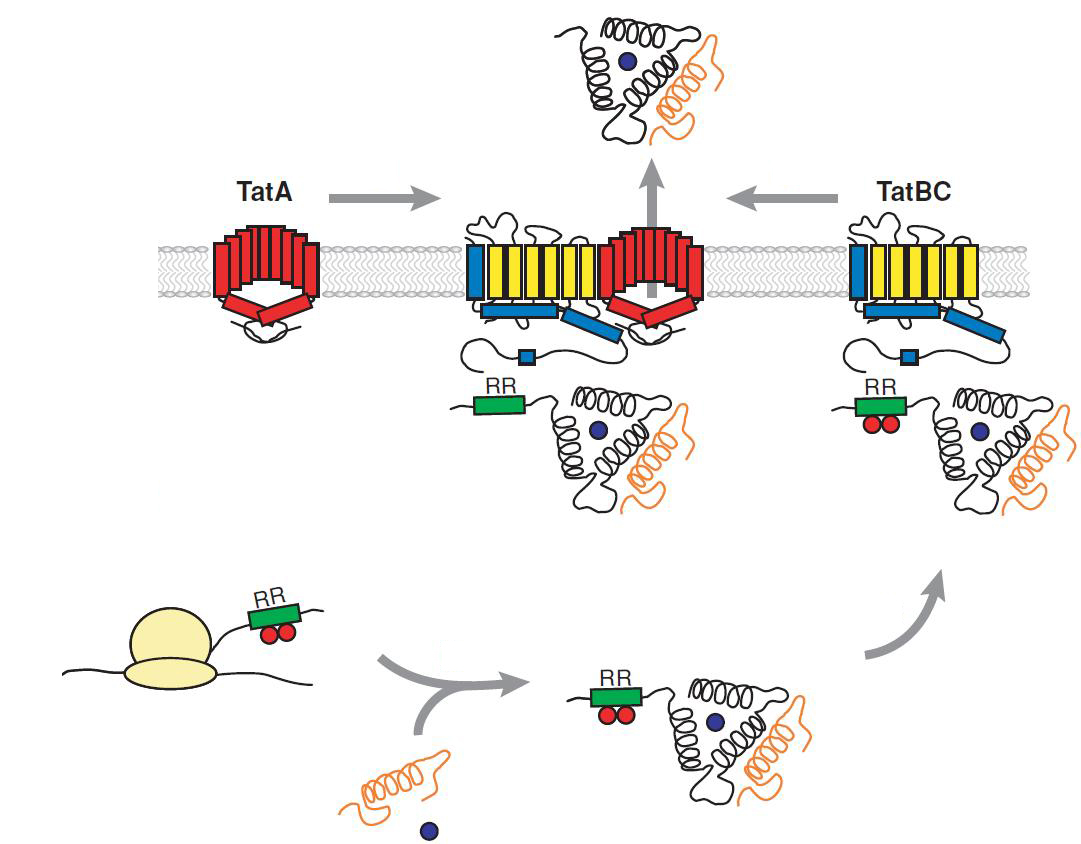
The translocon is composed of three parts: TatA, TatB and TatC. The transport-pore is proposed to be formed during substrate binding by these three parts. The TatA, TatB and TatC proteins may form complexes of different sizes, which on the other hand form pores matching the size of the folded substrate. The transport of proteins through the TAT pathway depends on the proton motive force. Calculations by Adler & Theg showed that the transport of one folded substrate molecule requires the release of approximately 7.9x104 protons, which equals 10.000 ATP molecules. The signal sequence is cleaved off after translocation.
The used signal sequence originates from the TorA protein (Trimethylamin-N-Oxid-Reductase). It is the main respiratory enzyme which reduces TMAO under anaerobic conditions in the periplasm, where it is transported by the TAT pathway.
Tobacco Etch Virus (TEV) protease
TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus. TEV protease is a useful reagent for cleaving fusion proteins. It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.
Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S.
P. (2007). Enhancing the stability and solubility of TEV protease using in silico design.
Protein Sci. 16: 2360-2367
Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D., and
Waugh, D. S. (2001). Tobacco etch virus protease: Mechanism of autolysis and rational
design of stable mutants with wild-type catalytic proficiency. Prot. Eng. 14: 993-1000.
Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable
form of recombinant tobacco etch virus protease. Biotechniques 30: 544-550.
14_3C-Protease
Selective human enterovirus and rhinovirus inhibitors: An overview of capsid-binding and protease-inhibiting molecules. Shih SR, Chen SJ, Hakimelahi GH, Liu HJ, Tseng CT, Shia KS.
The 14_3C protease originates from the human rhinovirus. Rhinoviruses are the most frequent reason for infections of the upper and lower respiratory tract, also known as the cold. Because the 3C protease of human rhinovirus is necessary for the cleavage of the polyprotein translated from viral RNA it may serve as a potential target for development of antiviral targets. The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln↓GlyPro. The small, 22-kDa size of the protease got its optimal activity at 4°C but is still very active at 37 °C. It is commonly used for an easy tag removal after the purification of recombinant proteins carrying his-tag. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.
Quelle: Human rhinovirus 3C protease as a potential target for the development of antiviral agents., Wanga QM, Chen SH., Curr Protein Pept Sci. 2007 Feb;8(1):19-27.
References
- Philip A. Lee, Danielle Tullman-Ercekand George Georgiou Annu. Rev. Microbiol. 2006. 60:373–95
- Olivier Genest, Marianne Ilbert, Vincent Méjean and Chantal Iobbi-Nivol April 22, 2005 The Journal of Biological Chemistry, 280, 15644-15648.
 "
"