Team:Potsdam Bioware/Project/Summary
From 2011.igem.org
Summary
Modification, Selection and Production of Cyclic Peptides for Therapy
One key task of biopharmaceuticals is the binding and blocking of deregulated proteins. Picking the right lead structure for biopharmaceuticals is very important for success. This year, we developed a novel system for the modification, selection and optimization of peptides showing potential for protease inhibition. These promising inhibitors are found in cyanobacteria and are called Microviridins. They belong to the class peptides characterized by unusual ω- ester and ω-amide bonds between the amino-acid side chains, which are also referred to as depsipeptides. These modifications are introduced post translationally by a set of enzymes and result in an extraordinary tricyclic cage structure.
Proteases are a large enzyme family and comprise about 647 human gene products. Therefore they are important drug targets. Additional many harmful bacteria, viruses and fungi use proteases in their reproduction cycle and growth. The ability of blocking these proteases is a highly relevant therapeutic application. Prominent examples of targets are the angiotensin-converting enzyme (ACE) and HIV proteases.
We chose a 6.5 kb Microviridin gene cluster named mdnABCDE from Mycrocystis aeruginosa NIES843. The mdn gene cluster comprises a gene encoding a precursor peptide named MdnA, which is then modified to form the Microviridin, two genes encoding ATP-grasp-type ligases named mdnB and mdnC, an ABC- transporter encoding gene named mdnE as well as one gene encoding an N- acetyltransferase of the GNAT family named mdnD (Ziemert et al., 2008). Our major aim was to modify the mdnA precursor peptide and optimize its protease inhibiting properties. Towards this goal, we synthesized semi-rational mdnA gene libraries with partially randomized oligonucleotides. The library was successfully cloned and verified by mass spectrometry.
To identify the best candidates inhibiting various given proteases, we established two different selection systems: Phage Display and a new developed in-vivo selection assay. Phage display is a frequently used technique in laboratories, yet to our knowledge, phage display of cellularly cyclized peptides is new. We constructed a fusion between the surface protein geneIII of the phage and the mdnA gene within the mdnA gene cluster. This was verified by an ELISA test and a trypsin assay.
We also constructed a recombinant in-vivo selection system linking protease degradation to antibiotic resistance. The designed device is divided into two parts: first a protease activity generator and second a protease detector. For the protease detector we fused various protease cleavage sites between a signal sequence and the antibiotic resistance conferring enzyme β-lactamse. β-Lactamse confers only resistance when transferred to the periplasm of E. coli. Thus, when the protease cleaves the signal sequence from the lactamase antibiotic resistance is abolished and cells die under selective pressure. However, when our Microviridin variants inhibit the protease, cells survive, allowing for easy and efficient inhibitor selection. We achieved different growth rates with increasing Ampicillin concentrations.
In addition to the practical work we established a mathematical model of the in-vivo selection system, which helped us to understand the selection process. We model the reaction kinetics by ordinary differential equations and coded them in matlab.
Currently we are selecting our Microviridin library against a panel of proteases by phage display and with our in-vivo selection system.
Highlights
Microviridin
The major aim of the microviridin group was to modify the mdnA such that the protease inhibiting activity is enhanced. Therefore we used random mutagenesis as well as focused oligonucleotids for creating a library, which is ready for being screened for mdnA with a therapeutically promising set of mutations. For further experiments we also fused the mdnA to a myc-tag. So in the future we will be able to purify and isolate the mdnA.
Due to the applicability of the whole mdn-cluster the creation of several BioBricks was possible. The construction was done using a given template vector containing the mdn-genes and sophisticated design of primers. Characterization of the BioBricks was done via HPLC analysis, mass spectrometry and western blot.
In a subproject we also tried to build auxiliary expression backbones with inducible promoters for easy cloning via the iGEM restriction enzyme sites. We already have the construct but the process of induction needs to be improved.[more]
Phage Display
Phage display is a powerful tool for selecting peptides or proteins that bind and regulate the function of target proteins. It is defined as a system in which the protein and its encoding gene are physically linked. Because of the therapeutic interest of microviridin as protease inhibitors a selection system for screening recombinant mdnA-libraries is of great importance. In our project the fundamental suitability of phage display for this purpose was shown. Therefore a appropriate phagemid was constructed. The production of phage particles carrying mdnA was determined by ELISA and phage display. [more]
In Vivo Selection
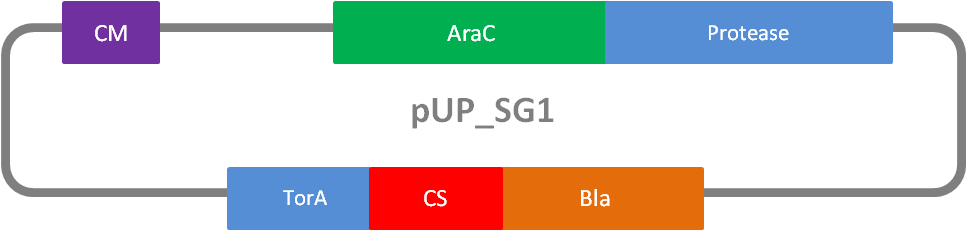
In addition to the Phage Display we developed a novel selection system. The design aimed for a cheap and time-saving alternative in contrast to an in vitro screen of protease inhibition kinetics. The assay allows us to select effective inhibitors for a any protease, among the billions of randomly generated mutants of the Microviridin. For this purpose we designed a plasmid containing two devices, first a protease activity detector and second a protease generator. We used the BioBrick <partinfo>I757010</partinfo> (β-lactamase) and <partinfo>K208005</partinfo> (ssTorA) and fused them together via a linker peptide to create our first device. The protease generator is an Arabinose inducible protease in iGEM standard.
The system works in a combined manner of the two devices. In order to confer β-lactam antibiotic resistance to cells the β-lactamase has to be exported in the periplasm. This transport is mediated by the TorA export sequence via the Twin-Arginine Translocation (TAT) system (DeLisa, 2008). The linker peptide between the TorA export sequence and the β-lactamase displays the corresponding protease cleavage site. In addition the linker peptide is chosen as short as possible to imitate folding because the TAT pathway only allows transport of correctly folded substrates. The linker peptide as well as the protease are designed as exchangeable parts.
If the protease device is functional, it will cleave the linker peptide between TorA and β-lactamase construct which leaves the cell without any antibiotic resistance. The construct was tested with increasing Ampicillin concentrations. The number of surviving colonies was depended on the export rate of the β-lactamase into the periplasm. A high cleavage rate of the linker peptide leads to a reduced Ampicillin resistance. With expressed protease a dramatically drop of the number of surviving colonies could be observed.
The survival assay was carried out with and without expressed protease. By increasing of the Ampicillin concentration we could detect a cutoff Ampicillin concentration. The colony forming units (CFU) were counted and compared.
Bilder über survival screen
We could show that our system works. The next transformation of the constructed microviridin library and our vector into competent E. coli.
Modeling
There is no synthetic biology without modeling, of course. We focused on systems modeling in which the reaction kinetics of the whole system is analyzed and outcomes are predicted. Thus a synthetic biology approach can be chosen because a better understanding of the system is achieved and further changes can be planed - just like in engineering.
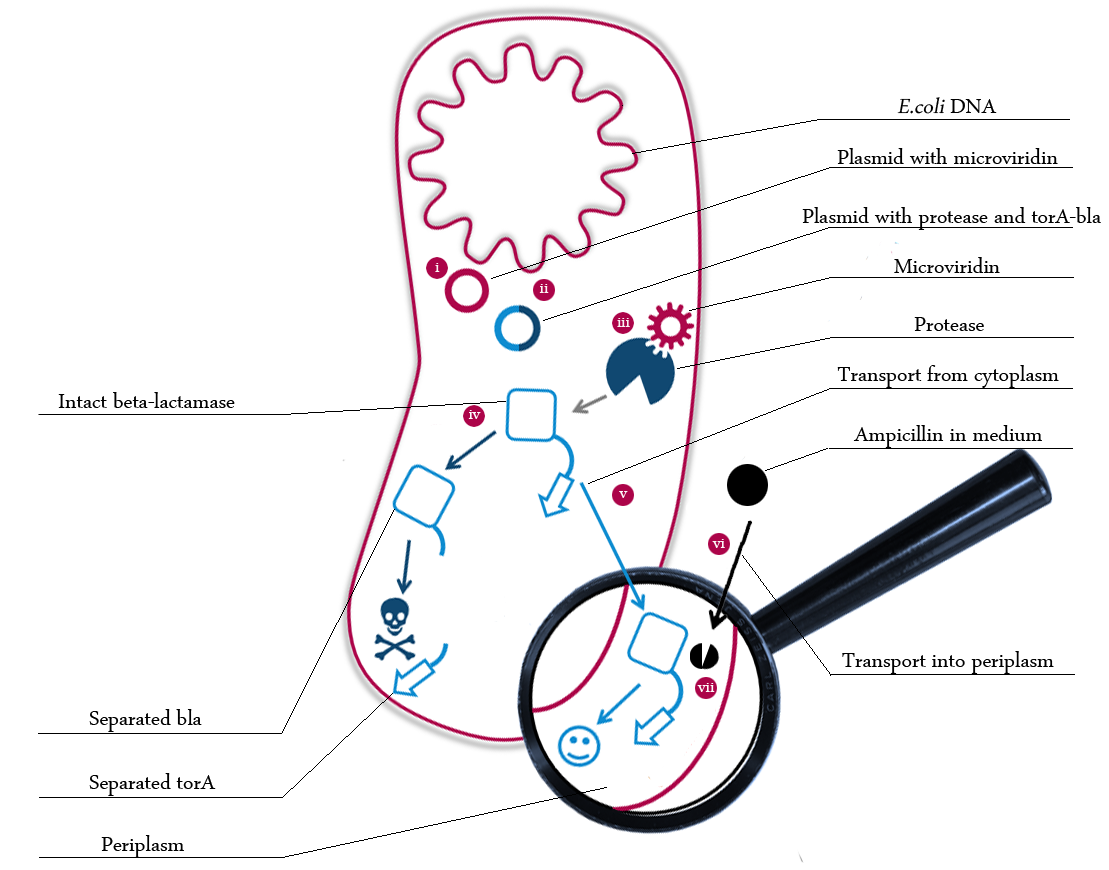
The following schema shows the major reactions taking place in our cell system. The Roman numbers indicate the place of reactions that were written down as equations and then numerically propagated through time. We were able to see that our system works very well in theory. We learned about correct time-scales for our triggering and we were able to identify expected cell-division rates as a reference for the lab work. [more]
Ethics
For information about an ethics seminar and a survey among politicians see [here].
Software
Have a look at the features and screenshots of our BioLog app on this [page].
 "
"