Team:Freiburg/Notebook/4 August
From 2011.igem.org

Contents |
green light receptor
Digest with Dpn1
Investigator: Jakob
5µl Buffer NEB4
5µl BSA (10x)
2µl Dpn1
38µl DNA from quickchange (CcaR)
incubate at 37°C for 1,5 h
inactivate at 80°C for 20 min.
Transformation
Investigator: Jakob
| Name:
Jakob | Date:
04.08.2011 |
| Continue from Experiment: 03.08.2011
Quickchange | |
| Project Name: Green light receptor CcaR | |
Documentation:
- (qc) CcaR in pSB1C3 (Cm resistance)
- Stored in the fridge
- To-do: pick some colonies
Testdigest of mutated CcaS
Investigator: Julia
done a miniprep of the quickchanged CcaS-colonies.
The one CcaS (labeled qC1) was changed with Primer P14 and 15, in order to get rid of EcoRI.
It was digested with EcoRI.
The other CcaS with Primer 16 and 17, was performed to delete SpeI site.
The transformation gave more positive colonies so we have 11 samples.
They were digested with SpeI
For Mastermix: 10+2extra
| 4μl | H2O | 48 | H2O |
| 1μl | Buffer, NEB4 | 12 | Buffer, NEB4 |
| 1μl | BSA (10x) | 12 | BSA (10x) |
| 0,5 μl | Enzym 1 | 1.5 | SpeI |
| 0,5 μl | Enzym 2 | - | |
| 3 μl | DNA |
10 μl total volume
Give 3 μl of DNA in an eppi and add 7μl of the mastermix. Incubate for about 1h at 37°C. Add 1 μl Loading dye buffer and load the gel.
Gelpicture: see red light receptor, lower lane.
Samples C3e and C3c were send to GATC for sequencing.
blue light receptor
NAME OF YOUR EXPERIMENT
Investigators:NAME
red light receptor
Testdigest of pcyA and terminator
Investigators:Julia
For Mastermix: 7+2extra
| 4μl | H2O | 36 | H2O |
| 1μl | Buffer, NEB4 | 9 | Buffer, NEB4 |
| 1μl | BSA (10x) | 9 | BSA (10x) |
| 0,5 μl | Enzym 1 | 1 | EcoRI |
| 0,5 μl | Enzym 2 | 1 | PstI |
| 3 μl | DNA |
10 μl total volume
Give 3 μl of DNA in an eppi and add 7μl of the mastermix.
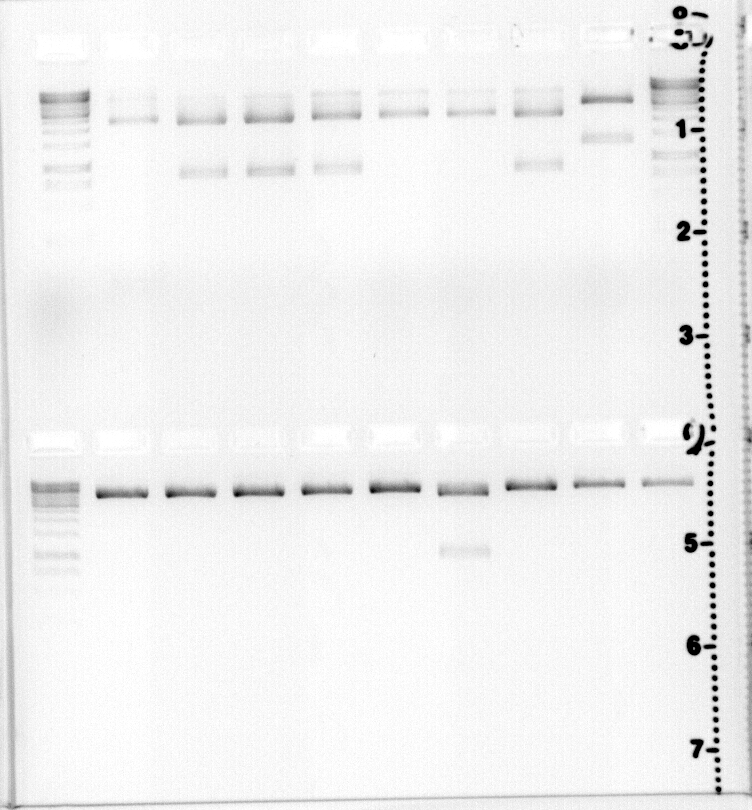
upper lane: (from left to right)Ladder 1kb, pcyA+terminator, a,b,d,e,f,g,h, last CcaS1 digested with EcoRI,Ladder 1kb
lower lane:Ladder 1kb,all qCcaS, from testdigest, see Greenlight recptor
samples named( from left to right) C3e,C2d,C3b,C3c,C3d,C3h,C2e,C2f,C2g
Lysis cassette
Transformation
Investigators: Jakob
Transformation
| Name:
Jakob | Date:
04.08.2011 |
| Continue from Experiment: 03.08.2011
Quickchange (Theo), 3A assembly | |
| Project Name: Lysis casette | |
Procedure
- take cells from -80°C freezer and put them on ice! (every eppi contains about 400 μl cells)
- thaw cells on ice 20 minutes
- pipette 50 μl cells and 2 μl DNA into eppi still on ice!
- Incubate for 30 minutes on ice
- Heat at 42°C for 60 sec
- Incubate on ice for 5 minutes
- Add 200 μl LB Broth
- Incubate for 2 hours at 37°C (cells with lysis cassette at 30°C!!)
- Plate 50 μl and 200μl on two different LB/Agar plates with appropriate antibiotic resistance
Documentation:
Why are you doing this experiment? Name of the sample? Where are they stored? Name the vector with inserts, antibiotika resistance etc.
| Name samples:
α β γ δ ε ζ with Kanamycin resistance in pSB1K3 Lys + RBS with Chloramphenicol resistance pSB1C3 Lys complete with tetracycline resistance pSB1T3 Stored: orange rack in the fridge |
Precipitator
NAME OF YOUR EXPERIMENT
Investigators: NAME
 "
"
 Contact
Contact