Team:NTNU Trondheim/Journal
From 2011.igem.org

Lab Journal
Thursday 23/6
Lab equipment was prepared: Pipette tips, 1.5 ml tubes, toothpicks, SOC, LA with 100 µg/ml ampicillin/ 100 µg/ml spectinomycin (from 1988). Recipies are given in recipies section.
Friday 24/6
Biobricks were taken out from kit by resuspending them in sterile water followed by transformation to E.coli DH5 alpha. The biobricks that were taken out are given below:
| Abbreviation | Name | Part number | Resistance |
|---|---|---|---|
| P1 | rrnB P1 | BBa_K112118 | Spec |
| lambda pR | lambda pR mod | BBa_R0051 | Amp |
| RFP | TetR + p(TetR)+RFP | BBa_K092600 | Amp |
| Lux | TetR + p(TetR)+RFP+luxI | BBa_K092700 | Amp |
Saturday 25/6
Plates with transformants were moved to fridge.
Sunday 26/6
Transformants were inoculated in 3 ml LB + proper antibiotics to prepare for isolation of plasmid.
Monday 27/6
Plasmids that contained P1, lambda pR, RFP and Lux biobricks were isolated by using miniprep kit from Promega.
Consentrations of the biobricks were measured and were as follows:
| Biobrick | Concentration (ng/µl) |
|---|---|
| P1 | 98.4 |
| lamda pR | 43.8 |
| RFP | 141.8 |
| Lux | 62.8 |
Transformants from Lux/RFP constructs were not red as expected. It was suggested that the phenotype could be explained by presence of TetR in the system. Since tetracycline (Tc) is an inhibitor of TetR we tried to grow the transformants in sublethal consentrations of Tc. E. coli with RFP were grown in 3 ml LB with 0, 0.1, 0.3, 0.6, 1.0, 1.5 and 100 µg/ml Tc.
Tuesday 28/6
It's Jon's birthday! Happy day!
Growing the bacteria in Tc did not result in red color of the culture
lambda pR was ligated to RFP backbone:
- RFP and Luc construct were cut with EcoRI and XbaI.
- lambda pR was cut with EcoRI og SpeI
Restriction digestion was done using [http://partsregistry.org/Help:Protocols/Restriction_Digest iGEM's protocol]
Since the restriction digestion gave unexpected fragments and we had not yet seen any pigmentation, we started looking for other designs that could give the result that we wanted. We came up with a different design that used pLac promoter and lacI repressor instead of the pTet/TetR system.
To start constructing the alternative system 3 new biobricks were transformed to E. coli:
| Abbreviation | Name | Part number |
|---|---|---|
| LacI | LacI + RBS | BBa_J24679 |
| pLac | Lac promoter hybrid | BBa_R0011 |
| mCherry | mCherry + RBS + term | BBa_J06702 |
Wednesday 29/6
Lambda Pr, RFP and Lux was cut:
- RFP with XbaI and PstI
- Lux with XbaI and PstI
- Lambda Pr with SpeI and PstI
The cutting gives RFP and Lux as insert and Lambda pR the backbone.
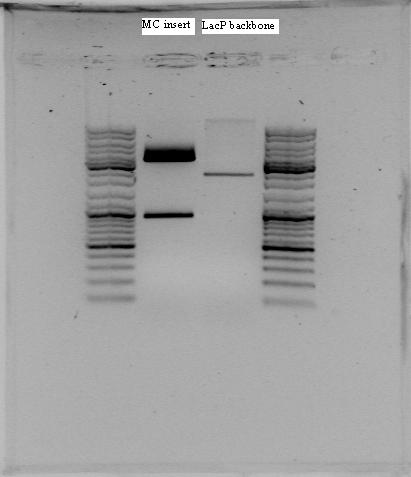
The restriction fragments were separated on agarose gel:
The restriction fragments for lambda pR were as expected but not for RFP or Lux.
Lambda pR was isolated from gel (2110 nt) by using DNA gel extraction kit.
DNA concentration of lambda pR was measured to 5.2 ng/µl
Biobricks that were transformed yesterday were inoculated in 3 ml LB with proper antibiotics to prepare for isolation of plasmids.
Anders ordered a biobrick that should contain LacI + RBS + double terminator from iGEM HQ: BBa_K292006
Thursday 30/6
Plasmid was isolated from lac-design biobricks. Concentration of DNA:
| Biobrick | Concentration (ng/µl) |
|---|---|
| pLac | 16.5 |
| lacI | 37.1 |
| mCherry | 36.7 |
pLac biobrick was cut with SpeI and PstI mCherry was cut with XbaI and PstI
Makes pLac backbone and mCherry insert
Tried direct ligation of restriction fragments using following reaction mix:
| Compound | Amount (µl) |
|---|---|
| H20 | 11 |
| Backbone | 2 |
| Insert | 2 |
| T4 ligase buffer | 2 |
| Ligase | 1 |
Reaction mix were mixed and incubated 30 min at 16C. Heat destruction of enzymes for 20 min at 80C. 2 µl of ligation mix were used for transformation. Red colonies were to be selected from transformation. Mix, spin down, 30 min 16C, heat kill 20 min 80C. Transform 2µL. Select for red colonies.
The rest of the ligation mix were separated on agarose gel and mCherry insert (895 bp) and pLac backbone (2116 bp) were isolated from gel using Quiaquick Gel Extraction Kit. DNA concentration was measured:
| Biobrick | Concentration (ng/µl) |
|---|---|
| pLac | 8.5 |
| mCherry | 2.1 |
The directly ligated restriction fragments transformed to E. coli were plated out on LA + Amp and LA + IPTG and grown ON.
The biobrick that has been ordered (BBa_K292006) was investigated and was found to be identical to the biobrick that we were planning to make, given that the sequence is as given in the registry.
relA
The nucleotide sequence of relA (codes for protein that synthesize ppGpp) was found in database. One hickup in making this gene a biobrick; there is a PstI site inside the gene (at 1383-1388 nt if added prefix). To overcome this problem we could use partial digestion or we could introduce a silent mutation that removes the restriction site. By changing CTGCAG → CTACAG or CTGCAA this could be done.
LacI-negative E.coli
To make the lac-system work we should use a strain that is lacI-negative. According to [http://partsregistry.org/Part:BBa_K177038 another iGEM group] the strain Top10 is lacI-negative.
Friday 1/7
mCherry insert and pLac backbone was isolated from gel using Quiquick Gel Extraction kit and then ligated using the following reaction mix:
| Compound | Amount (µl) |
|---|---|
| Insert | 14 |
| Backbone | 3 |
| T4 ligase buffer | 2 |
| Ligase | 1 |
The ligation mix was ligated at 16C for 1 hour.
Ligation mix was then transformed to LA + Amp, LA + Amp + IPTG and LA + Amp + more IPTG
Two new biobricks were also transformed and plated out on LA + Amp:
| Abbreviation | Biobrick | Part name |
|---|---|---|
| GFP | GFP + LVA | BBa_K082003 |
| RBS | Ribosome binding site | BBa_B0034 |
Monday 4/7
Plasmid isolation of GFP, RBS and LacP+mCherry. Concentrations were measured.
| Biobrick | Concentration (ng/µl) |
|---|---|
| GFP | 60.1 |
| RBS | 16.2 |
| pLac + mCherry | 29.1 |
Restriction cutting of RBS and GFP:
- RBS: SpeI and PstI making it the backbone
- GFP: XbaI and PstI making it the insert.
Gel Extraction:
Extracted RBS backbone, and GFP insert form the gel.
| Biobrick | Concentration (ng/µl) |
|---|---|
| GFP | 3.0 |
| RBS | 4.6 |
Plating:
Plating of pLac + mCherry on amp plates, amp plates with additional IPTG (7µL), amp + IPTG plates and amp + IPTG plates with additional IPTG (7 µL). Incubating on 30 and 37 degrees.
Tuesday 5/7
Ligation of RBS backbone and GFP insert.
- 10μL Ligation Mix
- 1.0 μL 10X T4 ligase buffer
- 6:1 molar ratio of insert to vector (~10ng vector)
- Add (8.5 - vector and insert volume)μl ddH2O
- 0.5 μL T4 Ligase
Used 2,5 µL RBS backbone and 6 µL GFP insert. Incubating at 16 degrees celsius, then heating at 80 degrees celsius.
PCR amplification of rrnB P1 from BioBrick
Resuspend primers in dH20. Primers will amplify the whole rrnB P1 BioBrick (rrnB.FWD and REV), and only the assumed promoter sequence (pro.FWD and REV), and will add normal prefix and suffix instead of BBb format.
Primers used:
| Primer | Type | Sequence |
|---|---|---|
| rrnB P1 F | Forward | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGACGTATCCTACGCCCGTGGT |
| rrnB P1 R | Reverse | GTTTCTTCCTGCAGCGGCCGCTACTAGTACGCCTTCCCGCTACAGAGTCA |
| proL F | Forward | GTTTCTTCGAATTCGCGGCCGCTTCTAGAGCCTCTTGTCAGGCCGGAATAACTCC |
| proL R | Reverse | GTTTCTTCCTGCAGCGGCCGCTACTAGTAGCGGCGTGTTTGCCGTTGTT |
PCR mix:
Work on ice
- Dillute P1 DNA 1µL + 9µL dH2O → ( 9,84 g/µL P1 DNA)
- 2 x PCR tubes 0,5 µL DNA
- 5 µL 10x PCR buffer 2 (w/MgCl)
- 0,5 µL 10 mM dNTPs
- 1 µL fwd primer (rrnBL og proL)
- 1 µL rev primer (rrnBL og proL)
- 0,5 µL polymerase
- 41,5 µL H20
Heat cycles:
Heated lid 103C
- Initial denaturation: 94C 2 min
- Denaturing: 94C 30 s
- Annealing 58C 30 s
- Elongation 72C 60 s
- Repeat 2-4 34 times
- Final elongation 72C 7 min
- Cooling 4 C HOLD
Running products on 1,5 % agarose to find short fragment (promoter sequence). Should give 552 bp (rrnB) and 129 bp (proL).
The products were not so good, so we are running a new PCR tomorrow with different scheme.
Inoculated "LacI with RBS", TERM and LacP+MC biobrick. LacP+MC with and without IPTG and incubating at 30 and 37 degrees celsius.
Wednesday 6/7
Running PCR with rrnBS and L and proS and L on the rrnB P1 BioBrick. 5 cycles of 58C annealing temp, and 20 cycles of 68C annealing temp to try adjust for the massively changed annealing temperatures when primers with pre- and suffixes bind to PCR products. The PCR products where separated by gel electrophoresis and cut out, ready for gel extraction tomorrow.
Isolated plasmids containing the "LacI with RBS" and "TERM" biobricks.
| Biobrick | Concentration (ng/µl) |
|---|---|
| LacI with RBS | 26.0 |
| TERM | 29.9 |
Restriction cutting of "LacI with RBS" with EcoRI and SpeI, and "TERM" with EcoRI and XbaI. The fragments where separated using gel electrophoresis.
The "LacI with RBS" insert and the backbone containing "TERM" where cut out of the gel, and extracted from it.
| Biobrick | Concentration (ng/µl) |
|---|---|
| LacI with RBS insert | 1.6 |
| TERM backbone | 3.9 |
RBS+GFP transformants where inoculated in 3 mL LB+amp.
Thursday 7/7
Ligation of TERM backbone and LacI with RBS insert extracted from gel 6/7.
10μL Ligation Mix:
- 1.0 μL 10X T4 ligase buffer
- 6:1 molar ratio of insert to vector (~10ng vector)
- Add (8.5 - vector and insert volume)μl ddH2O
- 0.5 μL T4 Ligase
Used 2,5 µL TERM backbone and 6 µL LacI with RBS insert. Incubated at room temperature for 30 minutes, then heated at 65 C for 15 minutes.
- Plasmid with GFP+RBS was isolated. Concentration was 26.7 ng/µl
Restriction cutting:
rrnB P1 BioBrick
The rrnB P1 BioBrick found in the distribution was in a different format (BBb), and was therefore not compatible with the other BioBricks we are using. PCR was used to amplify the BioBrick itself, adding the prefix and suffix found [http://openwetware.org/wiki/Synthetic_Biology:BioBricks/Part_fabrication here] making it compatible with the other parts, but losing the first 10 bp. We also amplified only the assumed promoter sequence, with the pre/suffixes.
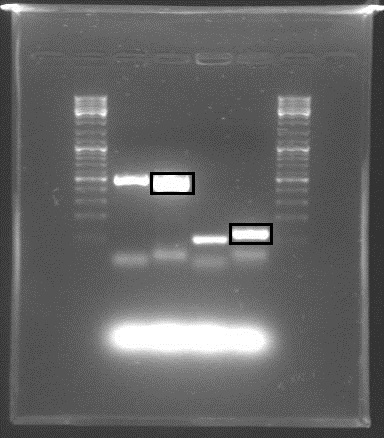
Gel electrophoresis with the PCR products showed that DNA of the right sequence length had been amplified, as shown in the figure. The full-length rrnB P1 amplicon had an approximate size of 500 bp, and the shorter promoter-only amplicon was approx. 100 bp. The shorter fragments shown were created using shorter primers with only X and S restriction sites, in case the long primers failed.
The amplicons with the complete pre/suffixes were cut out, as shown in the figure, and purified with QIAquick Gel Extraction Kit.
Both promoter variants were cut with E and P and also with E and S to create insertable pieces. Our RBS+GFP BioBrick was cut with E and X to create a backbone, and the linearized backbones psB1A3 and psb1C3 were cut to accomodate the cut PCR products.
Friday 8/7
It's Friday, we excited...
The following ligations were made:
| Testing activity | Ligate in BB plasmid |
|---|---|
| rrnBL+mCherry | rrnBL+backbone pSB1A3 |
| rrnBL+(RBS+GFP) | rrnBL+backbone pSB1C3 |
| ProL+mCherry | ProL+backbone psB1A3 |
| ProL+(RBS+GFP) | ProL+backbone psB1C3 |
 "
"