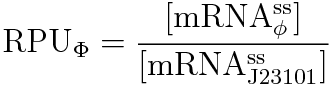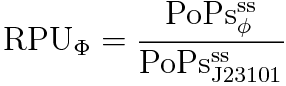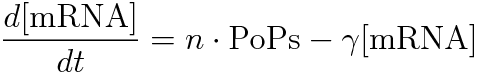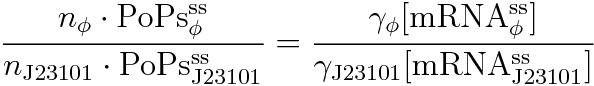Team:Kyoto/Hunger
From 2011.igem.org
Contents |
Project Hunger
Introduction
Carnivorous E.coli attracts insects by emitting light, but it is a burden for the E.coli to emit light. To reduce this burden, we use nitrogen regulatory proteins, NtrB and NtrC. They activate σ54 promoter when the supply of nitrogen is not enough. NtrB and NtrC are coded in glnL and glnG, respectively.
Ammonia is an essential nitrogen source for the bactria. When enteric bacteria are deprived of ammonia, they express glnA to produce glutamine synthetase(GS) under the σ54promoter. The transcription from σ54promoter is stimilated by phosphorylated form of NtrC(NtrC-P). The σ54 RNA polymerase binds to the glnA promoter, forming a closed complex, but cannot form an open complex and initiate transcription until it is activated by NtrC-P. NtrC is phosphorylated by NtrB-P, an autokinase which phosphorylates itself with ATPs. Phosphorylation and dephosphorylation of NtrB and C is controlled so that there are sufficient NtrC-P in the cell when under the low concentration of ammoniacal source.
The concentration of ammoniacal source is detected by the ratio of αketoglutarate to glutamine. If glutamine levels are low, less αketoglutarate is synthesized by GS and, as a result, Pii retains UMP and so cannot bind to NtrB. NtrB can then phosphorylate itself and transfer this phosphate to NtrC.
[GS Reaction Under Low NH3 Concentration]
Glutamate + NH3 + ADP → Glutamine + ADP + phosphate
GS
Glutamine + αketoglutarate → 2 glutamate
GS
Although this nitrogen controlled switch is complex,
Fig. NtrCがσ54,RNAポリメラーゼ,DNAの関係図(窒素源あり/なし) Fig. NtrBCの関係図 Fig. Ntr, Pii, GSの関係図
However, the σ54promoter has never been evaluated quantitatively. We characterize the σ54promoter by Relative Promoter Unit (RPU), because absolute promoter activity depends on test conditions and measuring instruments. RPU can reduce this Coefficient of Variation (CV) from 39.1% to 17.5% [2]. Therefore RPU can make it easier for us to share the data of promoter activity and use BioBricks.
We have used GFP fluorescence to measure RPU previously, but this time, we tried to calculate RPU using another way for the following reasons:
- We know that it is a lot of trouble to calculate RPU using GFP fluorescence.
- We imagined that the concentration of the glutamine changes because the σ54promoter relates to the metabolism of glutamine.
This is because when we calculate RPU using GFP, we need to measure the concentration of glutamine at two points in an exponential growth phase, but we have the following problems.
- before an exponential growth phase, the concentration of glutamine changed.
- the concentration of glutamine is different between one point and the other point.
We devised the new way of calculating RPU using the amount of mRNA, according to the following equation.
† NtrB and NtrC are otherwise called NRII, NRI
We can calculate the amount of mRNA easily by using this way.
ところでこの遺伝子はもともとBioBrickに登録されていたものか、今回新たに作ったもののどちらなのでしょう
http://partsregistry.org/wiki/index.php/Part:BBa_J64978
などがもう登録されているようですが
リンク先を見ましたがどうなんでしょうね とりあえずこの登録されているパーツのシーケンスは見ましたが今回我々が作ろうとしたglnLとは全然違うシーケンスでした 名に由来かは知りませんがE.coliのものじゃないでしょうねby Hashiya
Method
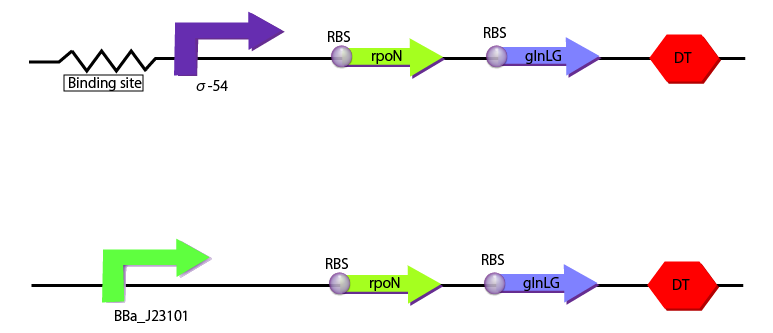
We created the following two constructions to characterize σ54 promoter with RPU depending on the concentration of glutamine. One construction includes BBa_J23101 as a promoter which is used as the standard and the other includes binding sites and σ54 promoter.
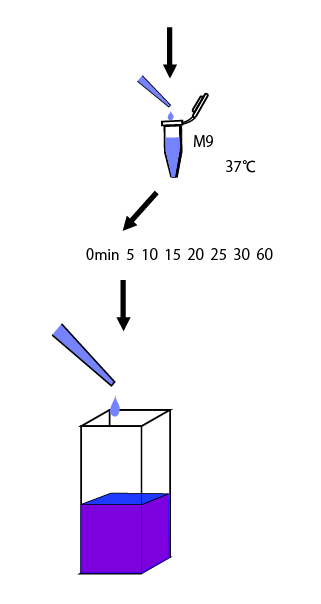
This time we intend to assay a promoter activity against glutamine concentration, it is desirable to induct after cultivation in the medium without glutamine. However, from our experience last year, we know that in a M9 medium without casamino acids, E.coli will grow really inefficiently. So, firstly we cultivated E.coli in the medium with casamino acids overnight, then changed the medium with the one without casamino acids. After that, we performed an experiment to determine how long they take to reach a steady state.
We also measured the amount of mRNA to calculate RPU of the steady state for each concentration of glutamine. First, we made the following preliminary experiment to measure the length of times before steady state.
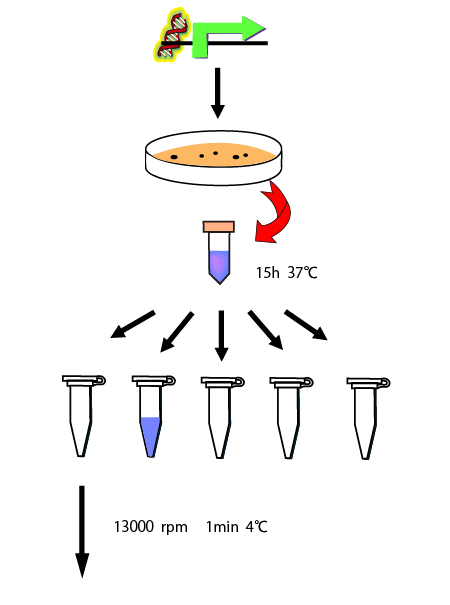
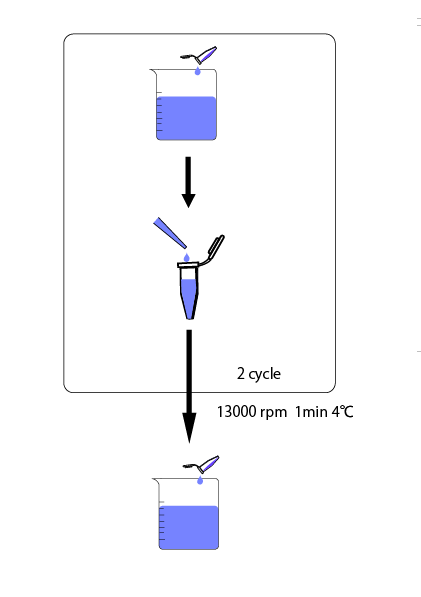
We cultivated E.coli in M9 media(+ casamino acids) for about 15 hours and dispenced 2.4ml to each tube. Then, we centrifuged these tubes (13,000 rpm , 4°C, 1min) and discarded the supernatant. We added 2.4ml media(- casamino acids) and centrifuged at 4°C twice. Again, we centrifuged these tubes and discarded the supernatant and added 2.4ml media(-casamino acids) at 37 °C. We brought up E.coli at 0,5,10,15,20,25,30,60min and extracted RNA and synthesized cDNA. Finally, we used real time PCR.
RPU is defined as follows.
This time, we decided to use equation(2).
PoPS (Polymerase Per Second) is the unit of absolute “promoter activity”. It is defined as the number of RNA polymerase molecules that pass by the final base pair of the promoter and continue along DNA as an elongation complex. PoPSSS is PoPS at the steady state of following system.
Where:
- [mRNA] is the concentration of mRNA,
- γ is the mRNA degradation rate,
- n is the copy number of the plasmid containing the promoter
Following equations related to promoterφ and J23101 are derived because d[mRNA]/dt = 0.
If the test promoter φ and the reference standard promoter are measured under the same culture conditions and both promoters are carried on the same backbone plasmid, following equations are approved.
Equation(4) divided by (5) is (8) and we can reduce (8) because of (6) and (7).
The equation (2) is derived from the above.
So, we can calculate RPU with this equation.
When we use this equation, we need the following data:
- the amount of mRNA which is corrected by internal control
- when an expression level reaches a steady state
Result
Gragh1 The data, which is not corrected by internal control
- Although the amount of RNA is different at first, we think that more than twice the difference is not an accidental error of the experiment.
- After 15 minutes, the expression level greatly changed, but from 0 to 10 min, the expression level reached a steady state.
- We think TBP and PGK are unsuitable for internal control.
- The behaviors of GAPDH and actin is analogous.
Gragh2~5
The data, which is corrected by internal control
- when we applied internal control to GAPDH, the changes of actin are little(×1.0~1.2)
- when we applied internal control to actin, the changes of actin are little(×0.8~1.0)
- We concluded that GAPDH and actin are suitable for internal control.
Discussion
Reference
[1] P. Jiang and A. J. Ninfa, “Escherichia coli PII signal transduction protein controlling nitrogen assimilation acts as a sensor of adenylate energy charge in vitro.,” Biochemistry, vol. 46, no. 45, pp. 12979-96, Nov. 2007.
[2] J. R. Kelly et al., “Measuring the activity of BioBrick promoters using an in vivo reference standard.,” Journal of biological engineering, vol. 3, p. 4, Jan. 2009.
[3] M. Al-Azri, H. Al-Azri, F. Al-Hashmi, S. Al-Rasbi, K. El-Shafie, and A. Al-Maniri, “Factors Affecting the Quality of Diabetic Care in Primary Care Settings in Oman: A qualitative study on patients’ perspectives.,” Sultan Qaboos University medical journal, vol. 11, no. 2. pp. 207-13, May-2011.
[4] L. J. Reitzer and B. Magasanik, “Transcription of glnA in E. coli is stimulated by activator bound to sites far from the promoter.,” Cell, vol. 45, no. 6, pp. 785-92, Jun. 1986.
[5] B. Magasanik, “Regulation of transcription of the glnALG operon of Escherichia coli by protein phosphorylation.,” Biochimie, vol. 71, no. 9-10, pp. 1005-12, 1989.
[6] J. Keener and S. Kustu, “Protein kinase and phosphoprotein phosphatase activities of nitrogen regulatory proteins NTRB and NTRC of enteric bacteria: roles of the conserved amino-terminal domain of NTRC.,” Proceedings of the National Academy of Sciences of the United States of America, vol. 85, no. 14, pp. 4976-80, Jul. 1988.
[7] J. R. Kelly et al., “Measuring the activity of BioBrick promoters using an in vivo reference standard.,” Journal of biological engineering, vol. 3, p. 4, Jan. 2009.
 "
"