From 2011.igem.org
(Difference between revisions)
|
|
| Line 111: |
Line 111: |
| | <LI>pSB1C3-tetR::HNS (FL) | | <LI>pSB1C3-tetR::HNS (FL) |
| | </OL> | | </OL> |
| | + | </OL> |
| | <LI>Enzyme digestion was carried out for the extracted and purified plasmid tetO2-0 and tetO2-3 | | <LI>Enzyme digestion was carried out for the extracted and purified plasmid tetO2-0 and tetO2-3 |
| | <OL> | | <OL> |
| | <LI>0.5uL of CIP alkaline phosphatase was added to prevent self-ligation to each enzyme digest reaction mixture. | | <LI>0.5uL of CIP alkaline phosphatase was added to prevent self-ligation to each enzyme digest reaction mixture. |
| | + | |style="font-family: georgia, helvetica, arial, sans-serif;font-size:2em;color:#01DF01;"|Lab Diaries |
| | + | |- |
| | + | |style="width:900px;"|'''Week 9''' |
| | + | <OL> |
| | + | <LI>tetO2 – 0 and tetO2 – 3 were ligated to form tetO2 – 0 – 3. |
| | + | <OL> |
| | + | <LI>tetO2 – 0 – 3 was transformed to DH10b competent cells. |
| | + | </OL> |
| | + | <LI>tetO2 – 0 –sfGFP |
| | + | <div ALIGN=CENTER> |
| | + | {| style="width:254px;background:#99EE63;text-align:center;font-family: georgia, helvetica, arial, sans-serif;color:#000000;margin- top:5px;padding: 2px;" cellspacing="5"; |
| | + | |- |
| | + | |[[Image:TetO2-0-sfGFP.jpg|250px]] |
| | + | |- |
| | + | |tetO2-0-sfGFP |
| | + | |} |
| | + | </div> |
Revision as of 17:44, 4 October 2011
| Lab Diaries
|
| Week 1
Transformation of reporter DNA (pEGFP-loxp-km-loxp) into DH10B (non-virulent strain E. coli) with antibiotic resistance (Chloramphenicol – Cm)
| Lab Diaries
|
Week 2
- tetO2 -1 (DNA binding site)
- Reverse PCR was used to insert tetO2 -1 into pEGFP-loxp-km-loxp (template DNA) by using forward and reverse primers which are with the tetO2 -1. By using reverse PCR, we can insert the tetO2 -1 site into the pEGFP while the plasmid produced is still in double-stranded circular form.
- pEGFP-loxp-km-loxp-tetO2-1 was then transformed into DH10B to greatly amplify the product by using bacterial cells.
- tetO2 -2 (DNA biding site)
- Reverse PCR was used to insert tetO2 -2 into pEGFP-loxp-km-loxp.
- 2uL of 10-fold diluted pEGFP-loxp-km-loxp- tetO2 -2 was transformed into DH10B to greatly amplify the product by using bacterial cells.
- tetO2 – 3 (DNA binding site)
- Reverse PCR was used to insert tetO2 – 3 into pEGFP-loxp-km-loxp.
- 2uL of 10-fold diluted pEGFP-loxp-km-loxp- tetO2 – 3 was transformed into DH10B to greatly amplify the product by using bacterial cells.
| Lab Diaries
|
Week 3
- Overlap PCR (involving 3 steps) was carried out to produce the fusion protein [tetR-HNS (any length)].
- PCR was carried out separately for 2 genes, tetR and HNS (full length, in this case).
- Primers used were as below:
- tetR: [forward] R-H-out-F; [reverse] R-H-tetR-R
- HNS (FL): [forward] R-H-tetR-F-out; [reverse] R-H(FL)-2-R
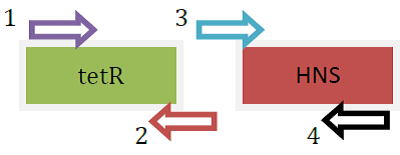
|
| Fusion protein genes produced separately using PCR
|
- Another PCR was carried out using primers with linker to insert the linker DNA to the tetR and HNS (FL) for the fusion of the 2 genes.
- PCR was used to further amplify the product from step 2 (fusion protein gene).
- PCR was used to obtain HNS(FL) at 5 different temperatures to test which temperature is optimal for annealing.
- Annealing phase at 5 different temperatures:
- 60.7C
- 62.2C
- 63.8C
- 65.4C
- 66.7C (less sample)
- Electrophoresis was carried to determine which temperature best suit the annealing phase. From the gel image, it was observed that a high concentration of products was resulted at annealing temperature between 60.7C and 62.2C.
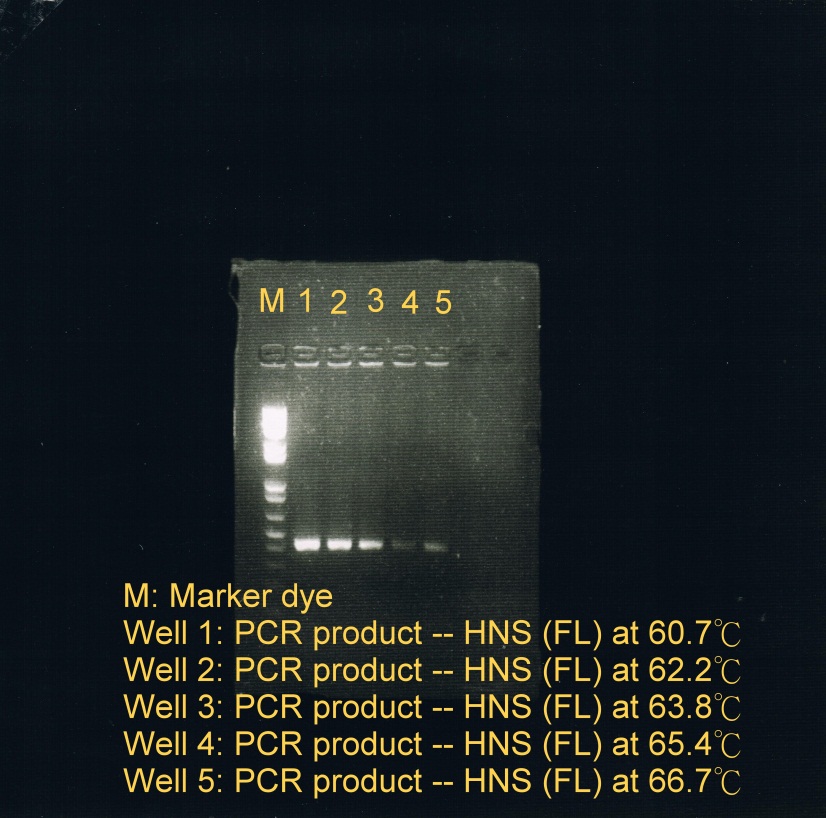
|
| Fusion protein genes annealed at 5 different temperatures
|
| Lab Diaries
|
| Week 4-5
tetO2 – 0 and tetO2 – 4
- Reverse PCR was used to insert tetO2 -0 and tetO2 – 4 into pEGFP-loxp-km-loxp.
- 2uL of pEGFP-loxp-km-loxp- tetO2 – 0 and pEGFP-loxp-km-loxp- tetO2 – 4 were transformed into DH10B separately to greatly amplify the product by using bacterial cells.
| Lab Diaries
|
| Week 6
Overlap PCR was carried out to produce fusion protein gene
- HNS::tetR
- HNS (1-46)
- HNS (1-83)
- HNS (1-90)
- HNS (FL)
- tetR::HNS
- HNS (2-46)
- HNS (2-83)
- HNS (FL)
| Lab Diaries
|
| Week 7
Overlap PCR was carried out to produce fusion protein gene tetR::HNS (2-90)
- The sticky ends of the enzyme products was transformed into blunt ends using PCR.
| Lab Diaries
|
Week 8
- pROT-HNS (for BioBricks) was produced
- Double enzyme digestion was carried out to digest the fusion protein genes and ligate them to pBS1C3 (plasmid backbone)
- 3 samples were transformed
- pSB1C3-tetR::HNS (2-83)
- pSB1C3-tetR::HNS (2-90)
- pSB1C3-tetR::HNS (FL)
- Enzyme digestion was carried out for the extracted and purified plasmid tetO2-0 and tetO2-3
- 0.5uL of CIP alkaline phosphatase was added to prevent self-ligation to each enzyme digest reaction mixture.
| Lab Diaries
|
Week 9
- tetO2 – 0 and tetO2 – 3 were ligated to form tetO2 – 0 – 3.
- tetO2 – 0 – 3 was transformed to DH10b competent cells.
- tetO2 – 0 –sfGFP
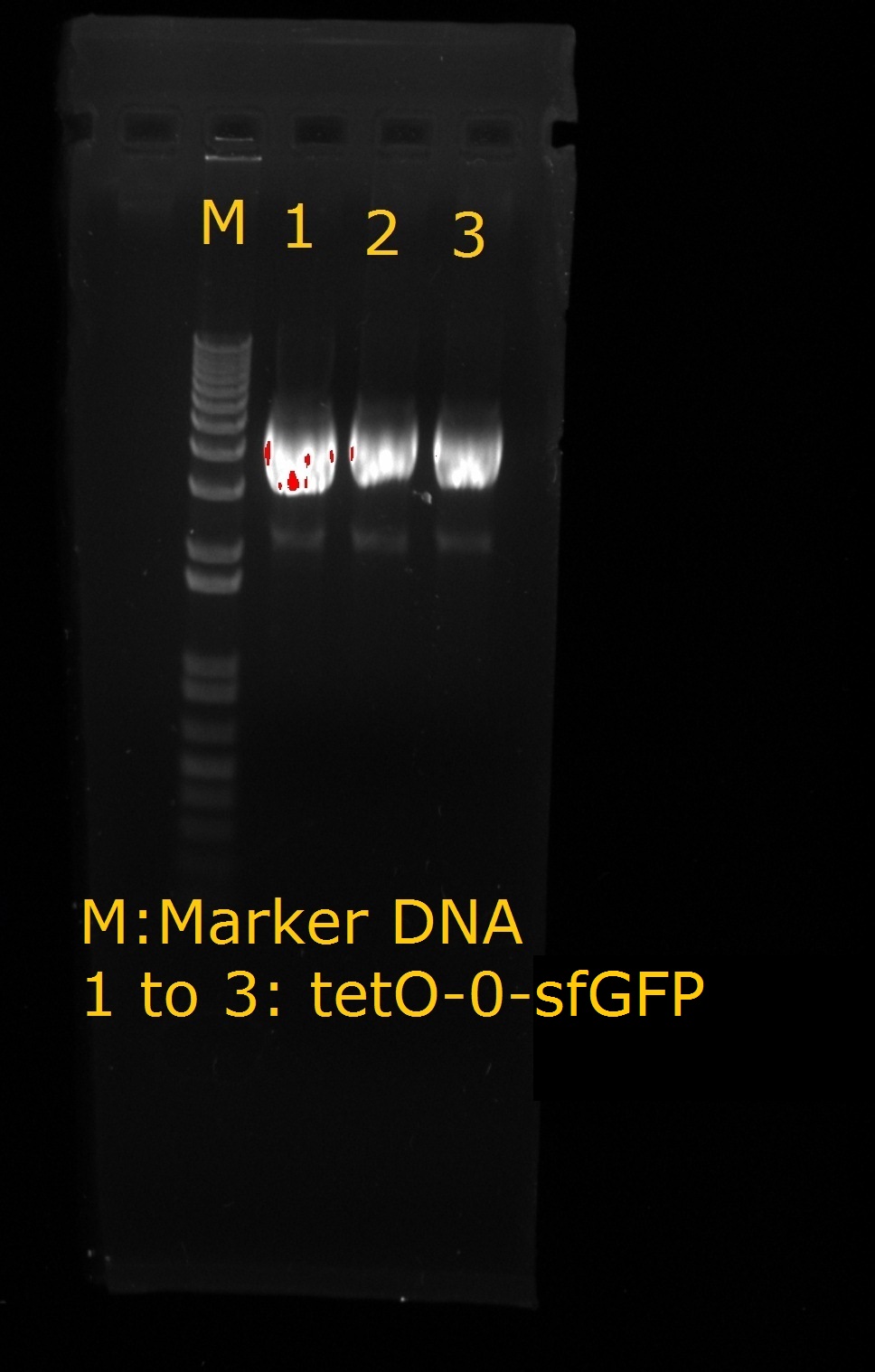
|
| tetO2-0-sfGFP
|
|
 "
"





