Team:MIT/Notebook/
From 2011.igem.org
| Line 1,457: | Line 1,457: | ||
<h1>Week 5: July 4 - July 10</h1> | <h1>Week 5: July 4 - July 10</h1> | ||
We began looking into using the Goldengate assembly method, but two gene elements contained a cut site that needed to be mutated out. We performed Site-Directed Mutagenesis using the Lightning Kit in the hopes of mutating out the cut site, but results were not successful due to mishandling during the protocol, so the procedure was set to be repeated. Gibson assembly of AVPR2-TEVs-GV16 continues as we re-PCRed the AVPR2 DNA segment and re-run the entire Gibson protocol, picking 20 colonies in a determined attempt to obtain a successful result. | We began looking into using the Goldengate assembly method, but two gene elements contained a cut site that needed to be mutated out. We performed Site-Directed Mutagenesis using the Lightning Kit in the hopes of mutating out the cut site, but results were not successful due to mishandling during the protocol, so the procedure was set to be repeated. Gibson assembly of AVPR2-TEVs-GV16 continues as we re-PCRed the AVPR2 DNA segment and re-run the entire Gibson protocol, picking 20 colonies in a determined attempt to obtain a successful result. | ||
| + | |||
| + | |||
| + | <h3><a name="DailyNotebook-July10"></a>July 10</h3> | ||
| + | |||
| + | <p>Jenny and Jon - SDM Colony counts were approximately 50 for the control, 20 for the TEV, and 0 for the Gal4VP16. There is a chance the TEV and Gal4VP16 labels were switched, which we'll find out after sequencing. Six colonies were picked from the TEV plate and inoculations were accidentally run for 22 hours at 37C.</p> | ||
| + | |||
| + | |||
| + | <h3><a name="DailyNotebook-July9"></a>July 9</h3> | ||
| + | |||
| + | <p>Jenny and Jon redid the SDM for Control, Gal4VP16, and TEV using the Lightning Kit, following pages 7 and 8 of this manual: www.qcbio.com/stratagene/210518.pdf<br/> | ||
| + | We used 1.5uL 100mM stock of the previously designed primers. The final reaction volume was only approximately 45uL, as we only had about 5uL of 5ng/uL dsDNA for both the Gal4VP16 and TEV reactions. We got the DNA from the dilutions that Divya and Tyler previously used.</p> | ||
| + | |||
| + | <p>For transformation, we followed the standard protocol instead of the one outlined in the manual and outgrew at 37C and incubated at 30C for 16 hrs.</p> | ||
| + | |||
| + | <h3><a name="DailyNotebook-"></a><font color="#660000"><b>July 8, 2011</b></font></h3> | ||
| + | |||
| + | <div class='table-wrap'> | ||
| + | <table class='confluenceTable'><tbody> | ||
| + | <tr> | ||
| + | <th class='confluenceTh'> Sample </th> | ||
| + | <th class='confluenceTh'> Assignees <br class="atl-forced-newline" /> </th> | ||
| + | <th class='confluenceTh'> Procedures <br class="atl-forced-newline" /> </th> | ||
| + | <th class='confluenceTh'> Enzyme(s) Used <br class="atl-forced-newline" /> </th> | ||
| + | <th class='confluenceTh'> Expected Bands <br class="atl-forced-newline" /> </th> | ||
| + | <th class='confluenceTh'> Results <br class="atl-forced-newline" /> </th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR 1xCI434 (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR 4xCI434 (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR 4xLexA (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR 4xMnt1 (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR 7xMnt1 (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR UAS:EBFP2 (A-C) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> ScaI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5000 <br class="atl-forced-newline" /> | ||
| + | 2000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR AVPR2 A (1-3) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> NcoI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1356 <br class="atl-forced-newline" /> | ||
| + | 3000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR AVPR2 B (1-3) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> NcoI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1356 <br class="atl-forced-newline" /> | ||
| + | 3000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR AVPR2 C (1-3) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> NcoI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1356 <br class="atl-forced-newline" /> | ||
| + | 3000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> EXPR AVPR2 D (1-3) <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Grant/Michelle/Charles <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> Miniprepped <br class="atl-forced-newline" /> | ||
| + | Nanodropped <br class="atl-forced-newline" /> | ||
| + | Restriction Digest </td> | ||
| + | <td class='confluenceTd'> NcoI <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1356 <br class="atl-forced-newline" /> | ||
| + | 3000 <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> </td> | ||
| + | </tr> | ||
| + | </tbody></table> | ||
| + | </div> | ||
| + | |||
| + | |||
| + | <h3><a name="DailyNotebook-July7%2C2011"></a>July 7, 2011</h3> | ||
| + | |||
| + | <p>Charles and Clara @ Knight lab: All four Kan plates had colonies, 40-50 colonies except for one of Deepak's gibson mix reactions, which had ~400 colonies. Inoculated 5 colonies from each reaction into LB-KAN. 20 Falcon tubes in 37C spinning. To be taken out at 2 AM by Grant.</p> | ||
| + | |||
| + | <h3><a name="DailyNotebook-July6%2C2011"></a>July 6, 2011</h3> | ||
| + | |||
| + | <p>Charles and Clara @ Weiss lab: Ran Gibson reaction with both DM and CH Gibson mixes each, total of 4 reactions run. 0.21 uL of 25.6 ng/uL AVPR2, 0.11 uL of 30 ng/uL GV16, 0.73 uL of 16 ng/uL L1L2 backbone added to each 15 uL Gibson mix. Incubated in thermocycler at 50 C for 1 hour. (Used formula to obtain required volume for 7 fmol: 7 x 0.65/(conc ng/uL) x length / 1000.</p> | ||
| + | |||
| + | <p>Charles and Clara @ Knight lab: Gibsons transformed into 10G cells (0.9mL SOC), incubated spinning for 1 hour at 37C. Centrifuged for 3 min max speed (14000), removed 500uL of supernatant, resuspended the remaining & Plated 100 uL of each tube on KAN. Incubating plates 37C overnight (6:30pm ~ right after morning meeting tmr). The leftover tubes (Gibson + transformation) are in the iGEM fridge.</p> | ||
| + | |||
| + | |||
| + | <p>Divya and Tyler's SDM:</p> | ||
| + | |||
| + | <p>We used the Agilent QuikChange II SDM Kit and followed the protocol in its manual (<span class="nobr"><a href="http://www.genomics.agilent.com/files/Manual/200523.pdf).&nbsp" class="external-link" rel="nofollow">http://www.genomics.agilent.com/files/Manual/200523.pdf).&nbsp<sup><img class="rendericon" src="/confluence/images/icons/linkext7.gif" height="7" width="7" align="absmiddle" alt="" border="0"/></sup></a></span>;</p> | ||
| + | |||
| + | <p><b><ins>GV16 Mutagenesis</ins></b> (to remove BsaI site)</p> | ||
| + | <div class='table-wrap'> | ||
| + | <table class='confluenceTable'><tbody> | ||
| + | <tr> | ||
| + | <th class='confluenceTh'> Amount of dsDNA </th> | ||
| + | <th class='confluenceTh'> 10 ng [A] </th> | ||
| + | <th class='confluenceTh'> 20ng [B] </th> | ||
| + | <th class='confluenceTh'> 50ng [C] </th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> dsDNA template </td> | ||
| + | <td class='confluenceTd'> 1.89 uL </td> | ||
| + | <td class='confluenceTd'> 3.78 uL </td> | ||
| + | <td class='confluenceTd'> 9.45 uL </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> 10x Reaction Buffer <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 5 uL </td> | ||
| + | <td class='confluenceTd'> 5 uL </td> | ||
| + | <td class='confluenceTd'> 5 uL </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> GV16 Mutagenesis Primer (Forward) </td> | ||
| + | <td class='confluenceTd'> 1.25 uL </td> | ||
| + | <td class='confluenceTd'> 1.25 uL <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1.25 uL <br class="atl-forced-newline" /> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> GV16 Mutagenesis Primer (Reverse) </td> | ||
| + | <td class='confluenceTd'> 1.25 uL <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1.25 uL <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1.25 uL <br class="atl-forced-newline" /> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> dNTP mix </td> | ||
| + | <td class='confluenceTd'> 1 uL </td> | ||
| + | <td class='confluenceTd'> 1 uL <br class="atl-forced-newline" /> </td> | ||
| + | <td class='confluenceTd'> 1 uL <br class="atl-forced-newline" /> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> H2O (to get to 50 uL) </td> | ||
| + | <td class='confluenceTd'> 39.61 uL </td> | ||
| + | <td class='confluenceTd'> 37.72 uL </td> | ||
| + | <td class='confluenceTd'> 32.05 uL </td> | ||
| + | </tr> | ||
| + | </tbody></table> | ||
| + | </div> | ||
| + | |||
| + | <p>Then added 1uL HF DNA Polymerase</p> | ||
| + | |||
| + | <p><b><ins>TEV Protease Mutagenesis</ins></b> (to remove BsaI site)</p> | ||
| + | |||
| + | <div class='table-wrap'> | ||
| + | <table class='confluenceTable'><tbody> | ||
| + | <tr> | ||
| + | <th class='confluenceTh'> <font color="#660000">Amount of dsDNA</font> </th> | ||
| + | <th class='confluenceTh'> <font color="#660000"><del>10 ng [A]</del></font> </th> | ||
| + | <th class='confluenceTh'> <font color="#660000">20ng [B]</font> </th> | ||
| + | <th class='confluenceTh'> <font color="#660000">50ng [C]</font> </th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">dsDNA template</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>1.98 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">3.96 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">9.92 uL</font> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">10x Reaction Buffer</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>5 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">5 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">5 uL</font> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">TEV Protease Mutagenesis Primer (Forward)</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>1.25 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1.25 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1.25 uL</font> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">TEV Protease Mutagenesis Primer (Reverse)</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>1.25 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1.25 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1.25 uL</font> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">dNTP mix</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>1 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">1 uL</font> </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td class='confluenceTd'> <font color="#000000">H2O (to get to 50 uL)</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000"><del>39.52 uL</del></font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">37.54 uL</font> </td> | ||
| + | <td class='confluenceTd'> <font color="#000000">31.58 uL</font><br class="atl-forced-newline" /> </td> | ||
| + | </tr> | ||
| + | </tbody></table> | ||
| + | </div> | ||
| + | |||
| + | <p>Then added 1uL HF DNA Polymerase</p> | ||
| + | |||
| + | <p>I accidentally added too much Primer to the 10ng tube, so it was scrapped.</p> | ||
| + | |||
| + | <p>After going through the Thermal Cycler, 1uL of DpnI was added to each remaining sample. The samples were then put in the incubator at 37 degrees C for 1 hour, then moved to the freezer.</p> | ||
| + | |||
| + | <h3><a name="DailyNotebook-"></a><font color="#660000"><b>July 5, 2011</b></font></h3> | ||
| + | |||
| + | <p><font color="#000000">AVPR2 re-PRC: Charles and Clara re-PCR-ed the AVPR2 using primers 13 and 14. This time, however, we used 20s extension time instead of 18s. </font></p> | ||
| + | |||
| + | <p><font color="#000000">Tyler: Did LR at 4:30 pm for pEXPR 4-5 UAS:EBFP2. Will transform tomorrow morning. Plan on also getting LacI (without the Krab) from Weiss lab for use as a weak repressor.</font></p> | ||
| + | |||
| + | </span> | ||
| + | |||
<!-----------------------------------------------------------------------------------------------------------------> | <!-----------------------------------------------------------------------------------------------------------------> | ||
</!> | </!> | ||
Revision as of 21:09, 28 September 2011
Overview
Here you can see what we did weekly in the lab!
Week 1: June 6 - June 11
In the first week we began constructing basic reporter DNA constructs that have various combinations of promoters and genes. The mix-and-matching of promoters and genes was done using the LR reaction following the Gateway protocol. Our promoters included TRE, minCMV, Hef1a, Hefl1a-LacO, and our genes included eYFP, mKate, and eBFP2. Transformed and inoculated colonies from the LR reactions and performed restriction digests. Approximately half of the LR reactions were successful and became DNA parts that we could use later on.June 7, 2011
| Date |
Assignee |
DEST_R4R2 |
L4-R1 Promoter |
L1-L2 Gene |
LR? |
Tube Number |
Colony Number, Antibiotic, Person |
Protocol Used |
|---|---|---|---|---|---|---|---|---|
| 6.7.2011 |
Michelle Louis |
2-3 |
Tre |
C1434VP16 |
Yes. 6.7.2011, 5pm |
11 |
5, AMP, KH |
Standard, 30C for 16h |
| 6.7.2011 |
Michelle Louis |
2-3 |
Tre |
Mnt1VP16 |
Yes. 6.7.2011, 5pm |
12 |
4, AMP, KH | Standard, 30C for 16h |
| 6.7.2011 |
Tyler |
2-3 |
Tre |
LacIKrab |
Yes. 6.7.2011, 5pm |
6 |
30, AMP, KH | Standard, 30C for 16h |
| 6.7.2011 |
Jenny Divya |
3-4 |
minCMV-7xMnt1 |
eYFP |
Yes. 6.7.2011, 5pm |
9 |
0, AMP, KH, needs to be redone |
Standard, 30C for 16h |
| 6.7.2011 |
Jenny Divya |
2-3 |
Tre |
LexAVP16 |
Yes. 6.7.2011, 5pm |
10 |
2, AMP, KH, needs to be redone |
Standard, 30C for 16h |
| 6.7.2011 | Mariola Kenneth |
3-4 |
minCMV 1xC1434 |
eYFP |
Yes. 6.7.2011, 5pm |
7 |
3, AMP, KH, needs to be redone | Standard, 30C for 16h |
| 6.7.2011 | Mariola Kenneth |
3-4 | minCMV 4xLexA |
eYFP |
Yes. 6.7.2011, 5pm |
8 |
0, AMP, KH, needs to be redone |
Standard, 30C for 16h |
LR Reactions
Utilizing protocol here: LR Protocol
| Date |
Assignee |
DEST_R4R2 |
L4-R1 Promoter |
L1-L2 Gene |
LR? |
Tube Number |
Colony Number, Antibiotic, Person |
Protocol Used |
|---|---|---|---|---|---|---|---|---|
| 6.9.2011 |
Grant |
3-4 |
minCMV 4xLexA | eYFP | Yes. 6.9.2011, 3pm |
12 |
10, AMP, GR |
Standard + Grown at 37C in LB not SOC. |
| 6.9.2011 |
Jon C |
3-4 |
minCMV-4xMnt1 |
eYFP |
Yes. 6.9.2011, 3:45pm | 2 |
13, AMP, GR |
Standard + Grown at 37C in LB not SOC. |
| 6.9.2011 |
Jon C |
3-4 |
minCMV-4xCI434 |
eYFP |
Yes. 6.9.2011, 3:45pm | 1 |
7, AMP, GR | Standard + Grown at 37C in LB not SOC. |
| 6.9.2011 |
Jon C |
3-4 |
Hef1a-LacO |
eYFP |
Yes, 6.9.2011, 4:20pm |
5 |
25, AMP, GR |
Standard + Grown at 37C in LB not SOC. |
| 6.9.2011 |
Semon |
4-5 |
Tre |
mKate |
Yes, 6.9.2011 4:27pm |
3 |
6, AMP, GR |
Standard + Grown at 37C in LB not SOC. |
| 6.9.2011 |
Semon |
1-2 |
Hef1a |
eBFP2 |
Yes, 6.9.2011, 4.27pm |
4 |
3, AMP, GR |
Standard + Grown at 37C in LB not SOC. |
Transformation
| Date |
Assignee |
DEST_R4R2 |
L4-R1 Promoter |
L1-L2 Gene |
LR? |
Colony Number, Antibiotic |
Protocol Used | |
|---|---|---|---|---|---|---|---|---|
| 6.9.2011 |
Mariola |
|
|
eYFP |
no. Replication stock. | |
2, AMP |
Transformation (~ 2 hr 20 min) using LB instead of SOC |
Miniprep
| Date | Assignee |
DNA |
Quantity |
Time collected |
|---|---|---|---|---|
| 6.9.2011 |
Divya-Jenny |
pEXPR_2-3_Tre:LexAVP16 |
52.2 ng/uL |
12pm |
| 6.9.2011 |
Kenneth |
pEXPR_3-4_minCMV-CI434:eYFP (A) |
70 ng/ul |
12pm |
| 6.9.2011 |
Kenneth | pEXPR_3-4_minCMV-CI434:eYFP (B) |
174 ng/uL |
12pm |
| 6.9.2011 |
Louis |
pEXPR_2-3_Tre:C1434VP16 |
234.6 ng/uL |
12pm |
| 6.9.2011 |
Tyler |
pEXPR_2-3_Tre:Lac/Krab |
130.7 ng/uL |
12pm |
| 6.9.2011 |
Michelle |
pEXPR_2-3_Tre:Mnt1VP16 |
110.0 ng/uL |
12pm |
| 6.9.2011 |
Michelle |
pDEST_2-3_ccdB |
117.6 ng/uL |
12pm |
Restriction Digests
| Assignee |
DNA |
Enzyme |
Expected Results |
Picture of Gel |
Time Incubated | Comments |
||
|---|---|---|---|---|---|---|---|---|
| Louis |
pEXPR_2-3_Tre:C1434VP16 | NdeI |
6700 bp 800 bp |
a.3 | 6/9/11 3:50 PM |
|
||
| Kenneth |
pEXPR_3-4_minCMV-CI434:eYFP (A+B) |
NcoI and SacII |
2450 bp 4650 bp |
A: a.6 B: a.7 |
6/9/11 3:00 PM |
A did not cut as expected. B looks good. Digestion Mix: 2 uL NEB4, 0.5 uL of each enzyme, A: 7.1 uL/B: 2.9 uL of DNA, 10.4 uL/14.6 uL of H20 |
||
| Michelle |
pEXPR_2-3_Tre:Mnt1VP16 |
BglI |
3700 bp 2200 bp 1300 bp |
a.1 | 6/9/11 4:00PM |
DNA appeared to be uncut. Need to redo, possibly with a new enzyme and more DNA. |
||
| Michelle |
pDEST_2-3_ccdB |
NcoI and NheI |
6100 bp 1700 bp |
a.2 | 6/9/2011 4:25 PM |
Attained bands at correct positions. Other band represents partially cut DNA in double digest. |
||
| Divya |
pEXPR_2-3_Tre:LexAVP16 |
HincII |
a.4 | 6/9/2011 4:45 PM |
- Bands didn't show up. Needs to be redone. - DNA may have degraded. Need to redo Nanodrop as well. |
|||
| Tyler | pEXPR_2-3_Tre:Lac/Krab | SalI | 3 bands: 5288 bp 1510 bp 1241 bp cuts gene |
a.5 | 6/9/2011 3:00 PM |
Digestion: 4 uL * 130.7 ng/uL = 522.8 ng DNA 2 uL NEB3 2 uL BSA 11 uL H2O 1 uL SalI Total: 20 uL Only saw one band at 3500 bp Needs to be redone possibly with another restriction enzyme |
Gels
June 10, 2011
Restriction Gels
| Assignee | DNA |
Enzyme |
Expected result |
Time Incubated |
Comments |
|---|---|---|---|---|---|
| Louis |
pEXPR_2-3_Tre:C1434VP16 | EcoRI (Buffer 2) |
5500 bp 1050 bp 650 bp 360 bp |
6/10/11 11:15 AM |
|
| Michelle |
pEXPR_2-3_TRE:Mnt1VP16 (A) |
NcoI and NheI |
5700 bp 1300 bp |
6/10/11 11:40 AM |
Digestion: 6 uL DNA 1 uL NcoI 1 uL Nhe1 2 uL NEB2 2 uL BSA 8 uL H2O Failed. |
| Michelle |
pEXPR_2-3_TRE:Mnt1VP16 (B) |
NheI and SphI |
1700 bp 5300 bp |
6/10/11 11:40 AM |
Digestion: 6 uL DNA 1 uL NheI 1 uL SphI 2 uL NEB2 2 uL BSA 8 uL H2O Failed. Redoing LR on Monday June 13th. |
| Tyler | pEXPR_2-3_Tre:Lac/Krab | SalI | 3 bands: 5288 bp 1510 bp 1241 bp cuts gene |
6/10/2011 3:00 PM |
Digestion: 4 uL * 130.7 ng/uL = 522.8 ng DNA 2 uL NEB3 2 uL BSA 11 uL H2O 1 uL SalI Total: 20 uL ***Worked (see gel below - b.1) Stored in -80C freezer |
Comments:
pEXPR_2-3_Tre:LexAVP16 and pEXPR_3-4_minCMV-7xMnt1:eYFP are being entirely redone.
pEXPR_2-3_Tre:LexAVP16 (post-miniprep) was lost. pEXPR_3-4_minCMV-7xMnt1:eYFP LR was unsuccessful (no bacteria grew).
LR and transformation will be done on Saturday. Miniprep and restriction mapping will be done on Sunday
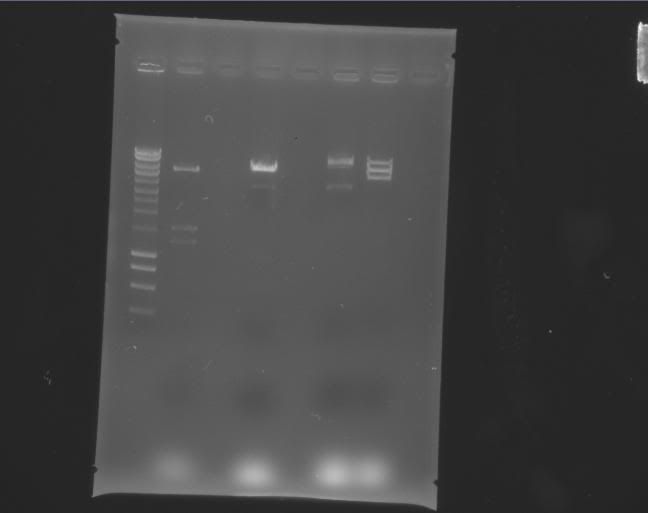
| Label | Gel |
Legend |
|---|---|---|
| b |
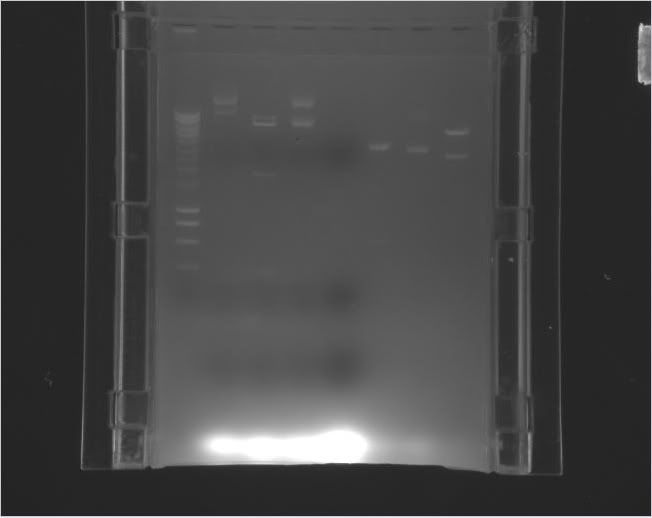 |
Column 0: HyperLadder Column 1: pEXPR_2-3_Tre:Lac/Krab Column 4: pEXPR_2-3_Tre:C1434VP16 Column 6: pEXPR_2-3_TRE:Mnt1VP16 (A) Column 7: pEXPR_2-3_TRE:Mnt1VP16 (B) |
June 11, 2011
Set of LRs from 6/9 (GR, JC, and SR) cell counts taken; inoculated by GR/JC. 3 cells taken per plate.
| Assignee |
DNA |
Enzyme |
Expected Results |
Time Incubated |
Comments |
|---|---|---|---|---|---|
| Sam |
pEXPR_1-2_Hef:eBFP |
NcoI & ApaL1 |
3750bp, 3180bp, 1250bp | 45 min | If it doesn't work there should be a whole mess of fragments (including a small 600bp one) 1mL each enz; 2 mL Buf; 1 mL BSA; 2mL DNA; 13mL H2O (Standard Protocol with extra BSA) |
| Sam |
pEXPR_4-5_Tre:mKate |
Bgl1 |
1270bp, 2630bp, 3380bp | 45 min | 1mL enz; 2mL DNA; 2mL BSA; 2mL Buf; 15mL H2O (Standard Protocol with extra BSA and water - not quite 10x) |
Week 2: June 12 - June 18
In the second week we attempted midipreps instead of minipreps of our available cell stocks. Midipreps were unsuccessful, likely due to centrifuge limitations. LR reactions to generate more usable promoter-gene pair parts continued, expanding to include the CI434-VP16, LexA-VP16, and Mnt-VP16 genes, whose expressed proteins activate the appropriate corresponding minCMV promoters. Most of the LRs done were not successful, reason unknown at this time.June 13, 2011
Status
| DNA | Status |
Comment |
|---|---|---|
| Mntcmv/1x cI434:eYFP |
good |
|
| Mnt cmv/4x LexA:eYFP |
bad | |
| HefIa/LacO:eYFP |
good | |
| Tre:LacI Krab |
good | |
| HefIaL EBFP2 |
bad |
| Date |
Assignee |
DEST_R4R2 |
L4-R1 Promoter |
L1-L2 Gene |
LR? |
Tube Number |
Colony Number, Antibiotic, Person |
Protocol Used |
|---|---|---|---|---|---|---|---|---|
| 6.13.2011 |
Jon |
2-3 |
Tre | C1434-VP16 | Yes. 6-13, 11am |
10 |
about 50, AMP, JC |
No |
| 6.13.2011 |
Tyler |
2-3 |
Tre |
LexA-VP16 |
Yes. 6-13, 11am | 9 |
about 30, AMP, GR |
No |
| 6.13.2011 |
Mariola |
2-3 |
Tre |
Mnt1-VP16 |
Yes. 6-13, 11am | Eppendorf |
0, AMP, GR | No |
| 6.13.2011 |
Semon |
4-5 |
Tre |
mKate |
Yes. 6-13, 11am |
11 |
about 10, AMP, SR |
No |
| 6.13.2011 |
Divya |
3-4 |
minCMV-7xMnt1 |
eYFP |
Yes 6/13, 1:30pm |
? |
1000s, AMP, GR |
No |
| 6.13.2011 |
Jenny |
3-4 |
minCMV-4xMnt1 |
eYFP |
Yes 6/13, 1:30pm |
? |
1000s, AMP, GR |
No |
| 6.13.2011 |
Jenny |
3-4 |
minCMV-4xC1434 |
eYFP |
Yes 6/13,1:30pm |
? |
1000s, AMP, GR |
No |
June 14, 2011
Highlights:
~10:30 AM: Miniprep of pDEST23-ccdB commenced and completed. DNA concentrations of the two samples were 175 and 310 ng/uL
~1:00 PM: Inoculation of cultures for pEXPR-23-TRE-CI434-VP16, pEXPR-23-TRE-LexA-VP16, pEXPR-45-TRE-mKate, pEXPR-34-minCMV-7xMnt-eYFP, pEXPR-34-minCMV-4xMnt-eYFP, and pEXPR-minCMV-4xCI434 were conducted. Colony counts from the previous transformations are noted above.
~1:30 PM: LR Gateway reactions were performed for the failed pEXPR-23-TRE-Mnt-VP16 and the new pEXPR-12-Hef1A-rtTA.
~5:00 PM: Inoculation of midiprep cultures for EXPR-34-minCMV4xLexA-eYFP, two samples of pEXPR-34-Hef1ALacO-eYFP, pEXPR-12-Hef1A-EBFP, pEXPR-23-TRE-LacIKrab, and pEXPR-34-minCMV1xcI434-eYFP commenced. We plan to midiprep the cells at about 10:00 AM.
~6:00 PM: Transformation of the aforementioned LR Gateway reactions commenced. Transformation completed around 9:00 PM.
~7:00 PM: Design for the TANGO primers complete.
June 15, 2011
~4AM: Grant did more LRs – 3-4_minCMV-4xLexA:eYFP, 1-2_Hef1a:rttA3, 1-2_Hef1a:eBFP2
~10AM: Minipreps done on the 6 LRs that succeeded from yesterday by Grant
Nanodrops
| Date |
Assignee |
Vector |
Concentrations (ng/uL) *denotes 260/280 below 1.8 |
|---|---|---|---|
| 6.14 11AM |
Jon/Michelle/Clara |
2-3_Tre:C1434-VP16 |
A: 284.0* (1.78), B: 391.3, C: 380.9, D: 205.5 |
| 6.14 11AM |
Jon/Michelle/Clara |
2-3_Tre:LexA-VP16 |
A: 361.4, B: 193.9* (1.64), C: 212.9* (1.58), D: 390.6 |
| 6.14 11AM |
Jon/Michelle/Clara |
4-5_Tre:mKate |
A: 176.9, B: 180.3, C: 188.7, D: 208.8 |
| 6.14 11AM |
Jon/Michelle/Clara |
3-4_minCMV-4xC1434:eYFP |
A: 173.8, B: 152.5* (1.60), C: 319.5, D: 192.5 |
| 6.14 11AM |
Jon/Michelle/Clara |
3-4_minCMV-7xMnt1:eYFP |
A: 147.7, B: 251.5, C: 147.6, D: 178.2 |
| 6.14 11AM |
Jon/Michelle/Clara |
3-4_minCMV-4xMnt1:eYFP |
A: 195.1, B: 241.2* (1.75), C: 193.7, D: 157.0 |
Restriction Maps, 6.14 5PM
| Assignee |
DNA |
Enzyme |
Expected Results |
Comments |
|---|---|---|---|---|
| Jon |
3-4_minCMV-4xMnt1:eYFP |
SalI-HF |
5288bp, 2044bp if works; 5288bp, 1600bp, 977bp if doesn't | All: 1uL SalI, 2uL Buf 4, 2uL BSA; A,B,C: 3uL DNA, D: 4uL DNA; DI up to 20 uL |
| Jon |
3-4_minCMV-7xMnt1:eYFP |
SalI-HF |
5288bp, 2257bp if works; 5288bp, 1600bp, 977bp if doesn't | All: 1uL SalI, 2uL Buf 4, 2uL BSA; A,B,C: 3uL DNA, D: 4uL DNA; DI up to 20 uL |
| Jon |
4-5_Tre:mKate |
SalI-HF |
5288bp, 1995bp if works; 5288bp, 1600bp, 977bp if doesn't | All: 1uL SalI, 2uL Buf 4, 2uL BSA; A,B,C: 3uL DNA, D: 4uL DNA; DI up to 20 uL |
| Mariola |
2-3_Tre:C1434-VP16 |
SalI-HF |
5288, 1862, 412 bp if works; 5288bp, 1600bp, 977bp if doesn't | All: 1uL SalI, 2uL Buf 4, 2uL BSA; A: 1.8uL DNA, B,C: 1.3uL DNA, D: 2.4uL DNA; DI up to 20 uL |
| Mariola |
3-4_minCMV-4xC1434:eYFP |
BamHI |
3757, 1855, 1166, 478 bp if works; no cut if doesn't | All: 1uL BamHI, 2uL Buf 4, 2uL BSA; A 2.9uL DNA, B: 3.2uL DNA, C: 1.6uL DNA, D: 2.6uL DNA; DI up to 20 uL |
| Tyler |
2-3_Tre:LexA-VP16 |
ScaI |
2 cuts: 4464 and 2729 bp if works; 3 cuts: 3535, 2729, 1589 if doesn't | All: 1uL SalI, 2uL Buf 4, 2uL BSA; A: 3uL DNA, B,D: 1.5uL DNA, C: 2.5uL DNA; DI up to 20 uL From Gel Below: A and C show the entry vector, B failed as well, D seems to have worked. There is uncut DNA around 7000 bp, but the other bands seem correct. D will be sent for sequencing. |

Midipreps were done by Kenneth and Tyler for sequenced verified constructs, 2-3_Tre:LacI-Krab, 3-4_minCMV 1xCI434:eYFP, and 3-4_Hef1ALac0:eYFP. Unfortunately DNA yields were low, as shown in the table below, so minipreps will be performed tomorrow, June 16, 2011.
| Date | Assignee | Vector | DNA Concentration (ng/uL) |
|---|---|---|---|
| 6.15.2011 | Tyler | 2-3_Tre:LacI-Krab | 54 |
| 6.15.2011 | Kenneth | 3-4_minCMV 1xCI434:eYFP | 216 |
| 6.15.2011 | Kenneth/Tyler | 3-4_Hef1ALac0:eYFP | failed |
June 16, 2011
~10am Minipreps
| Date | Assignee | Vector | DNA Concentration (ng/uL) |
|---|---|---|---|
| 6.16.2011 | Tyler | 2-3_Tre:LacI-Krab | 509.1 |
| 6.16.2011 | Kenneth | 3-4_minCMV 1xCI434:eYFP | 89.1 |
| 6.16.2011 | Kenneth/Tyler | 3-4_Hef1ALac0:eYFP | 385 |
June 17-18, 2011 Construction
Protocol for all Cells: Added DNA 5 minutes after removing cells from freezer. Incubated for 30 minutes. Heat shocked for 30 seconds at 42 Celsius. Incubated on ice for two minutes. Added 1 mL of SOC. Grew at 30 C for 2 hours. Plated 100 uL from total reaction on one half of plate & the rest of the reaction on the other half of plate.
| Date |
Assignee |
DEST_R4R2 |
Colonies (low/high conc.) |
|---|---|---|---|
| 6.18.2011 |
Grant | pEXPR12-Hef1A-eBFP2 | Order of <10 |
| 6.18.2011 |
Grant | pEXPR12-Hef1A-rtTA3 | Order of <10 |
| 6.18.2011 |
Grant | pEXPR23-TRE-CI434.VP16 | Order of 100-1000 |
| 6.18.2011 |
Grant | pEXPR23-TRE-LexA.VP16 | Order of 100-1000 |
| 6.18.2011 |
Grant | pEXPR23-TRE-Mnt.VP16 | Order of 100-1000 |
| 6.18.2011 | Grant |
pEXPR34-minCMV.4xCI434-EYFP | Order of 1000-10000 |
| 6.18.2011 | Grant | pEXPR34-minCMV.4xLexA-EYFP | Order of 1000-10000 |
| 6.18.2011 | Grant | pEXPR34-minCMV.4xMnt-EYFP | Order of 1000-10000 |
| 6.18.2011 | Grant | pEXPR34-minCMV.7xMnt-EYFP | Order of 1000-10000 |
| 6.18.2011 | Grant | pEXPR45-TRE-mKate | Order of <10 |
Conclusion: pDEST12 appears to be growing slowly in all cases, as does pDEST45. pDEST23 grows at the rate typically expected by Gateway reaction standards while pDEST34 grows at a rate much faster than these standards. Corroborated in Weiss lab?
Week 3: June 20 - June 24
In the third week we continued expanding our library of usable DNA parts by creating more combinations of genes and promoters using the LR reaction.June 20, 2011
Minipreps and nanodrops done on the plasmids from yesterday by Clara and Grant.
Restriction digesting done on the 10 transformations.
| Assignee |
DNA |
Enzyme |
Expected Results |
Picture of Gel |
Time Incubated |
Comments |
|---|---|---|---|---|---|---|
| Jon |
1-2_Hef1a-rtTA3 | SalI-HF |
5288, 2877bp if works; 5288, 1600, 977bp if it doesn't |
Gel 1 Lanes 2-4 |
2 hrs |
Being redone |
| Jon |
1-2_Hef1a-eBFP2 | SalI-HF |
5288, 2889bp if works; 5288, 1600, 977bp if it doesn't | Gel 1 Lanes 5-7 |
2 hrs | Being redone |
| Jon |
3-4_minCMV-7xMnt1:eYFP | SalI-HF |
5288bp, 2257bp if works; 5288bp, 1600bp, 977bp if doesn't | Gel 1 Lanes 8-10 |
2 hrs | A and C sent off for sequencing |
| Jon |
2-3_Tre_Mnt1-VP16 |
ScaI |
4440, 2729bp if works; 3535, 2729, 1589bp if doesn't |
Gel 1 Lane 11, Gel 2 Lanes 2-3 |
2 hrs | A sent for sequencing; also being redone |
| Michelle |
4-5_Tre_mKate |
SalI-HF | 5288bp, 1995bp if works; 5288bp, 1600bp, 977bp if doesn't | Gel 2 Lanes 10-11, Gel 3 Lane 2 |
2 hrs | B sent off for sequencing; also being redone |
| Michelle | 2-3_Tre:CI434-VP16 |
SalI-HF | 5288, 2270bp if works; 5288, 1600, 977bp if doesn't |
Gel 2 Lanes 7-9 |
2 hrs | Being redone |
| Michelle | 2-3_Tre:LexA-VP16 | SalI-HF | 5288, 1913bp if works; 5288, 1600, 977bp if doesn't | Gel 2 Lanes 4-6 |
2 hrs | B sent off for sequencing |
| Charles |
3-4_minCMV 4xLexA:eYFP | SalI-HF | 5288, 1849bp if works; 5288bp, 1600bp, 977bp if doesn't | Gel 3 Lanes 6-8 |
2 hrs | B sent off for sequencing; also being redone |
| Charles |
3-4_minCMV 4xMntI:eYFP | SalI-HF | 5288bp, 2044bp if works; 5288bp, 1600bp, 977bp if doesn't | Gel 3 Lanes 3-5 |
2 hrs | Being redone |
| Charles |
3-4_minCMV 4xCI434:eYFP | SalI-HF | 5288, 1960bp if works; 5288bp, 1600bp, 977bp if doesn't | Gel 3 Lanes 9-11 |
2 hrs | B sent off for sequencing |

Key: Ladder, 3x Hef1a rtTa3, 3x Hef1a eBFP2, 3x minCMV 7xMnt1:eYFP, Tre Mnt1 VP16 A, Ladder; Tre Mnt1 VP16 B+C, 3x Tre LexA VP16, 3x Tre CI434 VP16, Tre mKate A+B, Ladder

Key: Ladder, Tre mKate C, 3x minCMV 4xMnt1, 3x minCMV 4xLexA, 3x minCMV 4xCI434, Ladder
June 23, 2011
Colony Counts
These results are based on plating 100 uL of the outgrowth culture.
| Part Shorthand |
Colonies this Time |
Average Colonies Before |
Notes |
|---|---|---|---|
| TRE:mKate |
~10 |
~3 |
Using new TRE, Dest 23. |
| TRE:MntVP16 |
~50 |
~10 |
Using new TRE, Dest 23. |
| TRE:cI434VP16 |
~30 |
~1 |
Using new TRE, Dest 23. |
| Hef1A:rtTA3 |
~50 |
~1 |
Using new Hef1A, Dest 23. |
| Hef1A: eBFP2 |
~30 |
~3 |
Using new Hef1A, Dest 23. |
| 4xLexA:eYFP |
~1000s |
~1000s |
Using Dest 23. |
| 4xMnt:eYFP |
~1000s |
~1000s |
Using Dest 23. |
Conclusion: pENTR vectors probably weren't properly aliquoted the first time around, strongly reducing recombination efficiency.
Week 4: June 27 - July 1
In the fourth week we implemented a color-coded box system in our -20 freezer to accommodate for the growing need of organization of the growing number of DNA parts. We began learning and using the Gibson reaction to create the gene part AVPR2-TEVs-GV16, which is expressed to produce a vasopressin receptor bound to Gal4-VP16 by a TEV sequence that can be recognized by the TEV Protease. Gibson reaction results were not great, and it appeared that our AVPR2 DNA was not of sufficient quality, so we re-PCRed the AVPR2 DNA segment.June 28, 2011

June 29, 2011
~ Miniprepped at 10:30 AM
| Date |
Assignee | Vector |
DNA Concentration |
|---|---|---|---|
| 6.29.2011 |
Mariola |
pENTR_UAS |
A: 98.7 ng/ul B: 88.1 ng/ul |
| 6.29.2011 |
Mariola |
pENTR_Tet-LacO |
A: 84.7 ng/ul B: 83.2 ng/ul |
| 6.29.2011 |
Jenny |
pENTR_TEV-Protease |
A: 97.5 ng/ul B: 50.4 ng/ul |
| 6.29.2011 |
Jenny |
pENTR_LacI-miRRF4 |
A: 58.2 ng/ul B: 161.4 ng/ul |
| 6.29.2011 |
Michelle |
pENTR_GV16 |
A: 49.9 ng/ul B: 52.9 ng/ul |
June 30, 2011
11:45AM ~ 12:45PM (Charles, Clara)
Miniprep AVPR2 for TANGO: 270.9 ng/ul (successful)
3:00PM ~ 5:00 PM (Charles, Clara)
PCR primer # 13, 14, 17, 18, 19, 20
- AVPR2 - TEVs
Annealing temperature: 58.9 C (13 - 58.6 C; 14 - 55.9 C)
Extension time: 18 seconds (~1200 bp)
- Gal4 - VP16
Ta: 62.4 C (17 - 62.4 C; 18 - 59.4 C)
Ext: 13 seconds (~740 bp)
- L1L2 Backbone
Ta: 65.4 C (19 - 62.4 C; 62.4 C)
Ext: 30 seconds (~2500 bp)
5:00PM ~ 5:30PM (Charles, Clara)
Gibson assembly cont'd: inoculated and incubating samples overnight
July 1, 2011
| Time | Notes |
|---|---|
| 8:00 AM | Color-Coded inventory implemented in the -20 freezer. To be implemented in the -80 freezer. See below for details. |
| 9:00 AM | Morning meeting. |
| 11:00 AM | Design meeting. Significant amount of information to be added to wiki in terms of construction plans and proposed circuits. Do ASAP. |
| 1:30 PM | LR reactions of pDEST_45, pENTR EYFP-FF6, and the five inducible minCMV promoters complete. |
| 2:00 PM | Kenneth's next PCR of Delta started. |
| 2:00 PM | Cell stocks of the working pEXPR transactivators, TRE:mKate, and Hef1a:EBFP2 made. |
| 2:00 PM | Inoculation of the aforementioned pEXPR vectors for later transfections completed. |
1:00 PM ~ AVPR2 - TEVs - Gal4 VP16 cont'd:
Clara did PCR purification and nanodrop to check the concentrations.
The concentrations were relatively low:
AVPR2: 20.8 ng/uL
Gal4-VP16: 30ng/uL
L1L2 Gibson: 16 ng/uL
Used 10uL of each sample to run the gel.
The following are the pictures of the gel (Same picture but different exposure levels), in order of AVPR2; Gal4-VP16; and L1L2 Gibson:
Expected Length - AVPR2: ~1200 bp; Gal4-VP16: ~740 bp; L1L2 Gibson: ~2500 bp


Gal4-VP16 and L1L2 Gibson seem to be successful. Should PCR AVPR2 again.
As for the Gibson assembly practice, the practice samples were over-incubated so I had to throw them away. (Sorry, Charles!)
Color-Coded Inventory implemented in the -20 freezer.
Purple: Destination Vectors
Red: Gene Vectors
Green: Promoter Vectors
Yellow: Source/Ordered Vectors
Orange: Expression Vectors
Blue: Primers (5 uM)
Pink: Primer Stocks (100 uM)
Not yet ported to the -80 freezer cell stocks.
Week 5: July 4 - July 10
We began looking into using the Goldengate assembly method, but two gene elements contained a cut site that needed to be mutated out. We performed Site-Directed Mutagenesis using the Lightning Kit in the hopes of mutating out the cut site, but results were not successful due to mishandling during the protocol, so the procedure was set to be repeated. Gibson assembly of AVPR2-TEVs-GV16 continues as we re-PCRed the AVPR2 DNA segment and re-run the entire Gibson protocol, picking 20 colonies in a determined attempt to obtain a successful result.July 10
Jenny and Jon - SDM Colony counts were approximately 50 for the control, 20 for the TEV, and 0 for the Gal4VP16. There is a chance the TEV and Gal4VP16 labels were switched, which we'll find out after sequencing. Six colonies were picked from the TEV plate and inoculations were accidentally run for 22 hours at 37C.
July 9
Jenny and Jon redid the SDM for Control, Gal4VP16, and TEV using the Lightning Kit, following pages 7 and 8 of this manual: www.qcbio.com/stratagene/210518.pdf
We used 1.5uL 100mM stock of the previously designed primers. The final reaction volume was only approximately 45uL, as we only had about 5uL of 5ng/uL dsDNA for both the Gal4VP16 and TEV reactions. We got the DNA from the dilutions that Divya and Tyler previously used.
For transformation, we followed the standard protocol instead of the one outlined in the manual and outgrew at 37C and incubated at 30C for 16 hrs.
July 8, 2011
| Sample | Assignees |
Procedures |
Enzyme(s) Used |
Expected Bands |
Results |
|---|---|---|---|---|---|
| EXPR 1xCI434 (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR 4xCI434 (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR 4xLexA (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR 4xMnt1 (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR 7xMnt1 (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR UAS:EBFP2 (A-C) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
ScaI |
5000 2000 |
|
| EXPR AVPR2 A (1-3) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
NcoI |
1356 3000 |
|
| EXPR AVPR2 B (1-3) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
NcoI |
1356 3000 |
|
| EXPR AVPR2 C (1-3) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
NcoI |
1356 3000 |
|
| EXPR AVPR2 D (1-3) |
Grant/Michelle/Charles |
Miniprepped Nanodropped Restriction Digest |
NcoI |
1356 3000 |
July 7, 2011
Charles and Clara @ Knight lab: All four Kan plates had colonies, 40-50 colonies except for one of Deepak's gibson mix reactions, which had ~400 colonies. Inoculated 5 colonies from each reaction into LB-KAN. 20 Falcon tubes in 37C spinning. To be taken out at 2 AM by Grant.
July 6, 2011
Charles and Clara @ Weiss lab: Ran Gibson reaction with both DM and CH Gibson mixes each, total of 4 reactions run. 0.21 uL of 25.6 ng/uL AVPR2, 0.11 uL of 30 ng/uL GV16, 0.73 uL of 16 ng/uL L1L2 backbone added to each 15 uL Gibson mix. Incubated in thermocycler at 50 C for 1 hour. (Used formula to obtain required volume for 7 fmol: 7 x 0.65/(conc ng/uL) x length / 1000.
Charles and Clara @ Knight lab: Gibsons transformed into 10G cells (0.9mL SOC), incubated spinning for 1 hour at 37C. Centrifuged for 3 min max speed (14000), removed 500uL of supernatant, resuspended the remaining & Plated 100 uL of each tube on KAN. Incubating plates 37C overnight (6:30pm ~ right after morning meeting tmr). The leftover tubes (Gibson + transformation) are in the iGEM fridge.
Divya and Tyler's SDM:
We used the Agilent QuikChange II SDM Kit and followed the protocol in its manual (http://www.genomics.agilent.com/files/Manual/200523.pdf). ![]() ;
;
GV16 Mutagenesis (to remove BsaI site)
| Amount of dsDNA | 10 ng [A] | 20ng [B] | 50ng [C] |
|---|---|---|---|
| dsDNA template | 1.89 uL | 3.78 uL | 9.45 uL |
| 10x Reaction Buffer |
5 uL | 5 uL | 5 uL |
| GV16 Mutagenesis Primer (Forward) | 1.25 uL | 1.25 uL |
1.25 uL |
| GV16 Mutagenesis Primer (Reverse) | 1.25 uL |
1.25 uL |
1.25 uL |
| dNTP mix | 1 uL | 1 uL |
1 uL |
| H2O (to get to 50 uL) | 39.61 uL | 37.72 uL | 32.05 uL |
Then added 1uL HF DNA Polymerase
TEV Protease Mutagenesis (to remove BsaI site)
| Amount of dsDNA | |
20ng [B] | 50ng [C] |
|---|---|---|---|
| dsDNA template | |
3.96 uL | 9.92 uL |
| 10x Reaction Buffer | |
5 uL | 5 uL |
| TEV Protease Mutagenesis Primer (Forward) | |
1.25 uL | 1.25 uL |
| TEV Protease Mutagenesis Primer (Reverse) | |
1.25 uL | 1.25 uL |
| dNTP mix | |
1 uL | 1 uL |
| H2O (to get to 50 uL) | |
37.54 uL | 31.58 uL |
Then added 1uL HF DNA Polymerase
I accidentally added too much Primer to the 10ng tube, so it was scrapped.
After going through the Thermal Cycler, 1uL of DpnI was added to each remaining sample. The samples were then put in the incubator at 37 degrees C for 1 hour, then moved to the freezer.
July 5, 2011
AVPR2 re-PRC: Charles and Clara re-PCR-ed the AVPR2 using primers 13 and 14. This time, however, we used 20s extension time instead of 18s.
Tyler: Did LR at 4:30 pm for pEXPR 4-5 UAS:EBFP2. Will transform tomorrow morning. Plan on also getting LacI (without the Krab) from Weiss lab for use as a weak repressor.
<> "
"