Team:Wageningen UR/Project/CompleteProject1Description
From 2011.igem.org
(→Synchroscillator) |
(→Synchroscillator) |
||
| Line 49: | Line 49: | ||
'''Basic Components:''' | '''Basic Components:''' | ||
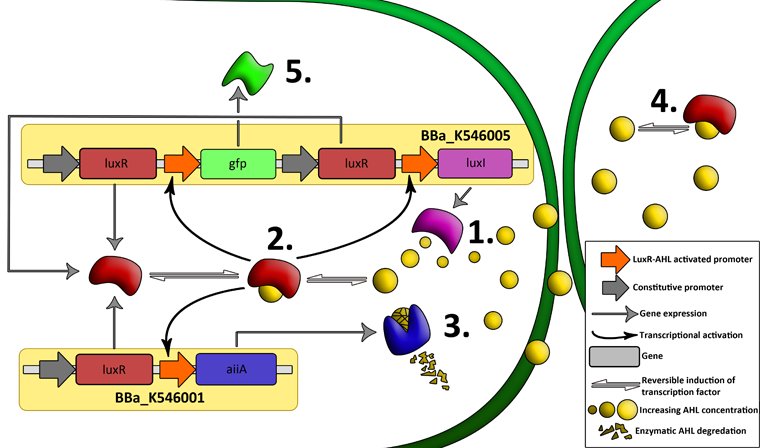
| - | '''LuxR''' is a transcriptional regulator in the bioluminescent quorum-sensing system of the symbiotic deep sea bacterium ''Vibrio fischeri''. It is induced by binding the | + | '''LuxR''' is a transcriptional regulator in the bioluminescent quorum-sensing system of the symbiotic deep sea bacterium ''Vibrio fischeri''. It is induced by binding the auto-inducer molecule N-(3-oxohexanoyl)-homoserine lactone (AHL). The AHL-LuxR complex controls expression of the lux regulon, which contains diverging pRight and pLeft promoter elements. The pRight element has low basal transcription, and is activated by AHL-LuxR; pLeft has higher basal expression, and is repressed by the AHL-LuxR complex. This dual activity makes LuxR a useful element for controlling interconnected genetic feedback loops. The unrestricted diffusion of AHL through the plasma membrane allows spatially proximate populations of cells to experience identical AHL conditions and synchronize AHL-dependent gene expression. |
The enzyme '''LuxI''' is an acyl-homoserine-lactone synthase which produces the intercellular signalling molecule N-(3-oxohexanoyl)-homoserine lactone (AHL). Placing LuxI under control of the pRight promoter results in a positive feedback loop: when increases in cell density cause the intracellular AHL concentration to rise above the activation threshold of the pRight promoter, the transcription rate of the LuxI gene is increased which in turn results in the production of more AHL. | The enzyme '''LuxI''' is an acyl-homoserine-lactone synthase which produces the intercellular signalling molecule N-(3-oxohexanoyl)-homoserine lactone (AHL). Placing LuxI under control of the pRight promoter results in a positive feedback loop: when increases in cell density cause the intracellular AHL concentration to rise above the activation threshold of the pRight promoter, the transcription rate of the LuxI gene is increased which in turn results in the production of more AHL. | ||
'''AiiA''' is an enzyme from B. subtilis which degrades AHL. Its biological function is to interfere with the quorum sensing signals of other bacteria. Placing it under control of the pRight promoter results in negative feedback as a response to increasing AHL concentrations. | '''AiiA''' is an enzyme from B. subtilis which degrades AHL. Its biological function is to interfere with the quorum sensing signals of other bacteria. Placing it under control of the pRight promoter results in negative feedback as a response to increasing AHL concentrations. | ||
| - | The reporter molecule Green Fluorescent Protein ('''GFP''')is also regulated by the pRight promoter and provides a quantitative (albeit delayed) indication of the AHL concentration the cell is exposed to at a given point in time. | + | The reporter molecule Green Fluorescent Protein ('''GFP''') is also regulated by the pRight promoter and provides a quantitative (albeit delayed) indication of the AHL concentration the cell is exposed to at a given point in time. |
| - | LuxI, AiiA and GFP are all tagged for rapid degradation (LVA-tag). Due to differences in the synthesis and degradation rates of LuxI and AiiA, there exists a space of conditions within which periodic oscillations in AHL concentration, and concomitant oscillatory protein expression can emerge. Under most conditions, the level of AHL within a population of cells will quickly reach a steady state. However, by simulating the system using a quantitative biochemical model, it is possible to predict conditions under which oscillations are likely to occur. See our [https://2011.igem.org/Team:Wageningen_UR/Project/ModelingProj1 | + | LuxI, AiiA and GFP are all tagged for rapid degradation (LVA-tag). Due to differences in the synthesis and degradation rates of LuxI and AiiA, there exists a space of conditions within which periodic oscillations in AHL concentration, and concomitant oscillatory protein expression can emerge. Under most conditions, the level of AHL within a population of cells will quickly reach a steady state. However, by simulating the system using a quantitative biochemical model, it is possible to predict conditions under which oscillations are likely to occur. See our [https://2011.igem.org/Team:Wageningen_UR/Project/ModelingProj1 modeling page] for details. |
=====3. Designs===== | =====3. Designs===== | ||
| Line 64: | Line 64: | ||
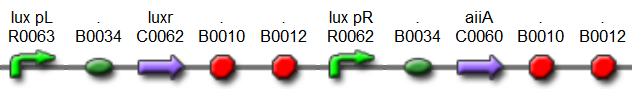
'''Fig.4.''' ''Synchronized Oscillator plasmids using natural quorum sensing system (Danino et al. 2010)'' | '''Fig.4.''' ''Synchronized Oscillator plasmids using natural quorum sensing system (Danino et al. 2010)'' | ||
| - | The first design we implemented was a BioBrick part based reconstruction of the plasmids used by Danino et al. We intended to make as accurate a replica as possible in order to confirm the previously published results, and to test the viability of our experimental [https://2011.igem.org/Team:Wageningen_UR/Project/Devices platform]. However, there are a few differences between the original Hasty system and “our replica“. While the Hasty system employs the natural lux promoter which contains divergent pLeft and pRight elements (Fig 4), the BioBrick parts we employed have both elements in the same orientation. Both the original and our system contain 3 copies of the luxR gene under control of the pLeft element. Furthermore, a different (high copy) backbone was used during the functional validation of the parts, as opposed to the low copy backbone employed by Hasty. | + | The first design we implemented was a BioBrick part based on reconstruction of the plasmids used by Danino et al. We intended to make as accurate a replica as possible in order to confirm the previously published results, and to test the viability of our experimental [https://2011.igem.org/Team:Wageningen_UR/Project/Devices platform]. However, there are a few differences between the original Hasty system and “our replica“. While the Hasty system employs the natural lux promoter which contains divergent pLeft and pRight elements (Fig 4), the BioBrick parts we employed have both elements in the same orientation. Both the original and our system contain 3 copies of the luxR gene under control of the pLeft element. Furthermore, a different (high copy) backbone was used during the functional validation of the parts, as opposed to the low copy backbone employed by Hasty. |
[[File:Hasty1.png|x67px|link=http://partsregistry.org/Part:BBa_K546005]] | [[File:Hasty1.png|x67px|link=http://partsregistry.org/Part:BBa_K546005]] | ||
Revision as of 18:30, 21 September 2011
 "
"