Team:EPF-Lausanne/Notebook/August2011
From 2011.igem.org
(→Friday, 12 August 2011) |
(→Friday, 12 August 2011) |
||
| Line 163: | Line 163: | ||
Lilia made colony PCR on TetR variants prepared by Alina and transformed on Wednesday. | Lilia made colony PCR on TetR variants prepared by Alina and transformed on Wednesday. | ||
| - | [[File:EPFL2011_colony-PCR_TetR-variants_mutants.png ]] | + | [[File:EPFL2011_colony-PCR_TetR-variants_mutants.png|800px]] |
Revision as of 18:45, 12 August 2011
Notebook: August 2011
Contents |
Tuesday, 02 August 2011
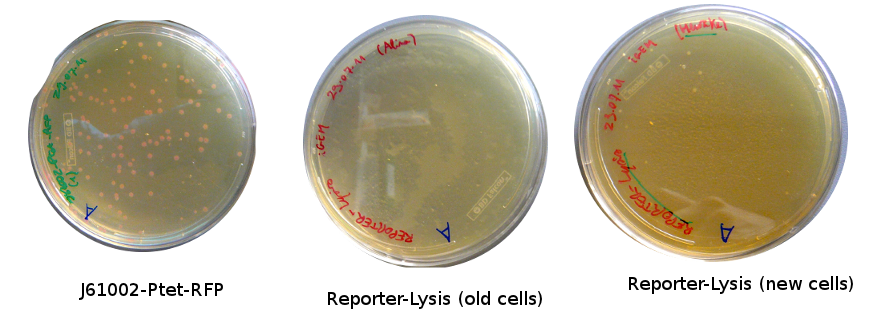
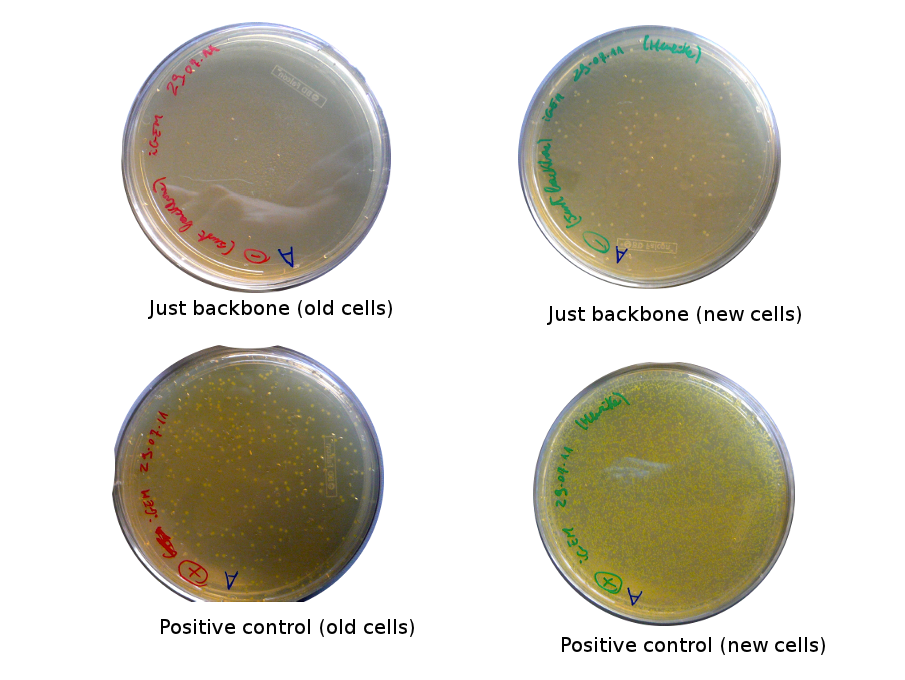
Lilia came on Saturday to take the plates and here are the results:
The old competents cells seem not to work very well and the Gibson assembly failed since in the plate with only backbone we have a lot of cells therefore the backbone can closes on itself.
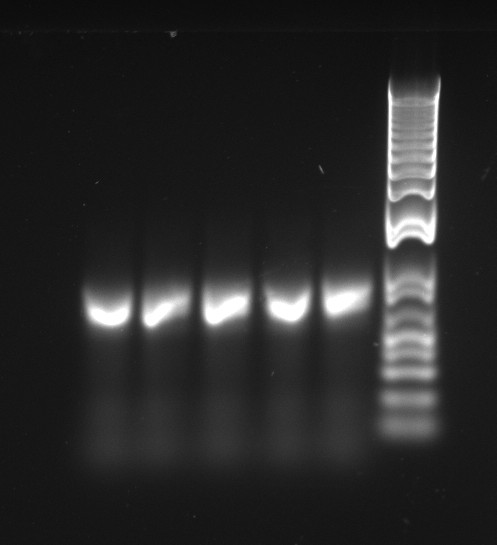
Alessandro make the PCR (gradient PCR in which the annealing temperature increases for each sample) to amplify TetR (putting Pconst in front of it) from Repressilator plasmid to make the Gibson for the Pconst-TetR plasmid. The result is illustrated on the right.
Since we have already amplified the backbone (pSB3K1) we can make the Gibson assembly to make the Pconst-TetR plasmid.
The latest mutation PCR yielded large amounts of product. The two bands might be a little worrying, but overall this is a lot better than any of the previous trials, with the old polymerase. The stitch PCR will be ran tomorrow.
On the side, more reporter and lysis plasmids were assembled (together with a control, containing only the backbone), from the previously made gibson fragments (PCR purified on 2nd August 2011).
Wednesday, 03 August 2011
Alessandro made glicerol stocks of Vincent's plasmid (J61002-Ptet-RFP) and purified the PCR product of the day before (Pconst-TetR) to make the Gibson assembly (using the already amplified backbone of pSB3K1). The Gibson assembly for the pSB3K1-Pconst-TetR (using either equimolar ratio of backbone-insert either 2 times insert) and for Reporter-RFP/Lysis has been plated.
Douglas ran the stitch PCR on the latest mutants, unfortunately yielding virtually no product. Only the PQ39LV31YM42 mutant has detectable amounts of product in the correct size, but then it also had a much larger concentration of template (or so it seems by eyeballing the mutation PCR's gel).
Likely causes:
- The common sequence DNA was lost during PCR purification (Doug accidentally used the high cut-off buffer), in which case the mutation PCR should be repeated from scratch...at least for the common sequence.
- The 3' final and 5' final primers are not suitable for the stitch step. Concentrations and heat cycles should be accurate, because they worked for the mutation PCR.
Nadine transformed and plated Doug's mutant TetR plasmids from 26.07. She prepared the X-gal and IPTG solutions needed for the control, but they still need to be filtered.
We tried a first colony PCR on J61002 Ptet-RFP cells, but nothing got amplified... Tomorrow we'll try the primers on purified plasmids to see it the problem comes from here.
Thursday, 04 August 2011
The mutant TetR plasmids plated yesterday yielded only one colony, even the mutation control didn't work. Nadine plated the rest of transformed cells, but probably the cells in the mutagenesis kit are too old to be competent.
Alessandro took plates from the day before and about 20 colonies grew for Reporter-Lysis with BL-21 and less using DH5alfa cells; for Reporter-RFP just 1 colony grew; about 10 colonies for Pconst-TetR. Alessandro screened some of them with colony PCR. The results are available only for Vincent's plasmid (our positive control) and for the pSB3K1-Pconst-TetR and they are positive (but we cannot estimate the size since the ladder seems to be very degraded). For Vincent's plasmid, together with cells Alessandro also made the PCR with the purified plasmid (to check whether the primers are good).
The following day the results for the other plates will be available and Alessandro will make co-transformation with Vincent's plasmid and with psB3K1-Pconst-TetR in order to assess the expression of RFP, controlled by TetR.
Friday, 05 August 2011
Nadine checked the 2nd plating of the mutagenesis experiment, and again we had cells only for the 4th mutagenesis(about 10 colonies). She made liquid cultures from 5 of them, the miniprep will be done tomorrow.
Colony PCRs for the Reporter-lysis plasmid have been made but they failed... so we have to start over again from Gibson assembly. Only one sample for Reporter-RFP has been tested, and the colony PCR also failed.
Alessandro and Alina made Kanamicine plates and Kanamicine+Ampicillin plates. Alessandro transformed:
- pSB3K1
- Vincent's plasmid (J61002-Ptet-RFP)
- pSB3K1 Pconst-TetR
- the old Gibson assembly for Reporter-RFP
- co-transformation with pSB3K1-Pconst-TetR and Vincent's plasmid
The transformation were performed for pSB3K1 because unexpectdely the liquid cultures for pSB3K1-Pconst-TetR (positive with colony-PCR) were red. On the registry the pSB3K1 backbone hasn't been documented to have RFP so the question why this cells were red is open. This new assembly has to be sequenced.
Monday, 08 August 2011
Lilia went on Saturday to miniprep the TetR mutants, and the concentrations are really high: from 150 ng/ul to 238 ng/ul.
Nadine made new Gibson reactions for the Reporter and Lysis plasmids: once with equimolar ratios and once with about 2x more parts than the backbone, plus a negative control containing only the backbone.
Alessandro prepared SOC medium and started liquid culture of pSB3K1 and pSB3K1-Pconst-TetR to make glycerol stocks. The cells transformed with pSB3K1 are red therefore that plasmid has RFP though not documented on the registry. A lot of colonies grew on the co-transformation plate; Alessandro will start a liquid culture the following day to test them in the plate-reader.
Vincent used the QuikChange Site-Directed Mutagenesis (SDM) Kit to try to produce TetR mutants. The pWhitescript 4.5-kb control plasmid provided by the Kit was used. The plasmid contains a stop codon (TAA) rather than a glutamine codon (CAA) in the beta-galactosidase gene of the pBluescript II SK(-) phagemid. When we transform the plasmid into XL10-gold ultracompetent cells, the resulting cells should look white on LB-ampicillin plates that have been treated with IPTG and X-gal, since the beta-galactosidase is no longer functional. However, if the mutagenesis works, there should be a point mutation that reverts the T into a C (TAA into CAA), meaning that the cells should appear blue on plates that contain IPTG and X-gal.
The protocol for Mutant Strand Synthesis (provided with the Kit) was followed meticulously. Vincent made a master mix for 5 mutants and made a separate control mixture. The DNA template concentration was 130 ng/uL (after being diluted from 160 ng/uL as used by Douglas) so we used .5 uL to get around 65 ng per mutant sample. The protocol asks for 125 ng of each primer. Since the primers all had the same 1 ug/uL concentration, we used 8 uL of each primer. The mutants were the following:
- Y42F
- V36F
- P29Q-Y42M
- P39Q-LV41
- P39K
Since the total strand is about 5 kb, we used 150 seconds at 68 C for the second segment of thermal cycling. The resulting cycling products were put in the fridge over night, since it was too late to start the digestion.
Tuesday, 09 August 2011
Plates with yesterday's Gibson assemblies were not concluent, as there were few colonies without RFP and not more than in the negative control. Alessandro and Nadine made a gradient Gibson assembly for reporter plasmid and a negative control (=only backbone), we will transform them as soon as the competent cells are ready.
The mutagenesis cycling products were digested using 2 uL of Dpn I restriction enzyme. The transformation of XL10-Gold cells was done normally. The ampicillin LB plates were melting, making beading a problem. The plates were put overnight.
Lilia run MITOMI on spotted de Brujin library with GFP-wtTetR expressed off-chip with SP6 ITT. Before the last scan, the chip was washed by mistake with opened buttons, so the interactions unfortunately might have been washed on the next images.
Wednesday, 10 August 2011
Henrike and Alessandro designed primers to put the T4 lysis cassette (BBa_K112808) under the control of the T7 promoter using as backbone: pSB3C5, pSB3K1, pSB3K5. This device will be used to test plasmid DNA recovery after lysis.
Nadine designed primers to add Ptet-LacI in the pSB3K1-Pconst-TetR palsmid, also adding a terminator before LacI. We decided not to inculde the ssrA degardation tag in LacI, to be sure that the cells won't lyse.
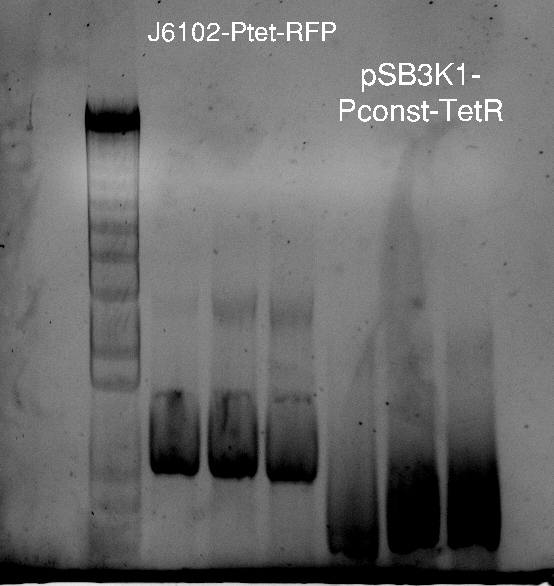
Since the pSB3K1 plasmid contains RFP, Nadine tried to cut it out by digesting it with XbaI and SpeI. Tomorrow we'll make a ligation, and as soon as the primers arrive we'll be able to add LacI with Gibson assembly. Ligation results show 2 bands, as expected, however they are too big...
Vincent got the plates containing the tetR mutants: only the control worked, and not spectacularly well at that. The LB plates were not very good, so a new batch of plates was made with what was left of the previous day's transformation (kept overnight in the 4 C fridge). Unlike the previous plates, the sample mutagenesis plates were not covered with IPTG and Xgal, since these mutageneses do not benefit from such treatment.
Thursday, 11 August 2011
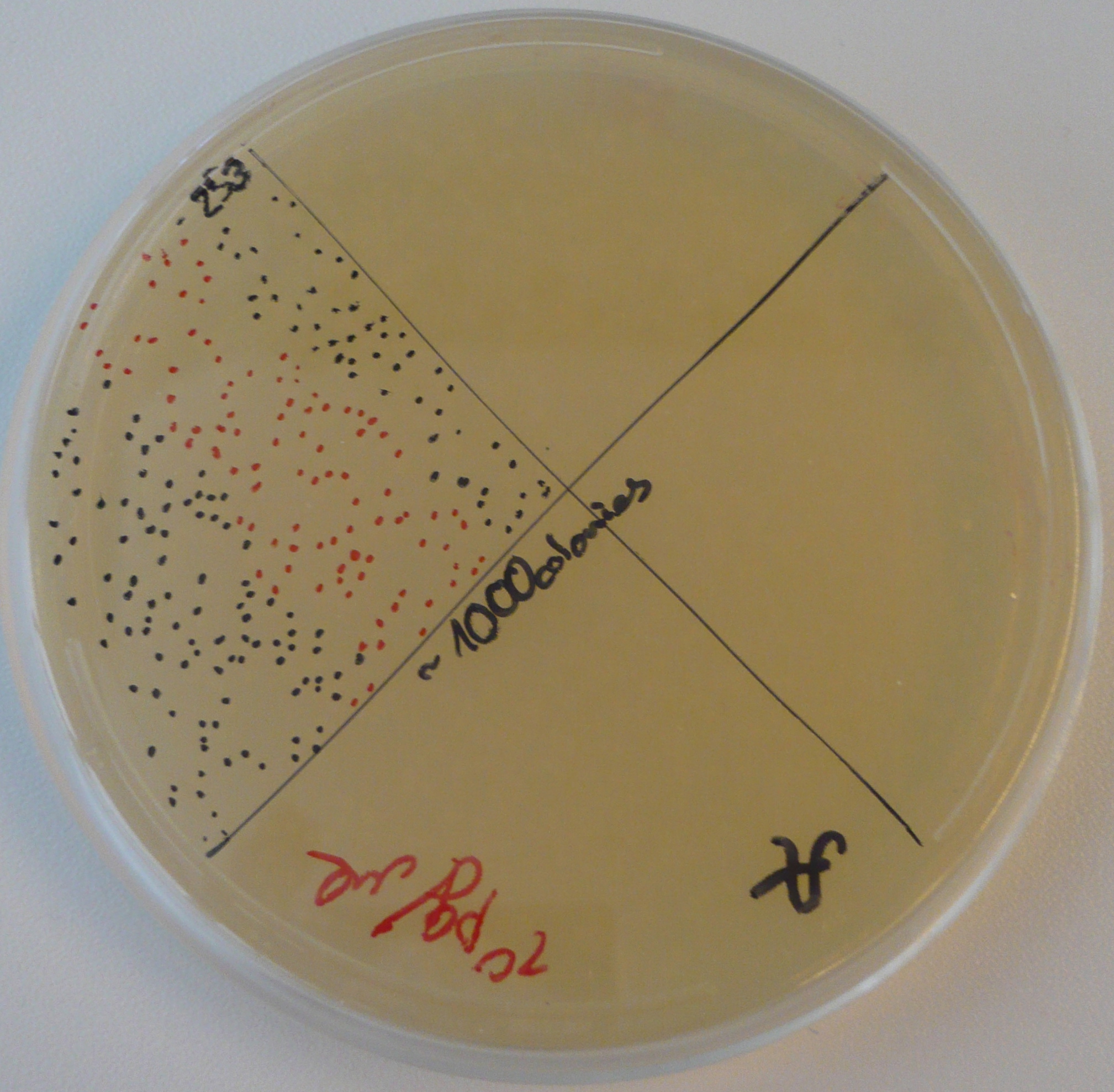
Lilia measured the competence of DH5alpha cells prepared by Clara, Lilia and Alina according to the second (Inoue) protocol for chemically competent cells, they were pipetted on Tuesday by Clara and Lilia. On Wednesday 20µl of cells were transfected with 100, 50 and 10pg of the plasmid containing GFP-TetR. The 10pg plate is shown on the picture. Competence is 10^8 CFU/µg.
Nadine ligated the pSB3K1 Pconst-tetR that should not contain RFP anymore. She also transformed it into Henrike's and our new competent cells.
Vincent got the fresh plates out of the incubator, only to find that the P39Q-Y42M was the only mutation that had grown colonies (only 2), which we consider to be an accident given that there ought to be hundreds of colonies. After poring through the protocol yet again, we spotted a mistake in the original dilution approach. Vincent went ahead and re-ran another PCR, digestion, transformation sequence, and we will see how the plates look tomorrow.
Alessandro searched literature for T7 variant with different strength to use them in the T7-Lysis plasmid.
Lilia run MITOMI on spotted de Brujin library with GFP-wtTetR expressed off-chip with SP6 ITT. Everything went well, but no strong interactions were seen.
Chip fabrication, spotting, alignement and the whole experiment was done by Lilia. Clara's wafers were used.
Kanamycin aliquotes of 50mg/ml in water and Ampicilin aliquotes 100mg/ml in 50% ddH2O and 50% ethanol were prepare by Lilia and put in the new box labelled "iGEM2011 Antibiotics" at -20°C in BM building.
Friday, 12 August 2011
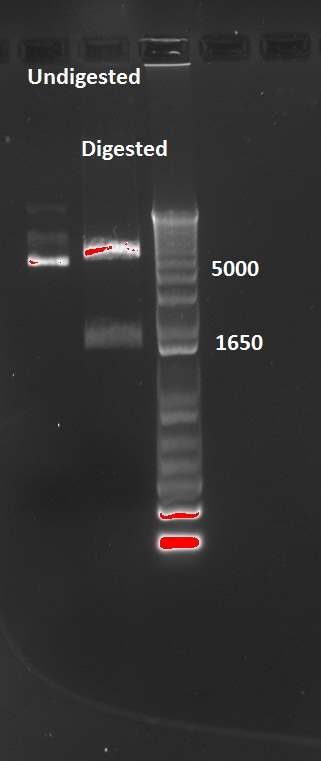
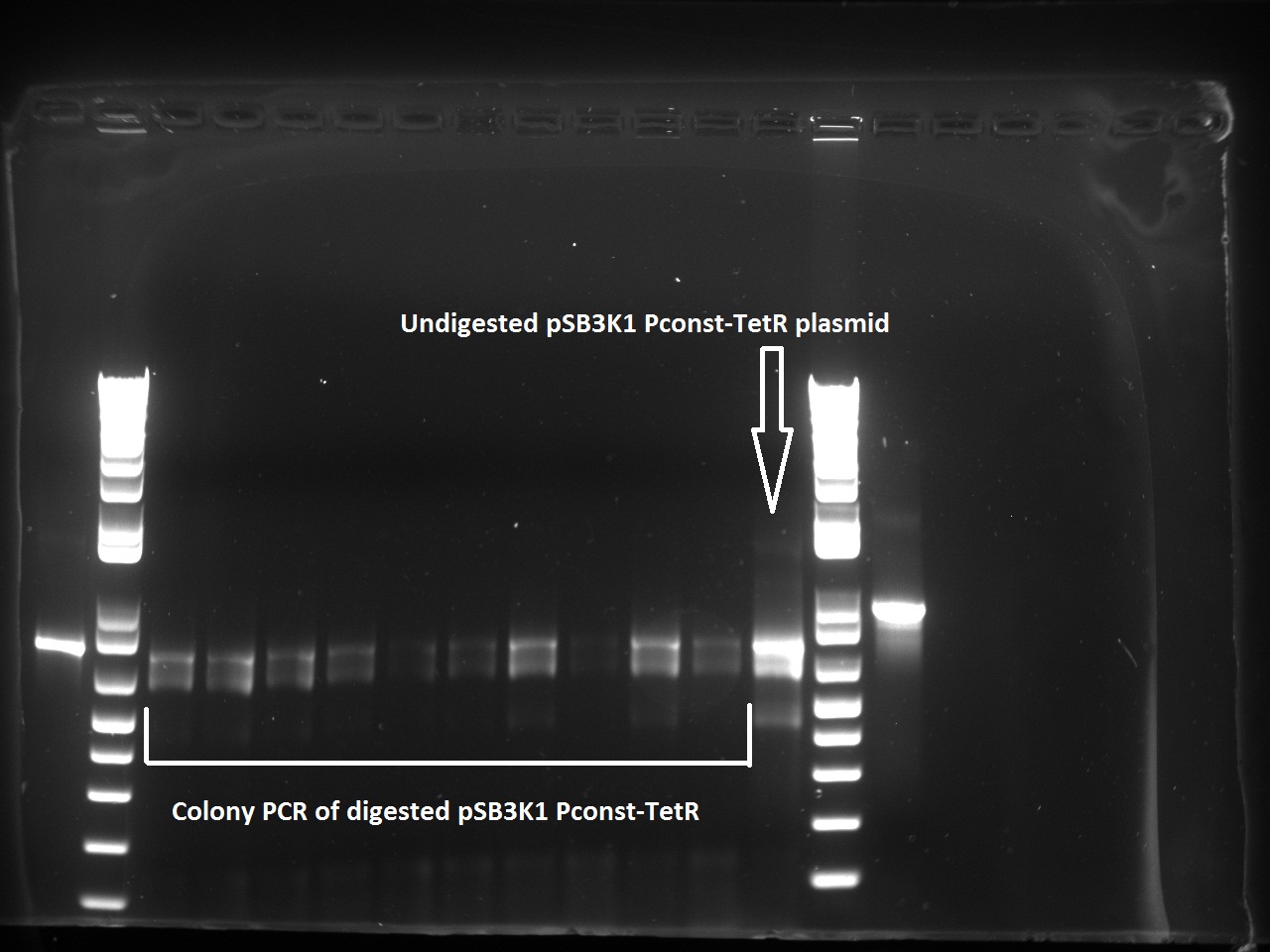
The transformed cells with digested pSB3K1 Pconst-tetR yielded only white colonies, which indicate that RFP actually got cut out. Nadine also made a colony PCR on 10 of the colonies, because the primers include the multiple cloning site. As a control, the same PCR was made on undigested pSB3K1 Pconst-tetR plasmid. The expected size for the digested would be 1200 kb, whereas with RFP it should be around 2000 bp. But the gel gives disappointing results, with every colony and the control showing the same ~800 bp band... Liquid cultures have been made, and if they don't turn red we'll miniprep them and send to sequencing.
Alessandro made the extension PCR on RPF to put it under the control of 4 different T7 promoter: T7const (almost 100%), T7lac (T7 promoter with different versions of a lac operator downstream). The gel will be run on Monday as well as the gene specific PCR for T4 lysis cassette.
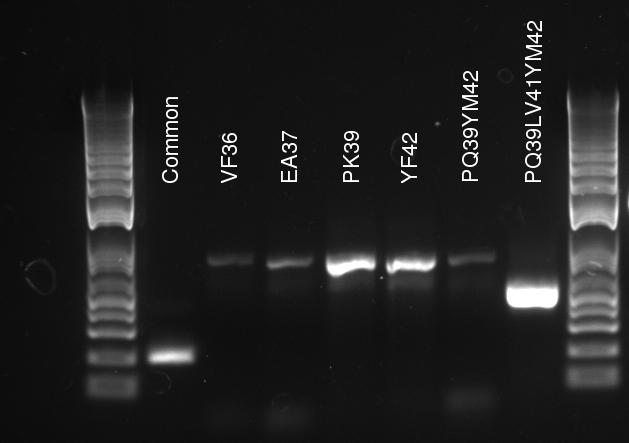
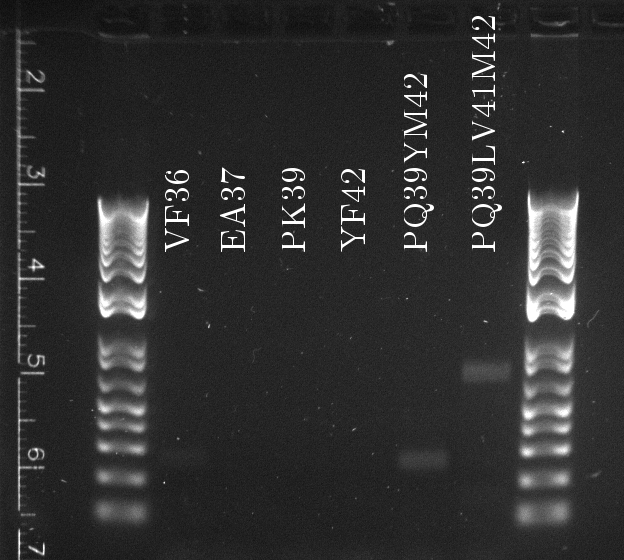
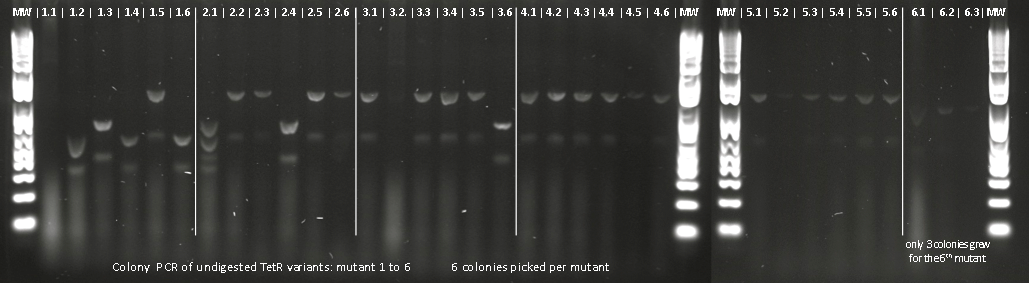
Lilia made colony PCR on TetR variants prepared by Alina and transformed on Wednesday.
 "
"