Team:Arizona State/Project/Software
From 2011.igem.org
Rubenacuna (Talk | contribs) (Fixed some typos.) |
Rubenacuna (Talk | contribs) (Adding text.) |
||
| Line 1: | Line 1: | ||
{{:Team:Arizona State/Templates/main|title=CRISPRstudio|content= | {{:Team:Arizona State/Templates/main|title=CRISPRstudio|content= | ||
| | ||
| - | [[File:ASU crisprstudio ss.jpg|thumb| | + | [[File:ASU crisprstudio ss.jpg|thumb|CRISPRstudio information panel]] |
We have developed a tool to assist in the development of synthetic CRISPR systems. | We have developed a tool to assist in the development of synthetic CRISPR systems. | ||
* Pick spacers from a source sequence, based on homology with the target genome, hairpinning potential, restriction sites, and known PAMs. | * Pick spacers from a source sequence, based on homology with the target genome, hairpinning potential, restriction sites, and known PAMs. | ||
| Line 34: | Line 34: | ||
'''Interface''' | '''Interface''' | ||
---- | ---- | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| + | <p>''All screen shots were taken on Windows 7, however, CRISPRstudio was designed with portability in mind and will look similar on other operating systems.''</p> | ||
| + | | ||
| + | <center>[[File:ASU crisprstudio ss2.jpg|450px|Update prompt]]</center> | ||
| + | <p>When first launching CRISPRstudio, users are greeted with the above screen. This screen prompts the user to update the local database of CRISPR information. Press y to update or n to continue without updating. An internet connection is required. Note that we directly interface with several online databases and changes in their formats may cause our updater to fail. In this case, the data provided with CRISPRstudio may be used. Release packages of CRISPRstudio include data that was current at time of release.</p> | ||
| + | | ||
| + | <center>[[File:ASU crisprstudio ss.jpg|500px|Information panel]]</center> | ||
| + | <center>[[File:ASU crisprstudio ss3.jpg||Menu]]</center> | ||
| + | <p>In the upper right side of the screen, you will see a menu bar. The view menu allows the user to switch between viewing CRISPR information and generating spacers. The settings menu controls how spacers are generated. A help menu that provides no actual is also available for piece of mind while using the software.</p> | ||
| + | | ||
| + | <center>[[File:ASU crisprstudio ss4.jpg|500px|Spacer creation panel]]</center> | ||
| + | <center>[[File:ASU crisprstudio ss5.jpg||Finalize Spacers]]</center> | ||
| + | <center>[[File:ASU crisprstudio ss6.jpg||Spacer Generation Settings]]</center> | ||
| + | The Spacer Generation Settings dialog controls which REs are considering when creating spacers. The spacer creator requires that a set of REs does not exist in potenial spacers. The set of REs is defined here. On the left is a list of currently selected REs, these may be removed by selecting an item and clicking Remove Site. On the right is a list of all REs known in BioPython. Any of these may be added by selecting on it and clicking Add Site. Alternatively, the name of a RE (e.g. EcoRI) may be entered in the text box and adding by clicking the button. The selected REs are stored locally and will be loaded next time CRISPRstudio is run. | ||
'''Data Sources''' | '''Data Sources''' | ||
Revision as of 00:52, 28 September 2011
|
|
We have developed a tool to assist in the development of synthetic CRISPR systems.
Downloads
Installation Using the provided binaries is the fastest way to get started with CRISPRstudio. Visit our GoogleCode page and select the downloads tab. Download the appropriate build based on your OS. Builds are provided using [http://cx-freeze.sourceforge.net/ cx_Freeze]. All builds are provided for x86 platforms. For real x64 support, you will need to run the source directly. x64 support has not shown significant speed increases for CRISPRstudio. Extract the compressed file to some location. You also need to download BLAST+ 2.2.25 or compatible. BLAST+ must be installed and included in the system path. On most systems, the BLAST+ installer will take care of this. After this, you may run MainFrame to start the program. Source
Pick os/arch as needed. Note that each of the python packages should list the version of Python they support - select the right one. NumPy must be installed before BioPython. The x64 release of wxPython doesn't seem to like Win7 so it may be better to use the x86 release. BLAST+ must be included in the system path. If BLAST+ is not available in the the system, the spacer creator will not function. The information browser will still function normally. The code itself is not fully documented. However, the code was intentionally written to be readable without comments. Project files are available for PyCharm 1.6. You may then run MainFrame.py to start the program. Interface All screen shots were taken on Windows 7, however, CRISPRstudio was designed with portability in mind and will look similar on other operating systems.
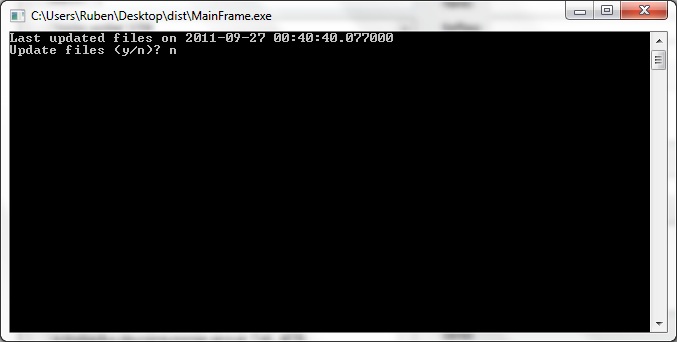 When first launching CRISPRstudio, users are greeted with the above screen. This screen prompts the user to update the local database of CRISPR information. Press y to update or n to continue without updating. An internet connection is required. Note that we directly interface with several online databases and changes in their formats may cause our updater to fail. In this case, the data provided with CRISPRstudio may be used. Release packages of CRISPRstudio include data that was current at time of release.
 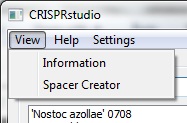 In the upper right side of the screen, you will see a menu bar. The view menu allows the user to switch between viewing CRISPR information and generating spacers. The settings menu controls how spacers are generated. A help menu that provides no actual is also available for piece of mind while using the software.
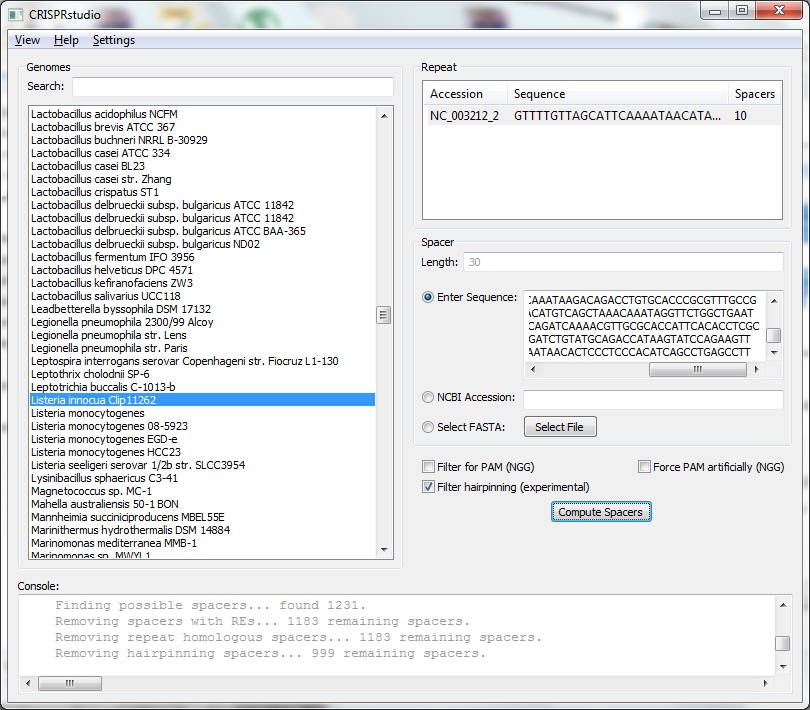 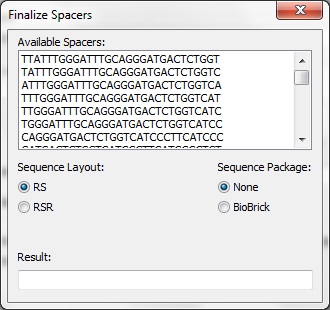 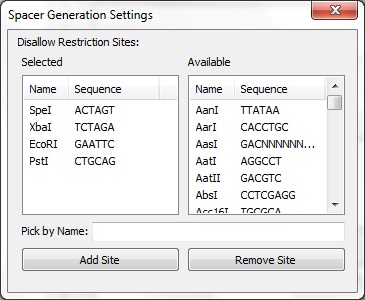 The Spacer Generation Settings dialog controls which REs are considering when creating spacers. The spacer creator requires that a set of REs does not exist in potenial spacers. The set of REs is defined here. On the left is a list of currently selected REs, these may be removed by selecting an item and clicking Remove Site. On the right is a list of all REs known in BioPython. Any of these may be added by selecting on it and clicking Add Site. Alternatively, the name of a RE (e.g. EcoRI) may be entered in the text box and adding by clicking the button. The selected REs are stored locally and will be loaded next time CRISPRstudio is run. Data Sources
|
 "
"
