results
Precipitator
Plastic binding domain
Green light receptor
We could successfully amplify the gene sequences for CcaS and CcaR and the promotor region PcpcG via
PCR from the whole Synechocystis sp. PCC6803 genome.
Later two parts ([http://partsregistry.org/Part:BBa_K608101 BBa_K608101] , [http://partsregistry.org/Part:BBa_K608102 BBa_K608102]), were inserted into the iGEM-Vector pSB1C3 and send to the registry.
Twice we tried to clone the promotor region PcpcG into a vector, but had no insert at all.
After a summer full of cloning we ran out of time to assemble
the green light receptor cassette like we planned to.
red light receptor
In August we decided to drop the red light receptor system.
The main reason beeing Jakob having to leave the team to start studying in Strasbourg.
So the FreiGEM team shrunk to five people.
We designed this composite in order to gain the enzyme pcyA (phycocyanobilin-ferredoxin oxidoreductase),
this design is necessary if the part is going be integrated into an assembly of various genes.
The enzyme plays an important role in the production of the PCB chromophore (phycocyanobilin)
which is essential for the green and red light receptor system.
Our sequencing confirmed that the part is correct in the pSB1C3 vector.
All physical DNA samples used for this composite come from the registry distribution.
Still we did submit one part for this system which is [http://partsregistry.org/Part:BBa_K608151 BBa_K608151]
the translational unit of pcyA
We talked to some of the iGEM teams like TU Munich also working with this system
and we are sure that the red light receptor is not lost.
Usage and Biology
For PCB chromophore production the enzyme [http://partsregistry.org/Part:BBa_I15008 ho1] (heme oxygenase)
in a similar design is necessary infront of this part
because both enzymes are needed to convert heme into PCB chromophore.
references
Ken Okada
"HO1 and PcyA proteins involved in phycobilin biosynthesis form a 1:2 complex with ferredoxin-1 required for photosynthesis"
FEBS letters, Volume 583, Issue 8, 17 April 2009, Pages 1251-1256
blue light receptor
The handling with the LOV domain and TrpR domain was very difficult.
On the one hand it was difficlut to digest the parts, because there was a SpeI restriction site in the tryptophan repressible promoter region. So we first had to mutate SpeI into NehI restriction site to digest our parts without digesting the promotor. In the beginning we also tried the Gibson assembly. Primer design was quite difficult because of the lenght, the usually very high annealing temperature and unspecifity of the primer sequences, combined to the mutations we had to include. These hassel leaded to trouble to amplify the Lov domain.The 3A-assembly with the mutated parts wasn´t successful either. We only cloned the Not-Gate in the pSB1C3 vector. We tried this experiment with different parameters but we weren´t successful.
Lysis cassette
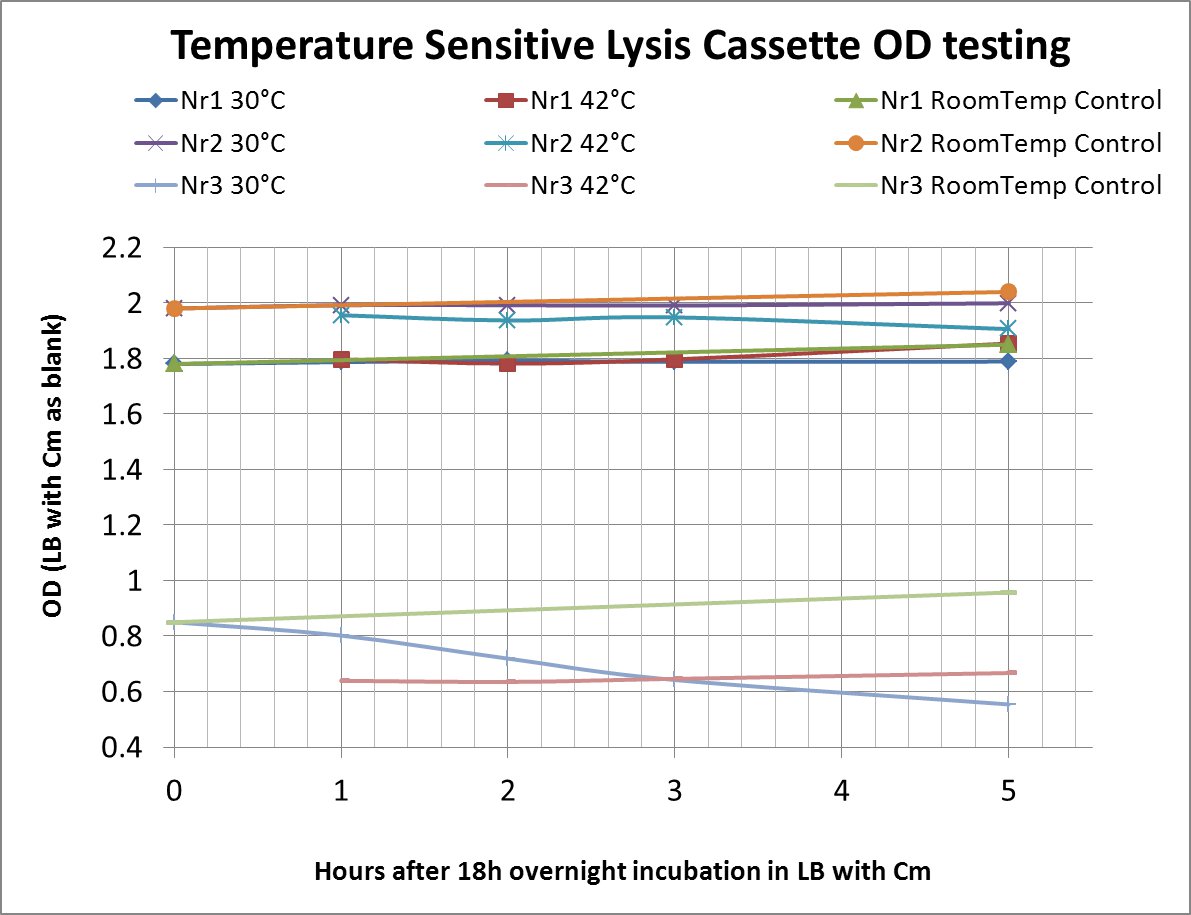
After a lot of mishappenings concerning the 2 parts making up the temperature sensitive lysis cassette, we made a last attempt to ligate them (link to workbook) and transform the novel composite over the last weekend before the Wiki-freeze. After putative positive colonies were picked, we managed to perform a last-minute, "blind" -ie not knowing if the sequence of the insert was correct-, and preliminary OD-measurement testing.
We later verified the part sequence. Since we cannot upload .ab1 files, we made 2 base-call files from them without changing absolutely anything (that means that there are some false bases at the beginning and end of every file).
- Forward Primer
- Reverse Primer
The OD values of the measurement are shown in the chart (please do bear in mind that this was a measurement done under deadline-stress, without the possibility to plan and perform something more concrete before the Wiki-Freeze)
- LB with Cm (Chloramphenicol) was used as a blank before every (hourly) measurement
Nr1: Insert is only part ([http://partsregistry.org/Part:BBa_K608351 BBa_K608351])
Nr2: Insert is only part ([http://partsregistry.org/Part:BBa_K608352 BBa_K608352])
Nr3: Both parts were correctly assembled (1:([http://partsregistry.org/Part:BBa_K608351 BBa_K608351]) + 2:([http://partsregistry.org/Part:BBa_K608352 BBa_K608352]))
Therefore the temperature sensitive lysis cassette seems to be pretty temperature-sensitive but also functioning properly. More tests are planned if time allows (ie wiki-unfreeze ;) )
Parts submitted to the registry
All our results are documented in our notebook and summarized in the partsregistry.
Here we shortly describe what happened to give you an idea about the current status of the different subprojects.
Please follow the links for more information.
Precipitator
[http://partsregistry.org/Part:BBa_K608404 BBa_K608404]
IPTG-inducible Promoter with plastic binding domain-tagged GFP
[http://partsregistry.org/Part:BBa_K608406 BBa_K608406]
Precipitator
Protein domain of Precipitator. Artificial Leucine Rich Repeat(LRR) with C and N-terminal hagfish domain fragments capping the artifical middle part. This part is one version of three different designed to bind nickel by histidines,
grouped together pointing away from the horseshoe shaped protein.
Bacterial LRR Consensus of the central LRR fragment:
LxxLxLxxNxLxxLPxxLPxx
Protein code:
CPSRCSCSGTEIRCNSKGLTSVPTGIPSS
ATRLELESNKLQSLPHGVFDK
LTQLTKSNNHLHSLPDNLPAS
LEVLDVSNNHLHSLPDNLPAS
LEVLDVSNNHLHSLPDNLPAS
LEVLDVSNNHLHSLPDNLPAS
LEVLDVSNNHLHSLPDNLPAS
LEVLDVSNNHLHSLPDNLPAS
LKELALDTNQLKSVPDGIFDR
LTSLQKIWLHTNPWDCSCPRIDY
LSRWLNKNSQKEQGSAKCSGSGKPVRSIICP
This protein can be used to complex Nickel or Cobalt.
The principal mechanism is comparable to Ni-NTA columns, as chelates the ions. Free binding sites of the ions are then exposed, so that a His-tagged protein can attach to them.
The design of the protein is of a particular interest, too. LRR are highly conserved motifs throughout evolution. They appear in all kingdoms of life in almost every thinkable role (Ligases, Receptors, Toxins etc.). Their core is highly conserved and provides a very stable backbone, while the intermediate, non-conserved aminoacids are almost freely interchangeable.
This protein can be used to complex up to 4 Nickel or Cobalt. However the principal design oft he protein is of a particular interest, too. LRR are highly conserved motifs throughout evolution. They appear in all kingdoms of life in almost every thinkable role (Ligases, Receptors, Toxins etc.). Their core is highly conserved and provides a very stable backbone, while the intermediate, non-conserved aminoacids are almost freely interchangeable.
We only submitted one of the three versions, to reduce redundancy in the registry. Please contact us for any questions.
 "
"
 Contact
Contact