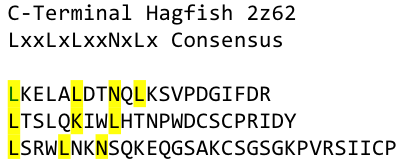Team:Freiburg/Modelling
From 2011.igem.org
(Difference between revisions)
(→Modelling: Rational protein design) |
(→Modelling: Rational protein design) |
||
| Line 14: | Line 14: | ||
Furthermore, many of the natural LRR proteins – of course – have their own biological function, we we were afraid could interfere in the cellular system we wanted to express them in. Luckily there are over a thousand already known structures of LRR proteins, what gave us a broad choice of motifs to choose and compare. | Furthermore, many of the natural LRR proteins – of course – have their own biological function, we we were afraid could interfere in the cellular system we wanted to express them in. Luckily there are over a thousand already known structures of LRR proteins, what gave us a broad choice of motifs to choose and compare. | ||
The Construction of a new protein | The Construction of a new protein | ||
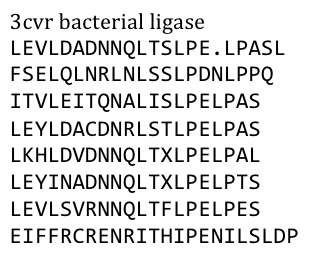
| - | After a long and detailed search, we found out that it is most reasonable to use bacterial LRR motifs, since they seemed very well conserved in sequence, and they are the shortest – with only 21 | + | After a long and detailed search, we found out that it is most reasonable to use bacterial LRR motifs, since they seemed very well conserved in sequence, and they are the shortest – with only 21 amino acids per LRR repeat. (Wei 2008, Kajava 1998). We wanted to have the protein as simple and as predictable in behavior and structure as possible. |
Several search inquiries led to the most conserved and shortest of all bacterial LRR (PDB: NO), unluckily this protein is a toxin derived from Yersinia pestis. We of course did not want to mess around with toxins, | Several search inquiries led to the most conserved and shortest of all bacterial LRR (PDB: NO), unluckily this protein is a toxin derived from Yersinia pestis. We of course did not want to mess around with toxins, | ||
{| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" align=right | {| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" align=right | ||
| Line 30: | Line 30: | ||
Caption | Caption | ||
|} | |} | ||
| - | By analyzing the logo it was obvious which positions of the LRR motif were conserved and which not – AND: which of the non conserved | + | By analyzing the logo it was obvious which positions of the LRR motif were conserved and which not – AND: which of the non conserved amino acids appeared in what kind of patterns. Were there positions in the protein that required a polar amino acid? Or non polar, hydrophobic /-philic, charged, non-charged?. We compared the consensus sequence with the 3D structure, using PYMOL, to extract as much information as possible and then came up with this ideal consensus sequence: |
{| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" | {| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" | ||
|[[File:Freiburg11_Seq2.png|300px]] | |[[File:Freiburg11_Seq2.png|300px]] | ||
| Line 36: | Line 36: | ||
|} | |} | ||
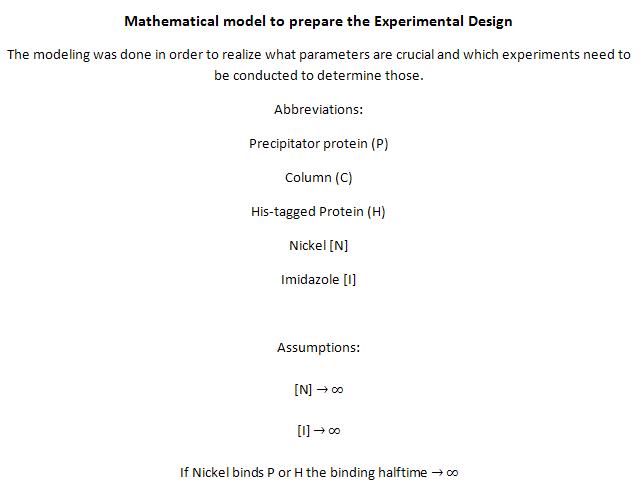
| - | The next consideration we had to do, was: how many Nickel do we need on the surface of our ideal Nickel binding protein, in what pattern, with what distances between, and at what angles towards each other to allow proper ion | + | The next consideration we had to do, was: how many Nickel do we need on the surface of our ideal Nickel binding protein, in what pattern, with what distances between, and at what angles towards each other to allow proper ion complexion? |
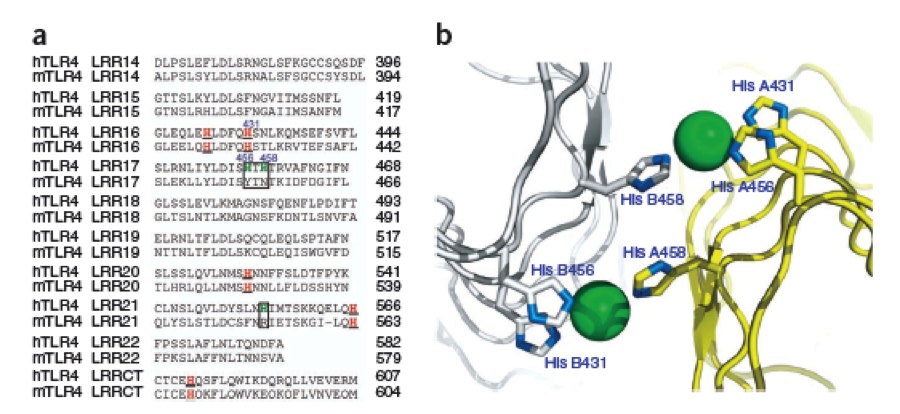
Nickel can be complexed by imidazole structures from 4 planar orthogonal directions, as well as to axial positions. It can however only take four ligands at once, preferably in a planar orientation. Cobalt has a bipyramidal setup for ligand-binding, too, but can take up to six ligands. The distances from a N-atom in the Imidazole ring to the ion had to be between 3 and 6 Angström. We found a crystal structure of a different protein (PDB:) which was resolved with three Histidines complexing a Nickel ion for a comparison, as well as some old publications that analyzed peptide ion bonds that taught us how the complex should look like. (Jordan 1974) | Nickel can be complexed by imidazole structures from 4 planar orthogonal directions, as well as to axial positions. It can however only take four ligands at once, preferably in a planar orientation. Cobalt has a bipyramidal setup for ligand-binding, too, but can take up to six ligands. The distances from a N-atom in the Imidazole ring to the ion had to be between 3 and 6 Angström. We found a crystal structure of a different protein (PDB:) which was resolved with three Histidines complexing a Nickel ion for a comparison, as well as some old publications that analyzed peptide ion bonds that taught us how the complex should look like. (Jordan 1974) | ||
We wanted our protein to be short. First, to cause as little unspecific binding as possible, second, to allow an easy expression and third to keep costs los for gene synthesis. For a proper purification of His-tagged proteins in Ni-NTA columns we knew that there are up to three Nickel involved in the interaction between the His tag (which consists of 6 or 7 Histidines) and the column. These have to be in close proximity to one another. There are always two Histidines of the tag binding to one ion. | We wanted our protein to be short. First, to cause as little unspecific binding as possible, second, to allow an easy expression and third to keep costs los for gene synthesis. For a proper purification of His-tagged proteins in Ni-NTA columns we knew that there are up to three Nickel involved in the interaction between the His tag (which consists of 6 or 7 Histidines) and the column. These have to be in close proximity to one another. There are always two Histidines of the tag binding to one ion. | ||
| Line 42: | Line 42: | ||
This was what we had to implement into our LRR backbone. | This was what we had to implement into our LRR backbone. | ||
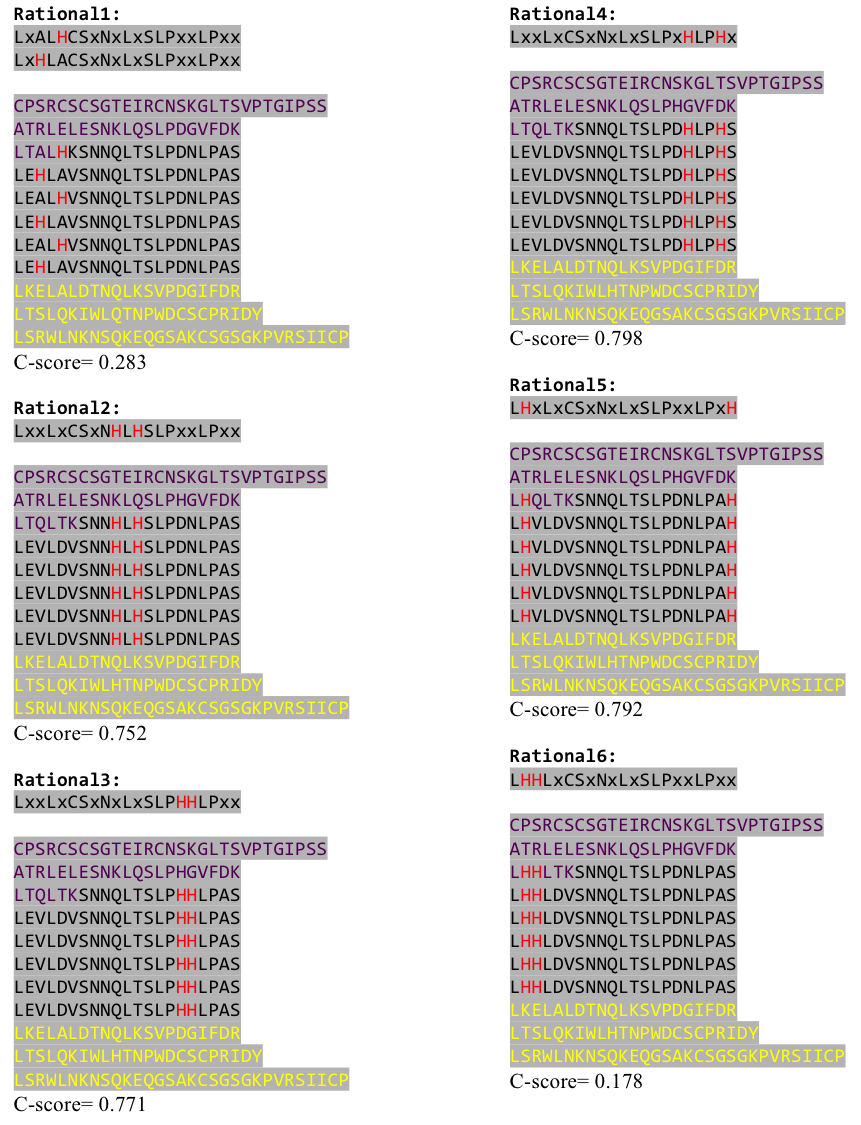
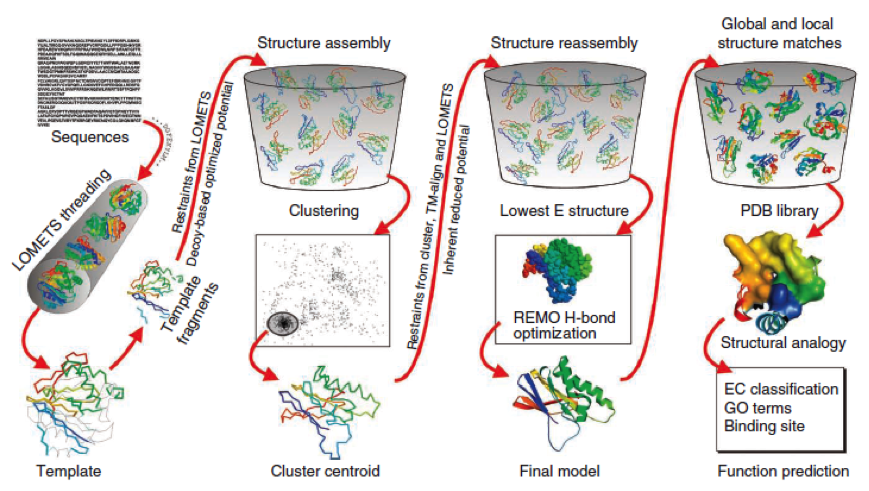
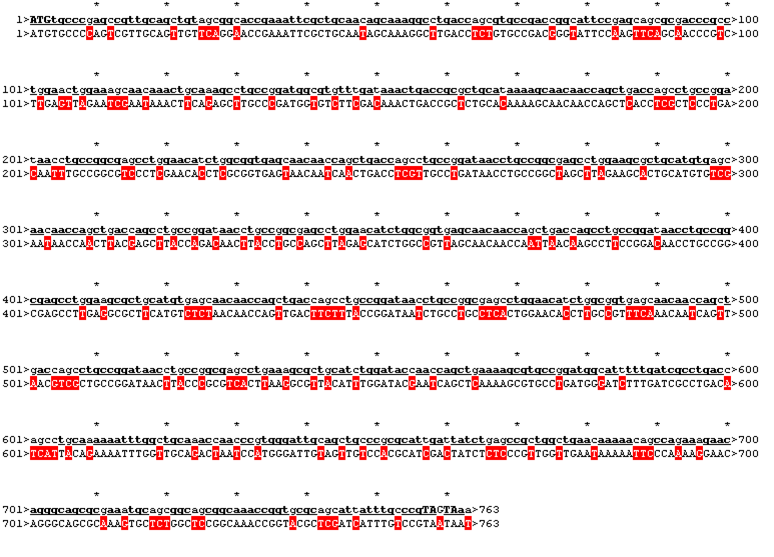
| - | To get a first impression how Histidines are positioned in a bacterial Ligase, we replaced all non conserved | + | To get a first impression how Histidines are positioned in a bacterial Ligase, we replaced all non conserved amino acids by Histidines and sent the sequence for a 3D sequence prediction to I-TASSER, an online structure prediction software tool. |
{| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" | {| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" | ||
|[[File:Freiburg11_Seq7.png|300px]] | |[[File:Freiburg11_Seq7.png|300px]] | ||
| Line 77: | Line 77: | ||
|} | |} | ||
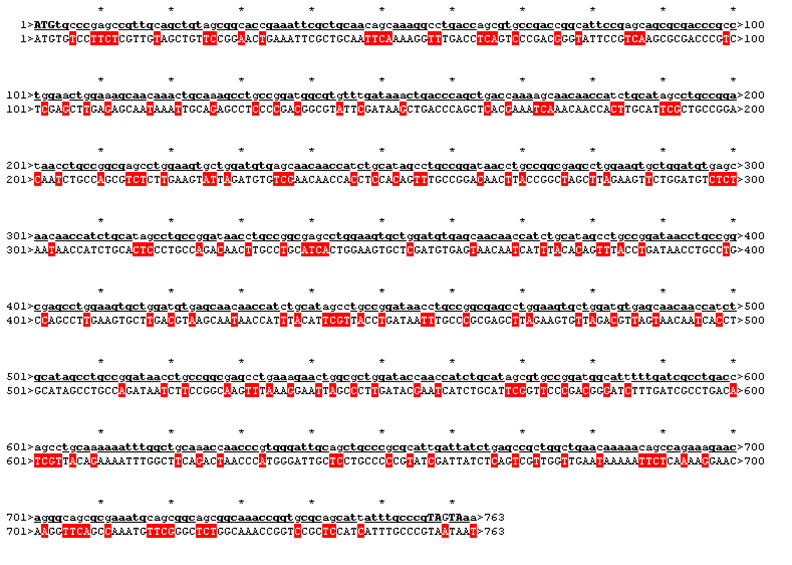
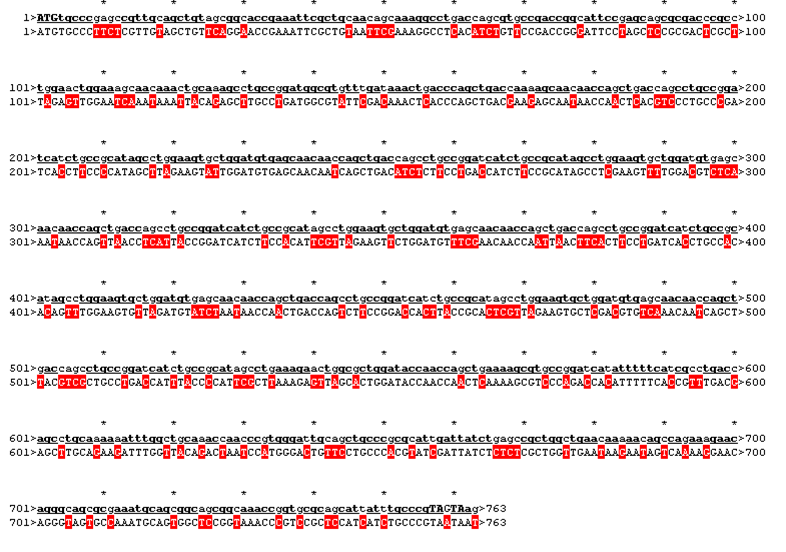
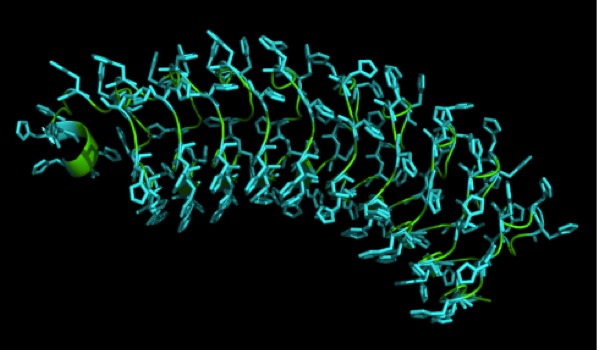
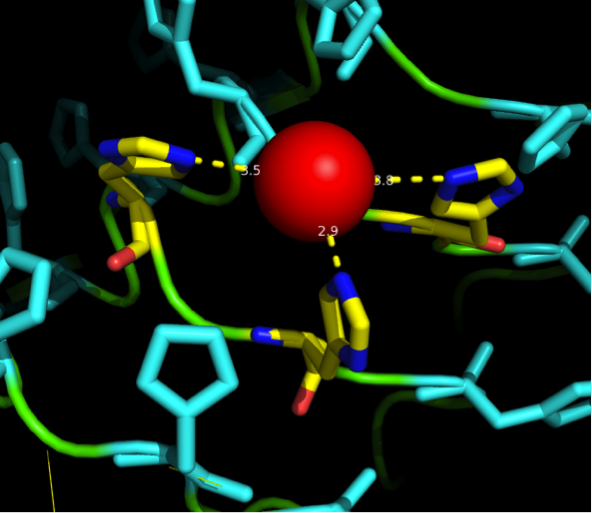
| - | We received this pdb file with a C-score= 0.53. Obviously this was not a realistic model, since the folding would surely be impaired by so many Histidines everywhere. But is was useful to find several good locations, were the Nickel could fit in in respect to the requirements it has for binding ligands.We manually fit in a Nickel ion, as it was crystallized in the mentioned PDB file, and measured the distances and evaluated the 3dimensional orientation of the Histidines towards the Nickel. Histidines can coordinate ligands with their free electron pair pointing planar away from the imidazole ring. What we further realized from the structure file was, that the end of the LRR segments were “open”, that means, the hydrophobic core of the protein was exposed and as it is visible in the prediction, curled in on one end into a sort of helix. This means the protein folding is not reliable and the structure needs some caps on both ends to stabilize the LRR core motif. | + | We received this pdb file with a C-score= 0.53. Obviously this was not a realistic model, since the folding would surely be impaired by so many Histidines everywhere. But is was useful to find several good locations, were the Nickel could fit in in respect to the requirements it has for binding ligands. We manually fit in a Nickel ion, as it was crystallized in the mentioned PDB file, and measured the distances and evaluated the 3dimensional orientation of the Histidines towards the Nickel. Histidines can coordinate ligands with their free electron pair pointing planar away from the imidazole ring. What we further realized from the structure file was, that the end of the LRR segments were “open”, that means, the hydrophobic core of the protein was exposed and as it is visible in the prediction, curled in on one end into a sort of helix. This means the protein folding is not reliable and the structure needs some caps on both ends to stabilize the LRR core motif. |
{| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" align="right" | {| style="color:black; background-color:lightgrey;" cellpadding="10%" cellpadding="15%" cellspacing="0" border="1" align="right" | ||
|[[File:Freiburg11Modelling5.png|250px]] | |[[File:Freiburg11Modelling5.png|250px]] | ||
| Line 112: | Line 112: | ||
| - | To the outer end the protein has a helix on the N-terminal side shielding off the core and the C-terminus the LRR disappears slowly turn by turn with more and more hydrophilic | + | To the outer end the protein has a helix on the N-terminal side shielding off the core and the C-terminus the LRR disappears slowly turn by turn with more and more hydrophilic amino acids replacing the LRR consensus sequence. |
Revision as of 21:32, 20 September 2011
 "
"
 Contact
Contact