Team:LMU-Munich/Project/Description
From 2011.igem.org
(→Project Details) |
|||
| Line 5: | Line 5: | ||
== Project Details== | == Project Details== | ||
| - | |||
This year’s project by the iGEM-team from Ludwig-Maximilians-University in Munich uses natural biosensors to detect the concentration of different metals. We have the vision to develop a set of bacterial metal sensors for easy qualitative and quantitative measurement of toxic metals just by reading the output after adding the water test sample. | This year’s project by the iGEM-team from Ludwig-Maximilians-University in Munich uses natural biosensors to detect the concentration of different metals. We have the vision to develop a set of bacterial metal sensors for easy qualitative and quantitative measurement of toxic metals just by reading the output after adding the water test sample. | ||
| Line 49: | Line 48: | ||
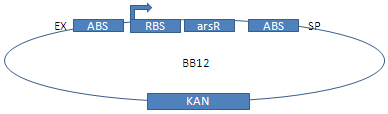
[[Image: Bild-pars.png | thumb |left]]'''pars''': The pars-promoter from Escherichia coli is regulated by the repressor arsR, which can bind As(III)- and Sb(III)-ions. Without these ions, the arsR binds to the pars-promoter, deactivating it. In the presence of As(III) or Sb(III), the repressor leaves the DNA, by this means enabling the promoter to work. For less leakiness of the repressed promoter, a second arsR-binding site was used. | [[Image: Bild-pars.png | thumb |left]]'''pars''': The pars-promoter from Escherichia coli is regulated by the repressor arsR, which can bind As(III)- and Sb(III)-ions. Without these ions, the arsR binds to the pars-promoter, deactivating it. In the presence of As(III) or Sb(III), the repressor leaves the DNA, by this means enabling the promoter to work. For less leakiness of the repressed promoter, a second arsR-binding site was used. | ||
| - | Since the regulator arsR does not exist in E. coli, it is coded on our plasmids. The pars-promoter with two binding sites has been multiplied via PCR with Phusion polymerase and primers E and F at an annealing temperature of 50°C. arsR with a constantly active promoter was | + | Since the regulator arsR does not exist in E. coli, it is coded on our plasmids. The pars-promoter with two binding sites has been multiplied via PCR with Phusion polymerase and primers E and F at an annealing temperature of 50°C. arsR with a constantly active promoter was fused using 3A-assembly.<br> |
| + | <br> | ||
| + | |||
| + | |||
| + | |||
| + | '''pnorB''': The fur-norB-System from Neisseria meningitidis is able to detect iron. Here fur is aktivating the norB-promoter if it has Fe-ions bound. Therefore the fur gene is used with a constitutive promoter and the inducible norB-promoter is fused with the reporter genes. The cloning details can be found in our [https://2011.igem.org/Team:LMU-Munich/Lab_Notebook lab notebook]. | ||
| + | <br>source(s):<br> | ||
| + | -[http://www.ncbi.nlm.nih.gov/pubmed/15130126 Fur functions as an activator and as a repressor of putative virulence genes in Neisseria meningitidis] Delany I, Rappuoli R, Scarlato V. | ||
| + | |||
| Line 60: | Line 67: | ||
The promoter pcueR from Bacillus subtilis controlled by the negative regulator CueR. CueR is capable of binding Cu(II)- and Cu(III)-ions by changing its conformation. In the absence of Cu (II/III)- ions cueR blocks the promoter region of pcueR by binding to the cueR binding site. If Cu - ions are available in the media, CueR releases the promoter region and transcription can be activated. For our project we fused the promotor region to a set of reporter genes in order to quantify the response of B. subtilis to high concentrations of copper ions. | The promoter pcueR from Bacillus subtilis controlled by the negative regulator CueR. CueR is capable of binding Cu(II)- and Cu(III)-ions by changing its conformation. In the absence of Cu (II/III)- ions cueR blocks the promoter region of pcueR by binding to the cueR binding site. If Cu - ions are available in the media, CueR releases the promoter region and transcription can be activated. For our project we fused the promotor region to a set of reporter genes in order to quantify the response of B. subtilis to high concentrations of copper ions. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| Line 82: | Line 85: | ||
| - | |||
| - | |||
| - | |||
Revision as of 17:59, 20 September 2011
Project Details
This year’s project by the iGEM-team from Ludwig-Maximilians-University in Munich uses natural biosensors to detect the concentration of different metals. We have the vision to develop a set of bacterial metal sensors for easy qualitative and quantitative measurement of toxic metals just by reading the output after adding the water test sample.
We use two different kinds of metal sensors. The ones in the first category work with reporter genes that lay downstream of an inducible promoter. The respective promoter is activated or deactivated by a specific metal-sensitive protein which binds to DNA dependent on the presence of that metal. As a consequence of this statistical event, there is a concentration-dependent transcription of the reporter gene, which is either GFP, luxAB or lacZ´.
The second kind of metal sensors directly uses the characteristics of special proteins to obtain a measurement of the metal, e.g. by analyzing the activity of an enzyme that needs a special metal ion as a cofactor.
For easy handling and compatibility we use the E. coli strain DH5α and B. subtilis for our two promoters taken from this organsim.
Reporter genes
Promoter-based Sensors
All genes are transcriptionally fused to the reporter gene, which means that approximately 50 nucleotides from the originally transcribed gene still remain with the promoter to ensure correct read-off. Downstream of this short open reading frame, the reporter gene with its own ribosome binding site is added.
source(s):
- [http://www.ncbi.nlm.nih.gov/pubmed?term=coordinating%20intracellular%20chivers Coordinating intracellular nickel-metal-site structure-function relationships and the NikR and RcnR repressors] Iwig JS, Chivers PT
- [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1236639/?tool=pubmed Complex transcriptional control links NikABCDE-dependent nickel transport with hydrogenase expression in Escherichia coli] RoweJS, Starnes GL, Chivers PT
- [http://www.ncbi.nlm.nih.gov/pubmed/10787413 Regulation of high affinity nickel uptake in bacteria] Ni2+-Dependent interaction of NikR with wild-type and mutant operator sites. Chivers PT, Sauer RT
- [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC93426/?tool=pubmed Isolation and characterization of the nikR gene encoding a nickel-responsive regulator in Escherichia coli] de Pina K, Desjardin V, Mandrand-Berthelot MA, Giordano G, Wu LF
source(s):
- [http://onlinelibrary.wiley.com/doi/10.1111/j.1365-2958.2006.05369.x/pdf Nickel homeostasis in Escherichia coli – the rcnR-rcnA efflux pathway and its linkage to NikR function] Iwig JS, Rowe JL, Chivers PT
- [http://www.ncbi.nlm.nih.gov/pubmed?term=coordinating%20intracellular%20chivers Coordinating intracellular nickel-metal-site structure-function relationships and the NikR and RcnR repressors] Iwig JS, Chivers PT
Since the regulator arsR does not exist in E. coli, it is coded on our plasmids. The pars-promoter with two binding sites has been multiplied via PCR with Phusion polymerase and primers E and F at an annealing temperature of 50°C. arsR with a constantly active promoter was fused using 3A-assembly.
pnorB: The fur-norB-System from Neisseria meningitidis is able to detect iron. Here fur is aktivating the norB-promoter if it has Fe-ions bound. Therefore the fur gene is used with a constitutive promoter and the inducible norB-promoter is fused with the reporter genes. The cloning details can be found in our lab notebook.
source(s):
-[http://www.ncbi.nlm.nih.gov/pubmed/15130126 Fur functions as an activator and as a repressor of putative virulence genes in Neisseria meningitidis] Delany I, Rappuoli R, Scarlato V.
pAseR: The paseR-promoter from Bacillus subtilis is regulated by the repressor AseR, which can bind As(II)- and As(III)-ions. In presence of these ions the repressor is releaved from the aseR binding site next to the promoter PaseR. The promoter region was amplified from genomic DNA of Bacillus subtilis subtilis W168, which is used as a laboratory wildtype of the gram-positive model organism. For the amplification the Primers XXXX and XXXX where used in a phusion PCR according to the standard PCR protocol displayed on this wiki. In our project we fused the promoter regions upstream of a set of reporter genes (lacZ', GFP and LuxAB) in order to measure its activity in the presence of As-ions.
pcueR:
The promoter pcueR from Bacillus subtilis controlled by the negative regulator CueR. CueR is capable of binding Cu(II)- and Cu(III)-ions by changing its conformation. In the absence of Cu (II/III)- ions cueR blocks the promoter region of pcueR by binding to the cueR binding site. If Cu - ions are available in the media, CueR releases the promoter region and transcription can be activated. For our project we fused the promotor region to a set of reporter genes in order to quantify the response of B. subtilis to high concentrations of copper ions.
Protein-based Sensors
These sensors are fused to his-tag or strep-tag domains to enable detection via antibodies as well.
pabA: pabA is coding for the glutamine aminotransferase that is an subunit of the PABA-synthase (4-Aminobenzoic acid). The synthase catalizes the reaction from chorismate and glutamate to PABA. PABA can be easily oxidized with Fe(III) and is so turning to an yellow-reddish color when adding oxygen.
The Experiments
working BioBrick erläutern
Results
welche BioBricks sind fertig und funktionieren hier reinschreiben!!!
 "
"