Team:Freiburg/Notebook/25 July
From 2011.igem.org
(Difference between revisions)
Juliimapril (Talk | contribs) (→NAME OF YOUR EXPERIMENT) |
|||
| (20 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | {{:Team:Freiburg/Templates/header}} | ||
| + | <html> | ||
| + | <div id="notebook-page-header"> | ||
| + | <div id="notebook-back" width="100px" > | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/23_July">Previous entry</a> | ||
| + | </div> | ||
| + | <div id="notebook-title"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook"> 25 July </a> | ||
| + | </div> | ||
| + | <div id="notebook-next"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/26_July">Next entry</a> | ||
| + | </div> | ||
| + | </div> | ||
| + | </html> | ||
| + | |||
| + | ==Commons== | ||
'''PCR''' | '''PCR''' | ||
| Line 76: | Line 92: | ||
| - | + | Labeled PSB 1 A3, PSB 1 C3, PSB 1 K3 and PSB 1 T3 stored in -20°C in last drawer | |
| - | |||
| - | |||
| - | |||
| - | |||
==<span style="color:green;">green light receptor</span>== | ==<span style="color:green;">green light receptor</span>== | ||
| - | ===Testdigest Ligation of CcaS into pSB1K3=== | + | ===Testdigest: Ligation of CcaS into pSB1K3=== |
'''Investigators:JULIA''' | '''Investigators:JULIA''' | ||
| Line 151: | Line 163: | ||
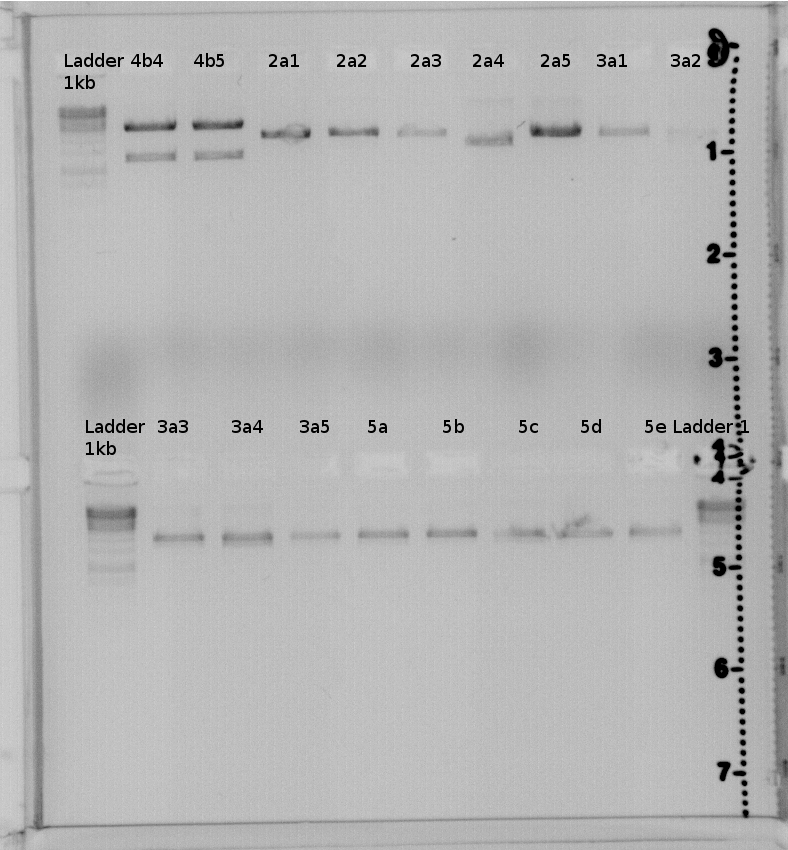
Picture of 1% gel: | Picture of 1% gel: | ||
| + | [[File:Freiburg11 07 25Testverdau Gel2 CcaS.Jpg|400px|400px]] | ||
| + | [[File:Freiburg11 7 26 Testverdau CcaS PR Gel1.Jpg|400px|400px]] | ||
| + | |||
| + | result:4a2,4b2,4b3,4b4,4b5 should have the right insert. | ||
| + | Send 4a2 and 4b2 to GATC for sequencing. | ||
==<span style="color:blue;">blue light receptor</span>== | ==<span style="color:blue;">blue light receptor</span>== | ||
| - | === | + | ===Gibson Not-Gate:Transformation=== |
| - | : | + | |
| + | |||
| + | '''Investigators: Sophie''' | ||
| - | |||
| - | |||
| - | |||
| Line 266: | Line 282: | ||
'''What program do you use? ''' | '''What program do you use? ''' | ||
| - | + | "57°C auf 70°C" (first annealing temperature:55°c, after 10 cycles 65°c) | |
| Line 277: | Line 293: | ||
I will do Gibson assemblz of the two parts next. | I will do Gibson assemblz of the two parts next. | ||
| + | <br/> | ||
| - | |||
| - | |||
| - | ''' | + | '''DNA-concentration measured with nanodrop:'''<br/> |
| + | {| cellpadding="10" cellspacing="0" border="1" | ||
| + | |sample | ||
| + | |DNA-concentration (ng/μl) | ||
| + | |- | ||
| + | |S35 | ||
| + | |75.3 | ||
| + | |- | ||
| + | |S45 | ||
| + | |27.2 | ||
| + | |} | ||
| + | [[File:Gimp_7_27_2011_Sophie.jpg]] | ||
| + | We will repeat the PCr, because of the bad result of S35. We will do a PCR with higher temperature. | ||
| - | ==<span style="color: | + | ==<span style="color:grey;">Precipitator</span>== |
| + | '''Digestion''' | ||
| - | |||
| - | + | {| style="border-spacing:0;" | |
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Ruediger | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 25.07 | ||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Date Name | ||
| + | Experiment | ||
| - | = | + | |- |
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: | ||
| - | + | |} | |
| + | Procedure | ||
| - | ''' | + | |
| + | # add H<sub>2</sub>O (38μl-DNA ) | ||
| + | # 5 μl NEB4 buffer (stored at iGEM’s, -20°C) | ||
| + | # 5 μl 10x BSA (used 1:10 diluted sample stored at iGEM’s, -20°C) | ||
| + | # DNA (500 ng) (Vector:Insert ratio 1:3 in following Ligation) | ||
| + | # 1 μl restriction enzymes (stored at iGEM’s, -20°C) | ||
| + | # heat for 1-2 hours 37°C (6 hours if time) | ||
| + | # heat for 20 minutes 80°C (inactivation of enzymes) | ||
| + | # keep at 4°C if you cannot continue | ||
| + | |||
| + | Measured DNA-concentration with Nanodrop to calculate the volume of DNA to do the digestion: | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Sample Name | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| DNA concentration (μg/μl) | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| P20 GFP | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 132 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| P19 GFP | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 145 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| P18 GFP | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 107 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| S39 PR | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 90 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| S43 PR | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 40 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Tet Vector | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 25 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |} | ||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| '''Components''' | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''Vector (μl)'''</center> | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''S39'''</center> | ||
| + | |||
| + | <center>'''Insert1: PR '''</center> | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''S43'''</center> | ||
| + | |||
| + | <center>'''Insert1: PR '''</center> | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''P18'''</center> | ||
| + | |||
| + | <center>'''Insert2: GFP+Pbd'''</center> | ||
| + | | style="border-top:0.0007in solid #000000;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''P19'''</center> | ||
| + | |||
| + | <center>'''Insert2: GFP+Pbd'''</center> | ||
| + | | style="border:0.0007in solid #000000;padding:0.0382in;"| <center>'''P20'''</center> | ||
| + | |||
| + | <center>'''Insert2: GFP+Pbd'''</center> | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| DNA (500ng) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 20 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''5.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''12.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''4.7'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''3.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| <center>'''3.8'''</center> | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| BSA (10x) (5μl) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| NEB4 Buffer (5μl) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| Enzyme 1 (1μl) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| E | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''E'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''E'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''X'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''X'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| <center>'''X'''</center> | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| Enzyme 2 (1μl) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| P | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''S'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''S'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''P'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''P'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| <center>'''P'''</center> | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| H<sub>2</sub>O (38 μl- DNA) | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| 18 | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''32.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''25.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''33.3'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| <center>'''34.5'''</center> | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| <center>'''34.2'''</center> | ||
| + | |||
| + | |- | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| In total 50 μl | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:none;padding:0.0382in;"| | ||
| + | | style="border-top:none;border-bottom:0.0007in solid #000000;border-left:0.0007in solid #000000;border-right:0.0007in solid #000000;padding:0.0382in;"| | ||
| + | |||
| + | |} | ||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Where are the samples stored? Antibiotica resistance, vector used etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 3A Assembly of linearized empty Tet Vector Psb1T3, PR Parts S39, S43 (CM resistance) and GFP Pbd from PCR with P18,19,20 | ||
| + | |||
| + | I did not digest linearized PsB1T3 with Dpn. | ||
| + | |||
| + | |} | ||
| + | Run a gel to verify if the part is cut out. | ||
Latest revision as of 01:00, 22 September 2011
 "
"
 Contact
Contact