Team:Potsdam Bioware/BioBricks
From 2011.igem.org
UP Sebastian (Talk | contribs) (→BioBrick ssTorA_CS-TEV_blaFL) |
UP Sebastian (Talk | contribs) (→pSB1A3 Ara and pSB1A3 lac) |
||
| (167 intermediate revisions not shown) | |||
| Line 66: | Line 66: | ||
|- | |- | ||
| - | | BBa_K627014 || A3_Ara_YFP | + | | BBa_K627014 || A3_Ara_YFP || 15 |
|- | |- | ||
| - | | BBa_K627015 || A3_lac_YFP | + | | BBa_K627015 || A3_lac_YFP || 16 |
| + | |- | ||
| + | |||
| + | | BBa_K627016 || TEV protease || 17 | ||
| + | |- | ||
| + | |||
| + | | BBa_K627017 || HRV14 3C protease || 18 | ||
|} | |} | ||
<br> | <br> | ||
| Line 99: | Line 105: | ||
Forward primer: TAAATGAATTCGCGGCCGCTTCTAGATGCCTCAATATACTACTAAAC<br> | Forward primer: TAAATGAATTCGCGGCCGCTTCTAGATGCCTCAATATACTACTAAAC<br> | ||
Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTATCAGCAAACCCTACTTAATTTC<br> | Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTATCAGCAAACCCTACTTAATTTC<br> | ||
| - | To insert mdnED in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes | + | To insert mdnED in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.<br> |
<br> | <br> | ||
| - | Because this BioBrick is an expression part, the | + | Because this BioBrick is an expression part, the adenine of mdnE gene's start codon is part of the XbaI |
recognition site.<br> | recognition site.<br> | ||
<br> | <br> | ||
| Line 107: | Line 113: | ||
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | ||
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | ||
| - | <b> | + | <b>Genbank file:</b><br> |
[[Media: UP_BioBrick_mdnED.gb]] | [[Media: UP_BioBrick_mdnED.gb]] | ||
<br> | <br> | ||
| Line 115: | Line 121: | ||
<b>Part name:</b> BBa_K627001<br> | <b>Part name:</b> BBa_K627001<br> | ||
<b>Part type:</b> Coding<br> | <b>Part type:</b> Coding<br> | ||
| - | <b>Short description:</b> | + | <b>Short description:</b> mdnA gene encoding the precursor peptide of the tricyclic microviridin<br> |
<br> | <br> | ||
<b>Full description:</b><br> | <b>Full description:</b><br> | ||
| Line 124: | Line 130: | ||
The following BioBrick mdnA encodes the ribosomal precursor peptide (MdnA), which is essential for microviridin production (Ziemert et al., 2008). | The following BioBrick mdnA encodes the ribosomal precursor peptide (MdnA), which is essential for microviridin production (Ziemert et al., 2008). | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RFC10 expression part, the | + | Because this BioBrick is a RFC10 expression part, the adenine of mdnB gene`s start codon is part of the XbaI recognition site. |
| + | <br><br> | ||
| + | <b>Usage and Biology:</b><br> | ||
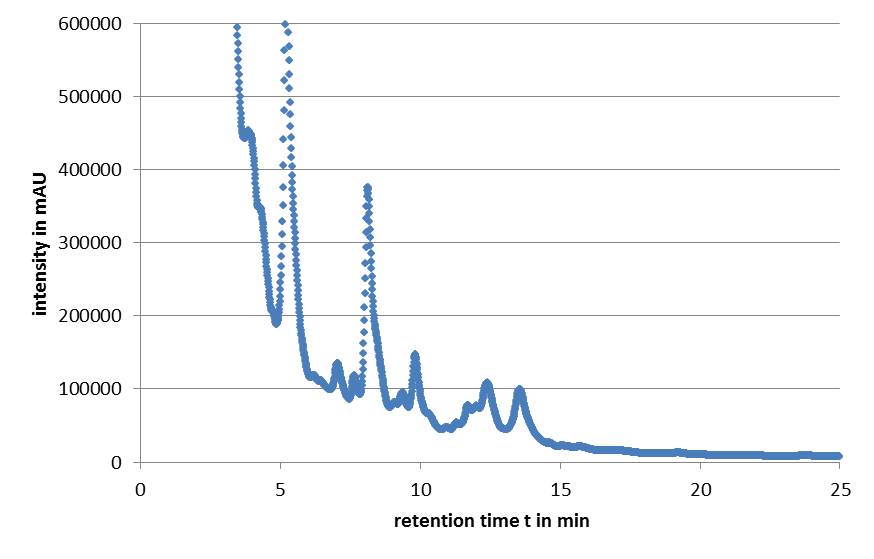
| + | Microviridin production in ''E. coli'' cells expressing the mdn genes was monitored by reverse phase HPLC. Qualitative analysis involves running a standard that contains the target analytes. The vectors pARW071 and pARW089 served as a control in our experiments because these vectors contain the original mdn-cluster. We could use the retention time as a way to determine the presence of the microviridin production in other samples. HPLC analysis of the purified compound yielded a high peak with a retention time of approximately 5 min. Minor peaks could be detected during the following 7 min (Fig. 1). | ||
| + | [[File:UP_modularization_HPLC.png|center|500px|thumb|'''Figure 1:''' HPLC chromatogram of mdnA]] | ||
| + | All HPLC chromatograms of the isolated mdnA showed reliable peaks with a retention time of 5 min together with a number of following minor peaks. | ||
| + | |||
| + | Fractions of the peaks were sampled and the identity of microviridin and also the presence of cyclization, which is important for the activity, was affirmed with mass spectrometry (Fig. 2). By using a mass spectrometer it is possible to determine both the elemental composition of a fraction and the chemical structure of molecules. | ||
| + | |||
| + | [[File:UP_modularization_MSII.gif|center|300px|thumb|'''Figure 2:''' Mass Spectrometry Data]] | ||
<br><br> | <br><br> | ||
<b>Source of the part:</b><br> | <b>Source of the part:</b><br> | ||
| Line 135: | Line 150: | ||
To insert mdnA in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | To insert mdnA in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is an expression part, the | + | Because this BioBrick is an expression part, the adenine of mdnA gene`s start codon is part of the XbaI recognition site. Further the sequence contains a AgeI recognition site after mdnA. |
<br><br> | <br><br> | ||
<b>Experiences:</b> | <b>Experiences:</b> | ||
<br><br> | <br><br> | ||
| - | <b>References:</b> | + | <b>References:</b><br> |
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9 | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9 | ||
| - | <br> | + | <br><br> |
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74 | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74 | ||
<br><br> | <br><br> | ||
| Line 161: | Line 176: | ||
The following BioBrick mdnB encodes a ATP-grasp-type ligase (Ziemert et al., 2008). | The following BioBrick mdnB encodes a ATP-grasp-type ligase (Ziemert et al., 2008). | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is an expression part, the | + | Because this BioBrick is an expression part, the adenine of mdnB gene`s start codon is part of the XbaI recognition site. |
<br><br> | <br><br> | ||
<b>Source of the part:</b><br> | <b>Source of the part:</b><br> | ||
| Line 172: | Line 187: | ||
To insert mdnB in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | To insert mdnB in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is an expression part, the | + | Because this BioBrick is an expression part, the adenine of mdnE gene's start codon is part of the XbaI recognition site. |
<br><br> | <br><br> | ||
| - | <b>References:</b> | + | <b>References:</b><br> |
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9 | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9 | ||
| - | <br> | + | <br><br> |
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74 | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74 | ||
<br><br> | <br><br> | ||
| Line 202: | Line 217: | ||
The following BioBrick mdnC encodes the ATP-grasp-type ligase (Ziemert et al., 2008). | The following BioBrick mdnC encodes the ATP-grasp-type ligase (Ziemert et al., 2008). | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RF10 expression part, the adenine of mdnC gene start codon is part of the XbaI | + | Because this BioBrick is a RF10 expression part, the adenine of mdnC gene`s start codon is part of the XbaI |
recognition site.<br> | recognition site.<br> | ||
<br> | <br> | ||
| Line 214: | Line 229: | ||
To insert mdnC in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | To insert mdnC in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RF10 expression part, the adenine of mdnA gene start codon is part of the XbaI recognition site. | + | Because this BioBrick is a RF10 expression part, the adenine of mdnA gene`s start codon is part of the XbaI recognition site. |
<br><br> | <br><br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | ||
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | ||
| - | <b> | + | <b>Genbank file:</b><br> |
[[Media: UP_BioBrick_mdnC.gb]] | [[Media: UP_BioBrick_mdnC.gb]] | ||
<br> | <br> | ||
| Line 254: | Line 269: | ||
To insert mdnD in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | To insert mdnD in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RF10 expression part the adenine of mdnD gene start codon is part of the XbaI recognition site. | + | Because this BioBrick is a RF10 expression part the adenine of mdnD gene`s start codon is part of the XbaI recognition site. |
<br><br> | <br><br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | ||
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | ||
| - | <b> | + | <b>Genbank file</b><br> |
[[Media: UP_BioBrick_mdnD.gb]] | [[Media: UP_BioBrick_mdnD.gb]] | ||
<br><br> | <br><br> | ||
| Line 282: | Line 297: | ||
The following BioBrick mdnE encodes an ABC transporter (Ziemert et al., 2008). | The following BioBrick mdnE encodes an ABC transporter (Ziemert et al., 2008). | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RF10 expression part the adenine of mdnE gene start codon is part of the XbaI | + | Because this BioBrick is a RF10 expression part the adenine of mdnE gene`s start codon is part of the XbaI |
recognition site.<br> | recognition site.<br> | ||
<br> | <br> | ||
| Line 294: | Line 309: | ||
To insert mdnE in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | To insert mdnE in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI. | ||
<br><br> | <br><br> | ||
| - | Because this BioBrick is a RF10 expression part the adenine of mdnE gene start codon is part of the XbaI recognition site. | + | Because this BioBrick is a RF10 expression part the adenine of mdnE gene`s start codon is part of the XbaI recognition site. |
<br><br> | <br><br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9<br><br> | ||
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74<br><br> | ||
| - | <b> | + | <b>Genbank file</b><br> |
[[Media: UP_BioBrick_mdnE.gb]] | [[Media: UP_BioBrick_mdnE.gb]] | ||
<br><br> | <br><br> | ||
| - | === BioBrick mdnA | + | === BioBrick mdnA-myc-gene III === |
<b>Part name:</b> BBa_K627006<br> | <b>Part name:</b> BBa_K627006<br> | ||
<b>Part type:</b> Coding<br> | <b>Part type:</b> Coding<br> | ||
| - | <b>Short description:</b> Fusion part of mdnA gene | + | <b>Short description:</b> Fusion part of mdnA gene (from mdn-cluster) with myc-tag and gene III<br> |
<br> | <br> | ||
| - | < | + | <br>'''Full description'''<br> |
| - | + | <br> | |
| + | This BioBrick is important for testing the suitability of a phage display system as an appropriate screening method for recombinant mdnA libraries. | ||
| + | The mdnA gene encodes the precursor peptide from the mdn-cluster, which is an essential building block for production of the protease inhibitor microviridin from Microcystis aeruginosa (Ziemert et al., 2008 and 2010). This peptide is known to be able to bind and inhibit proteases. Thus, variants of mdnA have a high potential for therapy as they can block disease-relevant proteases. | ||
| + | The gene III protein is a coat protein from the filamentous bacteriophage M13 (Smith, 1985). It appears only three to five times on the tip of the phage and is responsible for infection of bacterial cells. In phage display, genes from a DNA library are usually fused to the gene III and are displayed as fusion protein on the surface of the phages. Because it has been shown that the proteins of interest, which are fused to the C-terminus of gene-III can be functionally displayed (Krebber et al. 1997, Fuh and Sidhu, 2000, Rakonjac et al., 1999) only the sequence for this part has to be used. | ||
| + | The inserted sequence of the myc-tag enables easy detection and/or purification of the proteins of interest. In our case, the tag was used to verify display of the microviridin on the phage particle. | ||
| + | After transformation of E. coli with a vector containing the mdnA-myc-geneIII-fusion gene (in the context of the mdnA gene cluster) and co-infection with helper phages, E. coli cells were able to produce phage particles, which carry the microviridin peptide on their surface as confirmed by ELISA. These experiments proved the suitability of phage display as a screening method for variants of microviridin. | ||
<br><br> | <br><br> | ||
| + | <br>'''Control of expression of mdnA-myc-gene III in ''E. coli'' ''' | ||
| + | |||
| + | To control expression of the mdnA-myc-gene III fusion gene was analyzed by western blotting. E. coli cells transformed with the phagemid pPDV089 were harvested and lysated. The cell proteins were electrophoretically separated and transferred to a membrane. The mdnA-myc-gene III-fusion proteins were detected using specific anti-myc-antibodies and horseradish peroxidase (HRP)-linked antibodies as second antibodies. Enhanced chemiluminescence (ECL) was used to visualize the protein. ECL is based on the emission of light during the HRP -catalyzed oxidation of luminol, which was captured by a camera. The western blot analysis resulted in a band of a size just below the 30 kDa mark representing the mdnA-myc-gene III-fusion protein (24 kDa). | ||
| + | |||
| + | [[File:UP_coomassie+western blot.png|center|400px|thumb|'''Figure 3: Control of expression of mdnA-myc-gene III in ''E. coli'' by western blotting.''' For detection anti-myc-antibodies and secondary HRP-linked antibodies were used. The resulting band represents the mdnA-myc-gene III-fusion protein.]] | ||
| + | <br>'''Detection of phages carrying mdnA on their surface by ELISA'''<br> | ||
| + | <br> | ||
| + | The next step was the detection of the expression of the mdnA-myc-gene III-fusion gene on the surface of the phage. So ''E. coli'' cells strain XL1-Blue were first transformed with the phagemid pPDV089 before they were infected with helper phages. ''E. coli'' cells containing both plasmids were selected. An ELISA test was performed to determine whether these cells are able to produce phage particles carrying the mdnA peptide on their surface. To perform this test anti-c-myc-antibodies were immobilized on ELISA plates and incubated with purified phages. For detection a second antibody coupled with horse radish peroxidase (HRP) was used which binds the gene VIII protein of the phages. The HRP substrate o-phenyldiamine (OPD) was added and in case of binding a color reaction was expected. The color shift from achromatic to yellow in wells incubated with phages produced in XL1-Blue cells showed the successful expression of mdnA-c-myc-gene III-fusion protein on the phages.<br> | ||
| + | For more precise results the absorption at 492 nm was measured. The data were presented in a bar plot. As a negative control helper phages were added instead of produced phages. Furthermore two wells were prepared were the secondary antibody was not added. | ||
| + | The graphic shows clearly the much higher absorption measured in wells, which were incubated with phage particles of interest produced in XL1-Blue cells. As has already pointed out this shows the succeeded expression of mdnA-c-myc-gene III-fusion protein on the surface of the phages. | ||
| + | |||
| + | |||
| + | <div align="center"> | ||
| + | [[File:UP ELISA3 mit fehlerindikator.png|center|400px|thumb|'''Figure 4: Detection of phages carrying mdnA on their surface by ELISA.''' The bar plot shows the absorption at 492 nm. Anti-myc-antibodies were immobilized. For detection a second antibody coupled with horse radish peroxidase (HRP) was used which binds the gene VIII coat protein of the phages. The left bar shows the absorption of the wells containing helper phages (negative control), the right bar shows the absorption of wells containing mdnA carrying phages]]</div><br> | ||
| + | <br> | ||
| + | <br>'''Testing phage display with unmodified mdnA to examine its suitability as screening method '''<br> | ||
| + | <br> | ||
| + | To test the suitability of this screening method, phages representing unmodified mdnA on their surface and phages not representing mdnA (helper phages) in a ratio of one to one were incubated with immobilized trypsin which is known as a target of mdnA. The display was conducted in ELISA plates. The bound phages were eluted using a buffer with low pH value and neutralized afterwards. To check how many phages interacted with trypsin, ''E. coli'' cells XL1-Blue were re-infected with eluted phages and plated on agar with different antibiotics. Cells infected with phages carrying mdnA are able to grow on agar with ampicillin whereas cells infected with helper phages are able to grow on agar with kanamycin. To control the success of the panning round additionally ''E. coli'' cells were infected with phage mix before panning and plated on agar. Subsequent the number of clones grew on ampicillin and kanamycin before and after panning was compared. During the running of this step it was noticed that much more cells were infected with helper phages than with phages carrying mdnA despite of the engaged 1:1 ratio. This was surprising and indicated that mdnA on the surface of the phages may inhibit their infectivity. After controlling the plates an infection ratio of phages carrying mdnA to helper phages of 1:400 was calculated. This fact should be analyzed in further experiments.<br> | ||
| + | The results of the first phage display are plotted in the figure below. After one panning round an enrichment of phages carrying mdnA was expected. This is attributable to the fact that phage particles carrying mdnA c-myc geneIII fusion protein on their surface are expected to bind specifically to the immobilized trypsin. Unfortunately this was not observed in this experiment. The ratio of cells growing on kanamycin agar before (4000) to cells growing on kanamycin agar (cells containing helper phages) after panning (750) was determined as 5:1. The ratio of cells growing on ampicillin agar before (12) to cells growing on ampicillin agar (cells containing mdnA carrying phages) after panning (2) was nearly equal. Thus no enrichment of mdnA carrying phages occurred in the first experiment. So it was decided to repeat this experiment under improved conditions. Therefor the number of washing steps during the described experimental procedure was increased. Here the ratio of cells growing on kanamycin agar before (3000) to cells growing on kanamycin agar (cells containing helper phages) after panning (29) was determined as 103:1. The ratio of cells growing on ampicillin agar before (26) to cells growing on ampicillin agar (cells containing mdnA carrying phages) after panning (2) was determined as 13:1. Thus an enrichment factor of eight was reached for the phages displaying mdnA on their surface. <br> | ||
| + | These results indicate that the unmodified mdnA expressed on the phages binds specifically to the immobilized trypsin. Therefore it can be deduced that mdnA is presented in a functional 3D structure. These findings suggest that phage display in general is an appropriate method for screening a recombinant mdnA library. Further experiments are required to optimize this system. | ||
| + | |||
| + | |||
| + | [[Image:UP panning 3.png|center|400px|thumb|'''Figure 5: Optimization of phage display.''' After optimized conditions (right) a clear concentration of phages carrying mdnA after one panning round was noted. ''E. coli'' cells were infected with phage mix (helper phages and phages carrying mdnA) before and after panning and plated on agar containing kanamycin or ampicillin. The ratio of cells growing on ampicillin or kanamycin agar before panning to cells growing on ampicillin or kanamycin agar after was calculated. Helper phages which acted as negative control have a kanamycine resistance whereby phages carrying mdnA have an ampicillin resistance.]] | ||
| + | <br><br> | ||
| + | |||
| + | <br>'''Testing phage display with unmodified mdnA against further proteases''' | ||
| + | <br> | ||
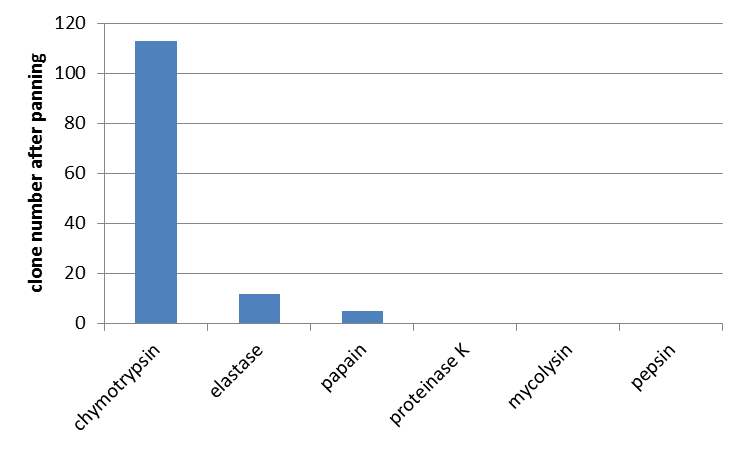
| + | Furthermore the interaction of unmodified mdnA with other proteases was determined. From the literature (Ziemert, 2010) the high inhibitory activity of microviridin L, besides trypsin, against chymotrypsin and elastase is also known. So a phage display with these enzymes was performed. Additionally papain, proteinase K, mycolysin and pepsin were tested for which no interaction was shown yet. For all enzymes an equal amount of E. coli cells and phages were used. In agreement with data from the literature interaction of microviridin with chymotrypsin and elastase was confirmed. Additionally an interaction with papain was noted. All other proteases were not bound by microviridin. | ||
| + | |||
| + | [[File:UP_test of different enzymes.png|center|400px|thumb|'''Figure 6: Phage display against different proteases.''' After panning the number of clones was counted. In agreement with data from the literature interaction of microviridin with chymotrypsin and elastase was confirmed. Additionally an interaction with papain was noted.]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
<b>Source of the part:</b><br> | <b>Source of the part:</b><br> | ||
The BioBrick mdnA is a part of the microviridin gene (mdn) cluster which was isolated from Microcystis aeruginosa strain NIES-843.<br> | The BioBrick mdnA is a part of the microviridin gene (mdn) cluster which was isolated from Microcystis aeruginosa strain NIES-843.<br> | ||
| Line 317: | Line 371: | ||
<br><br> | <br><br> | ||
<b>Design information:</b><br> | <b>Design information:</b><br> | ||
| - | Gene III was amplified from the vector | + | Gene III was amplified from the vector pak100blaKDIR using the following primers<br> |
Forward:TAAGCTTCTAGATGGCCGGCGAGCAGAAGCTGATCTCTGAGGAAGACCTGGGTGGTGGCTCTGGTTCC<br> | Forward:TAAGCTTCTAGATGGCCGGCGAGCAGAAGCTGATCTCTGAGGAAGACCTGGGTGGTGGCTCTGGTTCC<br> | ||
Reverse: TGCTTAGACGTCCTGCAGCGGCCGCTACTAGTATTAACCGGTAGACTCCTTATTACGCAGTA<br><br> | Reverse: TGCTTAGACGTCCTGCAGCGGCCGCTACTAGTATTAACCGGTAGACTCCTTATTACGCAGTA<br><br> | ||
| - | The mdnA gene was amplified from the vector pARW089 using the following | + | The mdnA gene was amplified from the vector pARW089 using the following primers<br> |
Forward: TTCCATGGCGCCAGAGGAATCTAGATGGCATATCCCAACGATC<br> | Forward: TTCCATGGCGCCAGAGGAATCTAGATGGCATATCCCAACGATC<br> | ||
Reverse: CTTCTGACTGGGAAGATTATACCGGTTAATACTAGTAGCGGCCGCTGCAGGACGTC<br> | Reverse: CTTCTGACTGGGAAGATTATACCGGTTAATACTAGTAGCGGCCGCTGCAGGACGTC<br> | ||
<br> | <br> | ||
| - | Gene III was amplified from pak100blaKDIR and mdnA from pARW089 by PCR. The primers were designed to enable the introduction of iGEM. The PCR product gene III was digested by whereas the PCR product mdnA was digested by AgeI. Thus a mdnA-gene III fusion part according to RFC25 was generated whereby AgeI and NgoMIV overhangs are compatible and placed in frame with the protein sequence. The ligation of AgeI and NgoMIV overhangs resulted in a scar coding for the threonine and glycine. Because the introduction of restriction sites before mdnA leaded to a great distance between ribosome binding site and mdnA a second RBS was inserted among SfoI and XbaI recognition sites to ensure a sufficiently expression rate of the mdnA- | + | Gene III was amplified from pak100blaKDIR and mdnA from pARW089 by PCR. The primers were designed to enable the introduction of iGEM. The PCR product gene III was digested by whereas the PCR product mdnA was digested by AgeI. Thus a mdnA-gene III fusion part according to RFC25 was generated whereby AgeI and NgoMIV overhangs are compatible and placed in frame with the protein sequence. The ligation of AgeI and NgoMIV overhangs resulted in a scar coding for the threonine and glycine. Because the introduction of restriction sites before mdnA leaded to a great distance between ribosome binding site and mdnA a second RBS was inserted among SfoI and XbaI recognition sites to ensure a sufficiently expression rate of the mdnA-gene III-fusion gene. Between mdnA and gene III myc sequence was inserted. |
<br><br> | <br><br> | ||
| - | Because this BioBrick is an expression part, the | + | Because this BioBrick is an RFC25 expression part, the adenine of mdnA gene's start codon is part of the XbaI recognition site. Further the sequence contains a AgeI recognition site after gene III. |
<br><br> | <br><br> | ||
| - | <b> | + | <b>References:</b><br> |
| + | *Fuh G., Sidhu S.S. (2000). Efficient phage display of polypeptides fused to the carboxy-terminus of the M13 gene-3 minor coat protein. FEBS Lett. 480(2-3):231-4 | ||
| + | *Krebber, A., Bornhauser, S., Burmester, J., Honegger, A., Willuda, J., Bosshard, H. R., Plückthun, A. (1997) Reliable cloning of functional antibody variable domains from hybridomas and spleen cell repertoires employing a reengineered phage display system. J. Immunol. Meth. 201(1):35-55 | ||
| + | *Rakonjac J., Feng J., Model P. (1999). Filamentous phage are released from the bacterial membrane by a two-step mechanism involving a short C-terminal fragment of pIII. J Mol Biol. 289(5):1253-65 | ||
| + | *Smith, G.P. (1985). Filamentous fusion phage: Novel expression vectors that display cloned antigens on the virus surface. Science 228: 1315-17 | ||
| + | *Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9 | ||
| + | |||
| + | *Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74 | ||
<br><br> | <br><br> | ||
| - | <b> | + | <b>Genbank file:</b><br> |
| - | + | [[Media: UP_Biobrick_mdnA_c-myc_geneIII.gb]] | |
| + | <br><br> | ||
| + | |||
| + | === BioBrick myc-gene III === | ||
| + | |||
| + | <b>Part name:</b> BBa_K627007<br> | ||
| + | <b>Part type:</b> Coding<br> | ||
| + | <b>Short description:</b> Fusion of c-myc-tag and gene III <br> | ||
<br> | <br> | ||
| - | + | <b>Full description:</b><br> | |
| + | The gene III protein is a coat protein from the filamentous bacteriophage M13 (Smith, 1985). It appears only 3-5 times on the tip of the phage and is responsible for infection of bacterial cells. It is usually fused to the gene of interest to determine protein-protein-interactions of its expression product using M13 phage display system. The gene III-protein consists of three domains: one C-termial and two N-terminal domains. Gene III was generated as biobrick several times for instance M13003 (2006) and M31537 (2007). It has been shown that proteins of interest fused to the carboxy-terminus of gene-III-proteins can be functionally displayed on the surface of phages (Fuh and Sidhu, 2000, Rakonjac et al., 1999). In the course of our project a phagemid was constructed carrying truncated gene III (only encoding C-terminal domain) as biobrick. To enable easy purification and detection of gene-III-fused proteins additionally a myc-tag-sequence was merged before gene –III-sequence. | ||
| + | <br><br> | ||
| + | <b>Source of the part:</b><br> | ||
| + | The gene-III-protein is a coat protein from the filamentous bacteriophage M13. | ||
| + | <br><br> | ||
| + | <b>Design information:</b><br> | ||
| + | Gene III was amplified from the vector pak100blaKDIR using the following primers<br> | ||
| + | Forward:TAAGCTTCTAGATGGCCGGCGAGCAGAAGCTGATCTCTGAGGAAGACCTGGGTGGTGGCTCTGGTTCC<br> | ||
| + | Reverse: TGCTTAGACGTCCTGCAGCGGCCGCTACTAGTATTAACCGGTAGACTCCTTATTACGCAGTA<br><br> | ||
<br> | <br> | ||
| - | + | Because myc-gene III is a fusion part (RFC25), no start codon is needed. Before the myc sequence a NgoMIV restriction site is located which has compatible overhangs to AgeI restriction sites. The ligation of AgeI and NgoMIV overhangs will result in a scar coding for threonine and glycine. So genes of interest containing AgeI restriction sites can be easily fused to gene III without frame shift. The gene III sequence contains no stop codon because there is one located between AgeI and SpeI recognition site directly merged behind gene III sequence. | |
| + | <br><br> | ||
| + | <b>References:</b><br> | ||
| + | Fuh G., Sidhu S.S. (2000). Efficient phage display of polypeptides fused to the carboxy-terminus of the M13 gene-3 minor coat protein. FEBS Lett. 480(2-3):231-4 | ||
| + | |||
| + | Rakonjac J., Feng J., Model P. (1999). Filamentous phage are released from the bacterial membrane by a two-step mechanism involving a short C-terminal fragment of pIII. J Mol Biol. 289(5):1253-65 | ||
| + | |||
| + | Smith, G.P. (1985). Filamentous fusion phage: Novel expression vectors that display cloned antigens on the virus surface. Science 228: 1315-17 | ||
<br><br> | <br><br> | ||
<b>Genbank file:</b><br> | <b>Genbank file:</b><br> | ||
| - | [[Media: | + | [[Media: UP_Biobrick_myc_gene III.gb]] |
<br><br> | <br><br> | ||
| - | |||
| - | <b>Part name:</b> BBa_K627008<br> | + | === BioBrick AraC-TEV === |
| - | <b>Part type:</b> | + | |
| - | <b>Short description:</b> Fusion part of | + | <b>Part name:</b> BBa_K627008, BBa_K627009, BBa_K627010 <br> |
| + | <b>Part type:</b> Device<br> | ||
| + | <b>Short description:</b> Fusion part of arabinose-inducible induction system and the TEV protease<br> | ||
| + | <b>RFC:</b> 10, 12, 23<br> | ||
<br> | <br> | ||
<b>Full description:</b><br> | <b>Full description:</b><br> | ||
| - | This BioBrick is a 2 part fusion of | + | |
| - | At 37°C, the TEV protease | + | <b>Introduction</b><br> |
| - | <br> | + | This BioBrick is a 2 part fusion of arabinose-inducible induction system and the TEV protease.<br><br> |
| + | <b>General Information of the single sub parts:</b><br><br> | ||
| + | <b>TEV protease</b><br> | ||
| + | TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus (TEV). It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.<br> | ||
| + | At 37°C, the TEV protease forms inclusion body, which leads to an inactive form. Incubated at 30 °C, the protease is expressed as soluble type and is highly active.<br><br> | ||
| + | <b>Arabinose-inducible induction system</b><br> | ||
| + | The arabinose-inducible induction system was amplified via PCR from the pBAD_iGEMexpress vector. Based on the inhibitionary funtion of the catabolic activator protein (CAP) it got a very low expression rate without induction by arabinose. For the induction concentration between 2 mM up to 50 mM arabinose can be used to get an high expression rate.<br><br> | ||
| + | <b>General Function:</b><br> | ||
| + | When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the TEV protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.<br><br> | ||
| + | <b>Performance and Summary:</b><br> | ||
| + | In combination with our created ssTorA_CS-TEV_blaFL device (BBa_K627012, [http://partsregistry.org/Part:BBa_K627012 More Information]) this protease generator was able to mediate the cell death of our transformed <i>E.coli</i> cells at ampicillin concentration up to 100 µg/ml.<br><br> | ||
<b>Source of the part:</b><br> | <b>Source of the part:</b><br> | ||
| - | Arabinose inducible operon form pBAD_iGEM_express, TEV protease from Gunther Stier | + | Arabinose inducible operon form pBAD_iGEM_express, TEV protease from Gunther Stier et al.<br> |
<br> | <br> | ||
<b>Design Notes</b><br> | <b>Design Notes</b><br> | ||
| - | This | + | This BioBrick was built by PCR using the following PCR primers:<br> |
<br> | <br> | ||
<b>Primer used for site directed mutagenesis:</b><br> | <b>Primer used for site directed mutagenesis:</b><br> | ||
| Line 382: | Line 478: | ||
<br> | <br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
| - | + | Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S. P. (2007). Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16: 2360-2367 | |
| - | + | <br><br> | |
| - | + | Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D., and Waugh, D. S. (2001). Tobacco etch virus protease: Mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Prot. Eng. 14: 993-1000. | |
| + | <br><br> | ||
| + | Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable form of recombinant tobacco etch virus protease. Biotechniques 30: 544-550.<br><br> | ||
| + | <b>Genbank file</b><br> | ||
| + | [[Media:BioBrick_AraC-TEV_comp.gb]] | ||
| + | <br><br> | ||
| + | Three parts are existing. | ||
| - | === BioBrick | + | === BioBrick AraC-14_3C protease=== |
| - | <b>Part name:</b> | + | <b>Part name:</b> BBa_K627011<br> |
| - | <b>Part type:</b> | + | <b>Part type:</b>Device<br> |
| - | <b>Short description:</b> Fusion part of | + | <b>Short description:</b> Fusion part of arabinose-inducible induction system and the HRV 14_3C protease<br> |
| + | <b>RFC:</b> 10, 12, 23<br> | ||
<br> | <br> | ||
<b>Full description:</b><br> | <b>Full description:</b><br> | ||
| - | |||
| - | |||
<br> | <br> | ||
| - | <b> | + | <b>Introduction</b><br> |
| - | Arabinose inducible | + | This BioBrick is a 2 part fusion of arabinose-inducible induction system and the HRV 14_3C protease.<br><br> |
| + | <b>General Information of the single sub parts:</b><br><br> | ||
| + | <b>HRV 14_3C</b><br> | ||
| + | The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln/GlyPro.<br> | ||
| + | The 22 kDa protease got its optimal activity at 4°C but also shows a high cleavage rate at 37°C. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.<br><br> | ||
| + | <b>Arabinose-inducible induction system</b><br> | ||
| + | The arabinose-inducible induction system was amplified via PCR from the pBAD_iGEMexpress vector. Based on the inhibitionary funtion of the catabolic activator protein (CAP) it got a very low expression rate without induction by arabinose. For the induction concentration between 2 mM up to 50 mM arabinose can be used to get an high expression rate.<br><br> | ||
| + | <b>General Function:</b><br> | ||
| + | When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the HRV 14_3C protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.<br><br> | ||
| + | <b>Performance and Summary:</b><br> | ||
| + | In combination with our created ssTorA_CS-14_3C_blaFL device (BBa_K627013, [http://partsregistry.org/Part:BBa_K627013 More Information]) this protease generator was able to mediate the cell death of our transformed <i>E.coli</i> cells at ampicillin concentration up to 100 µg/ml.<br> | ||
<br> | <br> | ||
| - | <b> | + | <b>Source of the part:</b><br> |
| - | + | Arabinose inducible operon form pBAD_iGEM_express was ampilied via PCR, 14_3C protease was isolated from the genome of human rhinovirus 14_3C and mutated with primers listed in part description BBa: to remove all iGEM restriction sites inside the protease gene.<br> | |
<br> | <br> | ||
| - | <b> | + | <b>Design Notes:</b><br> |
| - | <i>Fragment 1:</i>< | + | This BioBrick was built by PCR using the following PCR primers:<br> |
| - | * | + | <b>Site directed mutagenesis of HRV 14 3C protease:</b><br> |
| - | * | + | <i>Fragment 1:</i> |
| - | <i>Fragment 2:</i>< | + | * f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC |
| - | * | + | * r_14_3C_ACCAGC: ACCATTTGCTGGTACATCATCACCAGG |
| - | * | + | <i>Fragment 2:</i> |
| + | * f_14_3C_ACCAGC: CCTGGTGATGATGTACCAGCAAATGGT | ||
| + | * r_14_3C_tm_XbaI208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC | ||
| + | <i>Fragment 3:</i> | ||
| + | * f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG | ||
| + | * r_14_3C_tm_XbaI280_A-T: ATCCACACCTTCGAGATCTTCTGATAT | ||
| + | <i>Fragment 4:</i> | ||
| + | * f_14_3C_tm_Xba280_A-T: ATATCAGAAGATCTCGAAGGTGTGGAT | ||
| + | * r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | ||
| + | <b>Primers used for first assembly PCR of mutated fragments:</b><br> | ||
| + | <i>Fragment 5, containing Fragment 1 and 2:</i> | ||
| + | * f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC | ||
| + | * r_14_3C_tm_Xba208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC | ||
| + | <i>Fragment 6, containing Fragment 3 and 4:</i> | ||
| + | * f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG | ||
| + | * r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | ||
| + | <b>Primer used for final assembly PCR of mutated fragments:</b><br> | ||
| + | <i>Complete mutated 14_3C protease, containing Fragment 5 and 6:</i> | ||
| + | * f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC | ||
| + | * r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | ||
| + | <i>Primers used for amplification of the pBAD arabinose-inducible induction system:</i> | ||
| + | * f_AraC_iGEM_HindIII: TATAAGCTTGAATTCGCGGCCGCTTCTAGATTATGACAACTTGACGGCTACATCATT | ||
| + | * r_AraC_NgoMIV: ATAGCCGGCCTCCTTCTTAAAGTTAAACAAAATTATTTCTAGCCC | ||
| + | <i>Primers used for amplification and modification of HRV 14 3C protease:</i> | ||
| + | * f_14_3C_AraFusion_NgoMIV: ATATTGCCGGCATGGGACCAAACACAGAATTTGCACTATCC | ||
| + | * r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | ||
<br> | <br> | ||
| - | <b> | + | <b>Experience:</b><br> |
| - | + | ||
| - | + | ||
<br> | <br> | ||
| - | <b> | + | <b>References:</b><br> |
| - | + | Combination of Two Separate Binding Domains Defines Stoichiometry between Type III Secretion System Chaperone IpgC and Translocator Protein IpaB, Ravi Kumar Lokareddy, Michele Lunelli, Björn Eilers, Vivien Wolter, and Michael Kolbe, December 17, 2010 The Journal of Biological Chemistry, 285, 39965-39975. | |
| - | + | <br><br> | |
| + | Cleavage of Small Peptides In Vitro by Human Rhinovirus 14 3C Protease expressed in Escherichia coli, MICHAEL G. CORDINGLEY,* R. BRUCE REGISTER, PIA L. CALLAHAN, VICTOR M. GARSKY,AND RICHARD J. COLONNO JOURNAL OF VIROLOGY, Dec. 1989, p. 5037-5045 | ||
| + | <br><br> | ||
| + | <b>Genbank file:</b><br> [[Media:BioBrick_AraC-14_3C.gb]] | ||
| + | |||
| + | === BioBrick ssTorA_CS-TEV_blaFL === | ||
| + | |||
| + | <b>Part name:</b> BBa_K627012<br> | ||
| + | <b>Part type:</b> Composite Part<br> | ||
| + | <b>Short description:</b> Fusion of TorA sig-seq, TEV protease cleavage site and b-lactamase<br> | ||
| + | <b>RFC:</b> 10, 23, 25 <br> | ||
<br> | <br> | ||
| - | <b> | + | <b>Full description:</b><br> |
| - | + | <b>Introduction</b><br> | |
| - | + | This device is an detector for proteolytic activity of the matching protease (see cleavage site, here for TEV protease) inside the cytoplasm of <i>E.coli</i>. If there is an active protease, than the resistance capacity against ampicillin is lowered and the cells won't survive on higher antibiotic concentrations.<br><br> | |
| + | <b>General Information of the single sub parts:</b><br> | ||
| + | <b>ssTorA</b><br> | ||
| + | The TorA signal-sequence originate from the enzyme Trimethylamin-N-Oxid-Reduktase, which needs to be folded in the cytoplasm.<br><br> | ||
| + | <b>Lactamase:</b><br> | ||
| + | The beta lactamase is an enzyme which cleaves lactam rings and makes bacteria resistant to beta lactam antibiotics like ampicillin and penicillin.<br><br> | ||
| + | <b>Proteolytic cleavage site for TEV protease:</b><br> | ||
| + | The cleavage site for the TEV protease was created via oligo hybridization and ligated into the construct with cleavage site for XhoI and NheI, which makes this part modular and easy adaptable for any other protease.<br><br> | ||
| + | <b>Setting up the right order...</b><br> | ||
| + | This construct contains the cleavage site for the TEV protease flanked by the sequence of beta-lactamase at the end and the TorA signal sequence for the TAT (twin arginine transporter) pathway infront of the cleavage site.<br><br> | ||
| + | <b>General Function</b><br> | ||
| + | This protease activity detector device provides the signal sequence from TorA for the TAT pathway, called ssTorA. These signal sequence is used for the export of folded proteins into the periplasm of gram-negative bacteria like <i>E.coli</i>.<br> | ||
| + | The modular cleavage site, which can be exchanged with the enzymes XhoI and NheI, can be adapted to any protease. This feature allows the user to check for specific proteolytic activity in the cytoplasm.<br> | ||
| + | The beta lactamase, a popular resistance marker, is used to check the activity of the desired protease inside the cytoplasm. By using the TAT pathway to export the lactamase into the periplasm the survival of bacteria cells at high ampicillin concentrations shows a low protease activity. In contrast, the death of all colony forming units indicates a high proteolytic activity inside the cytoplasm, which leads to a reduced export capacity of beta lactamase via the TAT pathway.<br> | ||
| + | Also the modular construction of the cleavage site, which can be exchanged with the restriction enzymes XhoI and NheI, makes this protease activity detector highly adaptable to any desired protease.<br><br> | ||
| + | <b>Performance and Summary:</b><br> | ||
| + | With co-transformation of the TEV protease on a second plasmid, we can show the desired proof of principal: when the protease expression was activated, the transformed <i>E.coli</i> cells did not grow at ampicillin concentrations up to 100 µg/ml. The control colonies without induction of the protease survived up to concentrations of 400 µg/ml ampicillin.<br> | ||
<br> | <br> | ||
| - | < | + | [[Image:UP_TEV-Survival.png|center|600px|thumb|'''Figure 7:''' Survival test at different ampicillin concentrations: A) approx. 1000 cfu platet on agar plates containing chloramphenicol; B) Reduced amount of <i>E.coli</i> colonies, approx. 1000 cfu were platet on agar plates containing chloramphenicol, 1mM IPTG to induce ampicillin resistance via BBa_K627012 and 400µg/ml ampicillin; C)no culture growth of <i>E.coli</i> colonies, approx. 1000 cfu were platet on agar plates containing chloramphenicol, 1mM IPTG to induce ampicillin resistance via BBa_K627012 and 600µg/ml ampicillin ]] |
<br> | <br> | ||
| - | <b> | + | <b>Source</b><br> |
| - | + | TorA signal sequence and the lactamase was amplified via PCR from the vector pJC354, which originates from two iGEM compatible BioBricks: BBa_K208005 and BBa_I757010. The cleavage site was created, based on the information of the Brenda enzymes database, via 2 oligonucleotides ordered from SigmaAldrich.<br> | |
| - | * | + | <br> |
| - | + | <b>Design Notes:</b><br> | |
| + | This BioBrick was built by PCR using the following PCR primers:<br> | ||
| + | * p_TorA-f: CTTCTAGATGAACAATAACGATCTCTTTCAGGCATC | ||
| + | * o_Bla_igem_r: CTACTAGTATTAACCGGTCCAATGCTTAATCAGTGAGGCAC | ||
| + | <br> | ||
| + | <b>References</b><br> | ||
| + | ssTorA - http://partsregistry.org/Part:BBa_K208005<br> | ||
| + | blaFL - http://partsregistry.org/Part:BBa_I757010<br> | ||
| + | <br> | ||
| + | <b>Genbank file</b><br> | ||
| + | [[Media:BioBrick_ssTorA_CS-TEV_blaFL.gb]] | ||
| - | === BioBrick | + | === BioBrick ssTorA_CS-14_3C_blaFL === |
| - | <b>Part name:</b> | + | <b>Part name:</b> BBa_K627013<br> |
<b>Part type:</b> Composite Part<br> | <b>Part type:</b> Composite Part<br> | ||
| - | <b>Short description:</b> Fusion | + | <b>Short description:</b> Fusion of ssTorA, a cleavage site for HRV 14_3C protease and beta lactamase<br> |
| + | <b>RFC:</b> 10, 23, 25 <br> | ||
<br> | <br> | ||
<b>Full description:</b><br> | <b>Full description:</b><br> | ||
| - | This | + | <b>Introduction</b><br> |
| - | + | This device is an detector for proteolytic activity of the matching protease (see cleavage site, here for HRV 14_3C protease) inside the cytoplasm of <i>E.coli</i>. If there is an active protease, than the resistance capacity against ampicillin is lowered and the cells won't survive on higher antibiotic concentrations.<br><br> | |
| + | <b>General Information of the single sub parts:</b><br> | ||
| + | <b>ssTorA</b><br> | ||
| + | The TorA signal-sequence originate from the enzyme Trimethylamin-N-Oxid-Reduktase, which needs to be folded in the cytoplasm.<br><br> | ||
| + | <b>Lactamase:</b><br> | ||
| + | The beta lactamase is an enzyme which cleaves lactam rings and makes bacteria resistant to beta lactam antibiotics like ampicillin and penicillin.<br><br> | ||
| + | <b>Proteolytic cleavage site for HRV 14_3C protease:</b><br> | ||
| + | The cleavage site for the TEV protease was created via oligo hybridization and ligated into the construct with cleavage site for XhoI and NheI, which makes this part modular and easy adaptable for any other protease.<br><br> | ||
| + | <b>Setting up the right order...</b><br> | ||
| + | This construct contains the cleavage site for the HRV 14_3C protease flanked by the sequence of beta-lactamase at the end and the TorA signal sequence for the TAT (twin arginine transporter) pathway infront of the cleavage site.<br><br> | ||
| + | <b>General Function</b><br> | ||
| + | This protease activity detector device provides the signal sequence from TorA for the TAT pathway, called ssTorA. These signal sequence is used for the export of folded proteins into the periplasm of gram-negative bacteria like <i>E.coli</i>.<br> | ||
| + | The modular cleavage site, which can be exchanged with the enzymes XhoI and NheI, can be adapted to any protease. This feature allows the user to check for specific proteolytic activity in the cytoplasm.<br> | ||
| + | The beta lactamase, a popular resistance marker, is used to check the activity of the desired protease inside the cytoplasm. By using the TAT pathway to export the lactamase into the periplasm the survival of bacteria cells at high ampicillin concentrations shows a low protease activity. In contrast, the death of all colony forming units indicates a high proteolytic activity inside the cytoplasm, which leads to a reduced export capacity of beta lactamase via the TAT pathway.<br> | ||
| + | Also the modular construction of the cleavage site, which can be exchanged with the restriction enzymes XhoI and NheI, makes this protease activity detector highly adaptable to any desired protease.<br><br> | ||
| + | <b>Performance and Summary:</b><br> | ||
| + | With co-transformation of the HRV 14_3C protease on a second plasmid, we can show the desired proof of principal: when the protease expression was activated, the transformed <i>E.coli</i> cells did not grow at ampicillin concentrations up to 100 µg/ml. The control colonies without induction of the protease survived up to concentrations of 400 µg/ml ampicillin.<br> | ||
<br> | <br> | ||
| - | <b>Source | + | <b>Source</b><br> |
| - | + | TorA signal sequence and the lactamase was amplified via PCR from the vector pJC354, which originates from two iGEM compatible BioBricks: BBa_K208005 and BBa_I757010. The cleavage site was created, based on the information of the Brenda enzymes database, via 2 oligonucleotides ordered from SigmaAldrich.<br> | |
<br> | <br> | ||
| - | <b>Design Notes</b><br> | + | <b>Design Notes:</b><br> |
| - | This | + | This BioBrick was built by PCR using the following PCR primers:<br> |
| + | * p_TorA-f: CTTCTAGATGAACAATAACGATCTCTTTCAGGCATC | ||
| + | * o_Bla_igem_r: CTACTAGTATTAACCGGTCCAATGCTTAATCAGTGAGGCAC | ||
<br> | <br> | ||
| + | <b>Experience</b><br> | ||
| + | <br> | ||
| + | <b>References</b><br> | ||
| + | ssTorA - http://partsregistry.org/Part:BBa_K208005<br> | ||
| + | blaFL - http://partsregistry.org/Part:BBa_I757010<br> | ||
| + | <br> | ||
| + | <b>Genbank file</b><br> | ||
| + | [[Media:BioBrick_ssTorA_CS-14_3C_blaFL.gb]] | ||
| + | <br><br> | ||
| + | === pSB1A3 Ara and pSB1A3 lac=== | ||
| + | |||
| + | <b>Part name:</b> BBa_K627014<br> | ||
| + | <b>Part type:</b> Plasmid Backbone<br> | ||
| + | <b>Short description:</b> Plasmid backbone pSB1A3 carrying the arabinose induction system<br> | ||
| + | <br> | ||
| + | <b>Part name:</b> BBa_K627015<br> | ||
| + | <b>Part type:</b> Plasmid Backbone<br> | ||
| + | <b>Short description:</b> Plasmid backbone pSB1A3 carrying the lac operon induction system<br> | ||
| + | |||
| + | |||
| + | <b>Full description:</b><br> | ||
| + | In a subtask of our work we wanted to construct auxiliary expression backbones with inducible promotors. In contrast to constitutive systems inducible systems only express the protein of interest after adding an inducer, which can be for example AHL or arabinose. | ||
| + | We decided to construct IPTG- and arabinose-inducible systems. Therefore we amplified the promotor region and fused it to a reporter gene, which is constituted by YFP. Using this we have the option to verify the presence of the inducible promotor and also the function of the induction process by fluorescence. Our idea was to use the reporter gene as a placeholder. Via restriction enzyme digestion using the iGEM restriction enzyme site you will be able to replace the reporter gene with your gene of interest. As vector backbone we made use of the pSB1A3, which has ampicillin resistance (Fig. 8). The constructs using pSB1K3 (kanamycin resistance) and pSB1C3 (chloramphenicol resistance) are still on the anvil. | ||
| + | [[File:UP_expression_backbones_cloning.png|center|500px|thumb|'''Figure 8:''' Cloning Scheme of Expression Backbones. ''amp'' - gene for ampicillin resistance]] | ||
| + | |||
| + | We successfully tested the IPTG- and arabinose-inducible system (Fig 9). Using fluorescence microscopy we were able to detect the YFP-expression after an induction time of approximately 1.5 hrs. To prove the hypothesis we opposed an induced sample with a non-induced control for each promotor. By use of brightfield combined with differential interference contrast microscopy we traced the E. coli cells and switched then to the YFP detecting channel to investigate the fluorescence of these cells. | ||
| + | |||
| + | [[File:UP_Expression_Backbones_Fluorescence.png|center|800px|thumb|'''Figure 9:''' Fluorescenc Microscopy. control - not induced; induced - induction with IPTG or Arabinose, respectively]] | ||
| + | |||
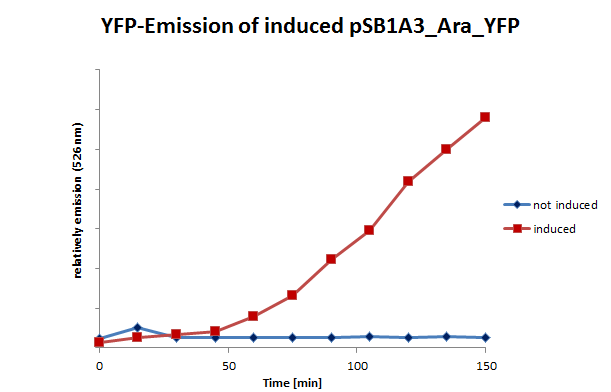
| + | We also tested the inducible systems by using fluorescence spectroscopy. For this experiment we induced both systems at a time when the cultures were located in phase of exponential growth. After inducing, the cultures were analyzed by fluorescence spectroscopy. In this analyze the cells were excitated by 500 nm. The resultant emission was measured in a spectrum between 510 and 580 nm. (Fig 10;11) | ||
| + | |||
| + | |||
| + | <table> <td>[[File:UP_psB1A3_Ara_YFPII.png|left|410px|thumb|'''Figure 10:''' Emission (526nm) of pSB1A3_Ara_YFP and control – E.coli cells (including pSB1A3_Ara_YFP) were induced and measured at certain moments.]] </td> <td>[[File:UP_psB1A3_IPTG_YFPII.png|right|410px|thumb|'''Figure 11:''' Emission (526nm) of induced pSB1A3_lac_YFP and control – E.coli cells (including pSB1A3_lac_YFP) were induced and measured at certain moments.]]</td> </table> | ||
| + | |||
| + | In contrast to the control of the arabinose inducible system the Lac-control also shows fluorescence. This demonstrates the leaky Lac-promotor, what is also a scientifically proven fact. A leaky promotor means that the promotor is not regulated in a leakproof manner. Even in repressed conditions the transcript can arise. | ||
| + | |||
| + | The lacking of the Lac-I gene on the vector is the reason for expression of YFP in both controls. For further research in this field it is necessary to clone the Lac-I gene inside the constructed vector. | ||
| + | <br> | ||
| + | <br> | ||
| + | <b>Source</b><br> | ||
| + | * pGA14mVenus Geneart | ||
| + | * Linearized BioBrick backbones | ||
| + | <br> | ||
| + | <b>Design Notes:</b><br> | ||
| + | We decided to construct IPTG- and arabinose-inducible systems. Therefore we amplified the promotor region and fused it to a reporter gene, which is constituted by YFP. Using this we have the option to verify the presence of the inducible promotor and also the function of the induction process by fluorescence. Our idea was to use the reporter gene as a placeholder. Via restriction enzyme digestion using the iGEM restriction enzyme site you will be able to replace the reporter gene with your gene of interest. As vector backbone we made use of the pSB1A3, which has ampicillin resistance (Fig. 8). The constructs using pSB1K3 (kanamycin resistance) and pSB1C3 (chloramphenicol resistance) are still on the anvil. | ||
| + | <br> | ||
| + | <br><br> | ||
| + | |||
| + | ===BioBrick TEV protease=== | ||
| + | <b>Part name:</b> BBa_K627016<br> | ||
| + | <b>Part type:</b> Coding part<br> | ||
| + | <b>Short description:</b> mutated TEV protease without iGEM restriction sites<br><br> | ||
| + | <b>General Informations</b><br> | ||
| + | TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus (TEV). It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.<br> | ||
| + | At 37°C, the TEV protease forms inclusion body, which leads to an inactive form. Incubated at 30 °C, the protease is expressed as soluble type and is highly active.<br><br> | ||
<b>Primer used for site directed mutagenesis:</b><br> | <b>Primer used for site directed mutagenesis:</b><br> | ||
<i>Fragment 1:</i><br> | <i>Fragment 1:</i><br> | ||
| Line 464: | Line 708: | ||
*f_TEV_AraFusion_NgoMIV: ATATTGCCGGCATGGGAGAAAGCCTGTTTAAGGGA<br> | *f_TEV_AraFusion_NgoMIV: ATATTGCCGGCATGGGAGAAAGCCTGTTTAAGGGA<br> | ||
*r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT<br> | *r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT<br> | ||
| - | |||
| - | |||
<br> | <br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
| - | + | Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S. P. (2007). Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16: 2360-2367 | |
| - | + | <br><br> | |
| - | + | Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D., and Waugh, D. S. (2001). Tobacco etch virus protease: Mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Prot. Eng. 14: 993-1000. | |
| + | <br><br> | ||
| + | Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable form of recombinant tobacco etch virus protease. Biotechniques 30: 544-550. | ||
| - | === BioBrick | + | ===BioBrick HRV14 3C protease=== |
| - | + | <b>Part name:</b> BBa_K627017<br> | |
| - | <b>Part name:</b> | + | <b>Part type:</b> Coding part<br> |
| - | <b>Part type:</b> | + | <b>Short description:</b> HRV14 3C protease<br> |
| - | <b>Short description:</b> | + | <b>HRV 14_3C</b><br> |
| - | <br> | + | The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln/GlyPro.<br> |
| - | <b> | + | The 22 kDa protease got its optimal activity at 4°C but also shows a high cleavage rate at 37°C. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.<br><br> |
| - | + | <b>General Function:</b><br> | |
| - | <br> | + | When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the HRV 14_3C protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.<br><br> |
| - | <b> | + | |
| - | + | ||
| - | <br> | + | |
<b>Design Notes:</b><br> | <b>Design Notes:</b><br> | ||
| - | This | + | This BioBrick was built by PCR using the following PCR primers:<br> |
<b>Site directed mutagenesis of HRV 14 3C protease:</b><br> | <b>Site directed mutagenesis of HRV 14 3C protease:</b><br> | ||
<i>Fragment 1:</i> | <i>Fragment 1:</i> | ||
| Line 516: | Line 757: | ||
* f_14_3C_AraFusion_NgoMIV: ATATTGCCGGCATGGGACCAAACACAGAATTTGCACTATCC | * f_14_3C_AraFusion_NgoMIV: ATATTGCCGGCATGGGACCAAACACAGAATTTGCACTATCC | ||
* r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | * r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA | ||
| - | |||
| - | |||
<br> | <br> | ||
<b>References:</b><br> | <b>References:</b><br> | ||
| - | + | Combination of Two Separate Binding Domains Defines Stoichiometry between Type III Secretion System Chaperone IpgC and Translocator Protein IpaB, Ravi Kumar Lokareddy, Michele Lunelli, Björn Eilers, Vivien Wolter, and Michael Kolbe, December 17, 2010 The Journal of Biological Chemistry, 285, 39965-39975. | |
| - | + | <br><br> | |
| - | + | Cleavage of Small Peptides In Vitro by Human Rhinovirus 14 3C Protease expressed in Escherichia coli, MICHAEL G. CORDINGLEY,* R. BRUCE REGISTER, PIA L. CALLAHAN, VICTOR M. GARSKY,AND RICHARD J. COLONNO JOURNAL OF VIROLOGY, Dec. 1989, p. 5037-5045 | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 23:32, 28 October 2011
BioBricks
| Label number | BioBrick Nickname | Tube label |
|---|---|---|
| BBa_K627000 | mdnED | 1 |
| BBa_K627001 | mdnA | 2 |
| BBa_K627002 | mdnB | 3 |
| BBa_K627003 | mdnC | 4 |
| BBa_K627004 | mdnD | 5 |
| BBa_K627005 | mdnE | 6 |
| BBa_K627006 | mdnA c-myc gene III | 7 |
| BBa_K627007 | c-myc gene III | 8 |
| BBa_K627008 | AraC TEV protease 1 | 9 |
| BBa_K627009 | AraC TEV protease 2 | 10 |
| BBa_K627010 | AraC TEV protease 3 | 11 |
| BBa_K627011 | AraC 14_3C protease | 12 |
| BBa_K627012 | ssTorA CS-TEV BlaFL | 13 |
| BBa_K627013 | ssTorA CS-14_3C BlaFL | 14 |
| BBa_K627014 | A3_Ara_YFP | 15 |
| BBa_K627015 | A3_lac_YFP | 16 |
| BBa_K627016 | TEV protease | 17 |
| BBa_K627017 | HRV14 3C protease | 18 |
BioBrick mdnED
Part name: BBa_K627000
Part type: Coding
Short description: ABC transporter and N-acetyltransferase from the mdn-cluster
Full description:
The BioBrick mdnED is a part of the whole microviridin gene (mdn) cluster, which encodes the protease
inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized
by the cyanobacteria Microcystis aeruginosa (Ziemert et al., 2010). They have a promising potential for therapy as they can block
disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L
biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes,
which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al.,
2008).
In the following BioBrick mdnED the genes mdnD (N-acetyltransferase of the GNAT family) and mdnE (ABC
transporter) is encoded (Ziemert et al., 2008).
Source of the part:
The BioBrick mdnDE as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: TAAATGAATTCGCGGCCGCTTCTAGATGCCTCAATATACTACTAAAC
Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTATCAGCAAACCCTACTTAATTTC
To insert mdnED in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is an expression part, the adenine of mdnE gene's start codon is part of the XbaI
recognition site.
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file:
Media: UP_BioBrick_mdnED.gb
BioBrick mdnA
Part name: BBa_K627001
Part type: Coding
Short description: mdnA gene encoding the precursor peptide of the tricyclic microviridin
Full description:
The BioBrick mdnA is a part of the whole microviridin gene (mdn) cluster, which encodes the protease inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized by Microcystis aeruginosa (Ziemert et al., 2010). They have a promising potential for therapy as they can block disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes, which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al., 2008).
The following BioBrick mdnA encodes the ribosomal precursor peptide (MdnA), which is essential for microviridin production (Ziemert et al., 2008).
Because this BioBrick is a RFC10 expression part, the adenine of mdnB gene`s start codon is part of the XbaI recognition site.
Usage and Biology:
Microviridin production in E. coli cells expressing the mdn genes was monitored by reverse phase HPLC. Qualitative analysis involves running a standard that contains the target analytes. The vectors pARW071 and pARW089 served as a control in our experiments because these vectors contain the original mdn-cluster. We could use the retention time as a way to determine the presence of the microviridin production in other samples. HPLC analysis of the purified compound yielded a high peak with a retention time of approximately 5 min. Minor peaks could be detected during the following 7 min (Fig. 1).
All HPLC chromatograms of the isolated mdnA showed reliable peaks with a retention time of 5 min together with a number of following minor peaks.
Fractions of the peaks were sampled and the identity of microviridin and also the presence of cyclization, which is important for the activity, was affirmed with mass spectrometry (Fig. 2). By using a mass spectrometer it is possible to determine both the elemental composition of a fraction and the chemical structure of molecules.
Source of the part:
The BioBrick mdnA as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: TTCCATGGCGCCAGAGGAATCTAGATGGCATATCCCAACGATC
Reverse primer: CTTCTGACTGGGAAGATTATACCGGTTAATACTAGTAGCGGCCGCTGCAGGACGTC
To insert mdnA in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is an expression part, the adenine of mdnA gene`s start codon is part of the XbaI recognition site. Further the sequence contains a AgeI recognition site after mdnA.
Experiences:
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file:
Media: UP_Biobrick_mdnA.gb
BioBrick mdnB
Part name: BBa_K627002
Part type: Coding
Short description: ATP-grasp-type ligase from mdn-cluster
Full description:
The BioBrick mdnB is a part of the whole microviridin gene (mdn) cluster, which encodes the protease inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized by Microcystis aeruginosa (Ziemert et al., 2010). They have a promising potential for therapy as they can block disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes, which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al., 2008).
The following BioBrick mdnB encodes a ATP-grasp-type ligase (Ziemert et al., 2008).
Because this BioBrick is an expression part, the adenine of mdnB gene`s start codon is part of the XbaI recognition site.
Source of the part:
The BioBrick mdnB as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: ATTATGAATTCGCGGCCGCTTCTAGATGAAAGAATCGCCTAAAGTTG
Reverse primer: TAATCTGCAGCGGCCGCTACTAGTATCAACCGAAGACTAAAAAATCAGCG
To insert mdnB in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is an expression part, the adenine of mdnE gene's start codon is part of the XbaI recognition site.
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file:
Media: UP_BioBrick_mdnB.gb
BioBrick mdnC
Part name: BBa_K627003
Part type: Coding
Short description: ATP-grasp-type ligase from the mdn-cluster
Full description:
The BioBrick mdnC is a part of the whole microviridin gene (mdn) cluster, which encodes the protease
inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized
by Microcystis (Ziemert et al., 2010). They have a promising potential for therapy as they can block
disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L
biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes,
which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al.,
2008).
The following BioBrick mdnC encodes the ATP-grasp-type ligase (Ziemert et al., 2008).
Because this BioBrick is a RF10 expression part, the adenine of mdnC gene`s start codon is part of the XbaI
recognition site.
Source of the part:
The BioBrick mdnC as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: TATTTGAATTCGCGGCCGCTTCTAGATGACCGTTTTAATTGTTAC
Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTATTATGAGTTAACTAGGATTTC
To insert mdnC in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is a RF10 expression part, the adenine of mdnA gene`s start codon is part of the XbaI recognition site.
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file:
Media: UP_BioBrick_mdnC.gb
BioBrick mdnD
Part name: BBa_K627004
Part type: Coding
Short description: N-acetyltransferase from the mdn-cluster
Full description:
The BioBrick mdnD is a part of the whole microviridin gene (mdn) cluster, which encodes the protease
inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized
by Microcystis (Ziemert et al., 2010). They have a promising potential for therapy as they can block
disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L
biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes,
which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al.,
2008).
The following BioBrick mdnD encodes a N-acetyltransferase of the GNAT family (Ziemert et al., 2008).
Because this BioBrick is a RF10 expression part the adenine of mdnD gene's start codon is part of the XbaI
recognition site.
Source of the part:
The BioBrick mdnD as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: TATATGAATTCGCGGCCGCTTCTAGATGAAAGCACTGGAAAAACTG
Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTATCAGCAAACCCTACTTAATTTC
To insert mdnD in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is a RF10 expression part the adenine of mdnD gene`s start codon is part of the XbaI recognition site.
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file
Media: UP_BioBrick_mdnD.gb
BioBrick mdnE
Part name: BBa_K627005
Part type: Coding
Short description: ABC transporter from the mdn-cluster
Full description:
The BioBrick mdnE is a part of the whole microviridin gene (mdn) cluster, which encodes the protease
inhibitor microviridin L. Microviridins are tricyclic depsipeptides, which are ribosomally synthesized
by Microcystis (Ziemert et al., 2010). They have a promising potential for therapy as they can block
disease-relevant proteases (Ziemert et al., 2008).
Microviridins are synthesized from a ribosomal precursor peptide (MdnA). Additionally, the microviridin L
biosynthesis gene cluster consists of genes encoding an ATP-grasp-type ligase (mdnB and mdnC) and genes,
which encode an ABC transporter (mdnE) and a N-acetyltransferase of the GNAT family (mdnD) (Ziemert et al.,
2008).
The following BioBrick mdnE encodes an ABC transporter (Ziemert et al., 2008).
Because this BioBrick is a RF10 expression part the adenine of mdnE gene`s start codon is part of the XbaI
recognition site.
Source of the part:
The BioBrick mdnE as a part of the microviridin gene (mdn) cluster was isolated from Microcystis aeruginosa strain NIES-843.
Design information:
This BioBrick was built by PCR using the following PCR primers
Forward primer: TAAATGAATTCGCGGCCGCTTCTAGATGCCTCAATATACTACTAAAC
Reverse primer: ATTTCTGCAGCGGCCGCTACTAGTACTATATTCTCACCCATTTTAAG
To insert mdnE in the vector pSB1C3, the resulting PCR product and the vector were digested with the restriction enzymes EcoRI and SpeI.
Because this BioBrick is a RF10 expression part the adenine of mdnE gene`s start codon is part of the XbaI recognition site.
References:
Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file
Media: UP_BioBrick_mdnE.gb
BioBrick mdnA-myc-gene III
Part name: BBa_K627006
Part type: Coding
Short description: Fusion part of mdnA gene (from mdn-cluster) with myc-tag and gene III
Full description
This BioBrick is important for testing the suitability of a phage display system as an appropriate screening method for recombinant mdnA libraries.
The mdnA gene encodes the precursor peptide from the mdn-cluster, which is an essential building block for production of the protease inhibitor microviridin from Microcystis aeruginosa (Ziemert et al., 2008 and 2010). This peptide is known to be able to bind and inhibit proteases. Thus, variants of mdnA have a high potential for therapy as they can block disease-relevant proteases.
The gene III protein is a coat protein from the filamentous bacteriophage M13 (Smith, 1985). It appears only three to five times on the tip of the phage and is responsible for infection of bacterial cells. In phage display, genes from a DNA library are usually fused to the gene III and are displayed as fusion protein on the surface of the phages. Because it has been shown that the proteins of interest, which are fused to the C-terminus of gene-III can be functionally displayed (Krebber et al. 1997, Fuh and Sidhu, 2000, Rakonjac et al., 1999) only the sequence for this part has to be used.
The inserted sequence of the myc-tag enables easy detection and/or purification of the proteins of interest. In our case, the tag was used to verify display of the microviridin on the phage particle.
After transformation of E. coli with a vector containing the mdnA-myc-geneIII-fusion gene (in the context of the mdnA gene cluster) and co-infection with helper phages, E. coli cells were able to produce phage particles, which carry the microviridin peptide on their surface as confirmed by ELISA. These experiments proved the suitability of phage display as a screening method for variants of microviridin.
Control of expression of mdnA-myc-gene III in E. coli
To control expression of the mdnA-myc-gene III fusion gene was analyzed by western blotting. E. coli cells transformed with the phagemid pPDV089 were harvested and lysated. The cell proteins were electrophoretically separated and transferred to a membrane. The mdnA-myc-gene III-fusion proteins were detected using specific anti-myc-antibodies and horseradish peroxidase (HRP)-linked antibodies as second antibodies. Enhanced chemiluminescence (ECL) was used to visualize the protein. ECL is based on the emission of light during the HRP -catalyzed oxidation of luminol, which was captured by a camera. The western blot analysis resulted in a band of a size just below the 30 kDa mark representing the mdnA-myc-gene III-fusion protein (24 kDa).
Detection of phages carrying mdnA on their surface by ELISA
The next step was the detection of the expression of the mdnA-myc-gene III-fusion gene on the surface of the phage. So E. coli cells strain XL1-Blue were first transformed with the phagemid pPDV089 before they were infected with helper phages. E. coli cells containing both plasmids were selected. An ELISA test was performed to determine whether these cells are able to produce phage particles carrying the mdnA peptide on their surface. To perform this test anti-c-myc-antibodies were immobilized on ELISA plates and incubated with purified phages. For detection a second antibody coupled with horse radish peroxidase (HRP) was used which binds the gene VIII protein of the phages. The HRP substrate o-phenyldiamine (OPD) was added and in case of binding a color reaction was expected. The color shift from achromatic to yellow in wells incubated with phages produced in XL1-Blue cells showed the successful expression of mdnA-c-myc-gene III-fusion protein on the phages.
For more precise results the absorption at 492 nm was measured. The data were presented in a bar plot. As a negative control helper phages were added instead of produced phages. Furthermore two wells were prepared were the secondary antibody was not added.
The graphic shows clearly the much higher absorption measured in wells, which were incubated with phage particles of interest produced in XL1-Blue cells. As has already pointed out this shows the succeeded expression of mdnA-c-myc-gene III-fusion protein on the surface of the phages.
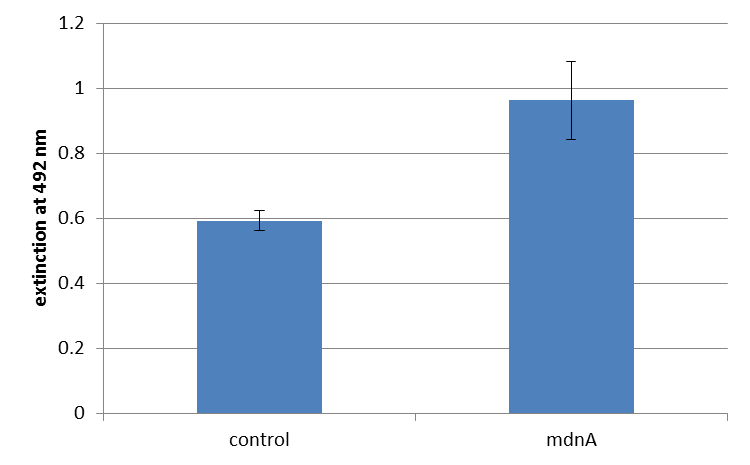
Testing phage display with unmodified mdnA to examine its suitability as screening method
To test the suitability of this screening method, phages representing unmodified mdnA on their surface and phages not representing mdnA (helper phages) in a ratio of one to one were incubated with immobilized trypsin which is known as a target of mdnA. The display was conducted in ELISA plates. The bound phages were eluted using a buffer with low pH value and neutralized afterwards. To check how many phages interacted with trypsin, E. coli cells XL1-Blue were re-infected with eluted phages and plated on agar with different antibiotics. Cells infected with phages carrying mdnA are able to grow on agar with ampicillin whereas cells infected with helper phages are able to grow on agar with kanamycin. To control the success of the panning round additionally E. coli cells were infected with phage mix before panning and plated on agar. Subsequent the number of clones grew on ampicillin and kanamycin before and after panning was compared. During the running of this step it was noticed that much more cells were infected with helper phages than with phages carrying mdnA despite of the engaged 1:1 ratio. This was surprising and indicated that mdnA on the surface of the phages may inhibit their infectivity. After controlling the plates an infection ratio of phages carrying mdnA to helper phages of 1:400 was calculated. This fact should be analyzed in further experiments.
The results of the first phage display are plotted in the figure below. After one panning round an enrichment of phages carrying mdnA was expected. This is attributable to the fact that phage particles carrying mdnA c-myc geneIII fusion protein on their surface are expected to bind specifically to the immobilized trypsin. Unfortunately this was not observed in this experiment. The ratio of cells growing on kanamycin agar before (4000) to cells growing on kanamycin agar (cells containing helper phages) after panning (750) was determined as 5:1. The ratio of cells growing on ampicillin agar before (12) to cells growing on ampicillin agar (cells containing mdnA carrying phages) after panning (2) was nearly equal. Thus no enrichment of mdnA carrying phages occurred in the first experiment. So it was decided to repeat this experiment under improved conditions. Therefor the number of washing steps during the described experimental procedure was increased. Here the ratio of cells growing on kanamycin agar before (3000) to cells growing on kanamycin agar (cells containing helper phages) after panning (29) was determined as 103:1. The ratio of cells growing on ampicillin agar before (26) to cells growing on ampicillin agar (cells containing mdnA carrying phages) after panning (2) was determined as 13:1. Thus an enrichment factor of eight was reached for the phages displaying mdnA on their surface.
These results indicate that the unmodified mdnA expressed on the phages binds specifically to the immobilized trypsin. Therefore it can be deduced that mdnA is presented in a functional 3D structure. These findings suggest that phage display in general is an appropriate method for screening a recombinant mdnA library. Further experiments are required to optimize this system.
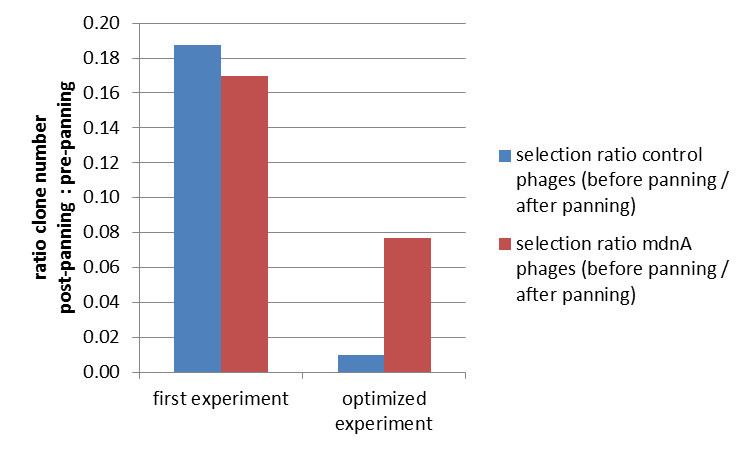
Testing phage display with unmodified mdnA against further proteases
Furthermore the interaction of unmodified mdnA with other proteases was determined. From the literature (Ziemert, 2010) the high inhibitory activity of microviridin L, besides trypsin, against chymotrypsin and elastase is also known. So a phage display with these enzymes was performed. Additionally papain, proteinase K, mycolysin and pepsin were tested for which no interaction was shown yet. For all enzymes an equal amount of E. coli cells and phages were used. In agreement with data from the literature interaction of microviridin with chymotrypsin and elastase was confirmed. Additionally an interaction with papain was noted. All other proteases were not bound by microviridin.
Source of the part:
The BioBrick mdnA is a part of the microviridin gene (mdn) cluster which was isolated from Microcystis aeruginosa strain NIES-843.
The gene III protein is a coat protein from the filamentous bacteriophage M13.
Design information:
Gene III was amplified from the vector pak100blaKDIR using the following primers
Forward:TAAGCTTCTAGATGGCCGGCGAGCAGAAGCTGATCTCTGAGGAAGACCTGGGTGGTGGCTCTGGTTCC
Reverse: TGCTTAGACGTCCTGCAGCGGCCGCTACTAGTATTAACCGGTAGACTCCTTATTACGCAGTA
The mdnA gene was amplified from the vector pARW089 using the following primers
Forward: TTCCATGGCGCCAGAGGAATCTAGATGGCATATCCCAACGATC
Reverse: CTTCTGACTGGGAAGATTATACCGGTTAATACTAGTAGCGGCCGCTGCAGGACGTC
Gene III was amplified from pak100blaKDIR and mdnA from pARW089 by PCR. The primers were designed to enable the introduction of iGEM. The PCR product gene III was digested by whereas the PCR product mdnA was digested by AgeI. Thus a mdnA-gene III fusion part according to RFC25 was generated whereby AgeI and NgoMIV overhangs are compatible and placed in frame with the protein sequence. The ligation of AgeI and NgoMIV overhangs resulted in a scar coding for the threonine and glycine. Because the introduction of restriction sites before mdnA leaded to a great distance between ribosome binding site and mdnA a second RBS was inserted among SfoI and XbaI recognition sites to ensure a sufficiently expression rate of the mdnA-gene III-fusion gene. Between mdnA and gene III myc sequence was inserted.
Because this BioBrick is an RFC25 expression part, the adenine of mdnA gene's start codon is part of the XbaI recognition site. Further the sequence contains a AgeI recognition site after gene III.
References:
- Fuh G., Sidhu S.S. (2000). Efficient phage display of polypeptides fused to the carboxy-terminus of the M13 gene-3 minor coat protein. FEBS Lett. 480(2-3):231-4
- Krebber, A., Bornhauser, S., Burmester, J., Honegger, A., Willuda, J., Bosshard, H. R., Plückthun, A. (1997) Reliable cloning of functional antibody variable domains from hybridomas and spleen cell repertoires employing a reengineered phage display system. J. Immunol. Meth. 201(1):35-55
- Rakonjac J., Feng J., Model P. (1999). Filamentous phage are released from the bacterial membrane by a two-step mechanism involving a short C-terminal fragment of pIII. J Mol Biol. 289(5):1253-65
- Smith, G.P. (1985). Filamentous fusion phage: Novel expression vectors that display cloned antigens on the virus surface. Science 228: 1315-17
- Ziemert, N., Ishida, K., Liaimer, A., Hertweck, C. & Dittmann, E. (2008). Ribosomal synthesis of tricyclic depsipeptides in bloom-forming cyanobacteria. Angewandte Chemie (International ed. in English) 47, 7756-9
- Ziemert, N., Ishida, K., Weiz, A., Hertweck, C. & Dittmann, E. (2010). Exploiting the natural diversity of microviridin gene clusters for discovery of novel tricyclic depsipeptides. Applied and environmental microbiology 76, 3568-74
Genbank file:
Media: UP_Biobrick_mdnA_c-myc_geneIII.gb
BioBrick myc-gene III
Part name: BBa_K627007
Part type: Coding
Short description: Fusion of c-myc-tag and gene III
Full description:
The gene III protein is a coat protein from the filamentous bacteriophage M13 (Smith, 1985). It appears only 3-5 times on the tip of the phage and is responsible for infection of bacterial cells. It is usually fused to the gene of interest to determine protein-protein-interactions of its expression product using M13 phage display system. The gene III-protein consists of three domains: one C-termial and two N-terminal domains. Gene III was generated as biobrick several times for instance M13003 (2006) and M31537 (2007). It has been shown that proteins of interest fused to the carboxy-terminus of gene-III-proteins can be functionally displayed on the surface of phages (Fuh and Sidhu, 2000, Rakonjac et al., 1999). In the course of our project a phagemid was constructed carrying truncated gene III (only encoding C-terminal domain) as biobrick. To enable easy purification and detection of gene-III-fused proteins additionally a myc-tag-sequence was merged before gene –III-sequence.
Source of the part:
The gene-III-protein is a coat protein from the filamentous bacteriophage M13.
Design information:
Gene III was amplified from the vector pak100blaKDIR using the following primers
Forward:TAAGCTTCTAGATGGCCGGCGAGCAGAAGCTGATCTCTGAGGAAGACCTGGGTGGTGGCTCTGGTTCC
Reverse: TGCTTAGACGTCCTGCAGCGGCCGCTACTAGTATTAACCGGTAGACTCCTTATTACGCAGTA
Because myc-gene III is a fusion part (RFC25), no start codon is needed. Before the myc sequence a NgoMIV restriction site is located which has compatible overhangs to AgeI restriction sites. The ligation of AgeI and NgoMIV overhangs will result in a scar coding for threonine and glycine. So genes of interest containing AgeI restriction sites can be easily fused to gene III without frame shift. The gene III sequence contains no stop codon because there is one located between AgeI and SpeI recognition site directly merged behind gene III sequence.
References:
Fuh G., Sidhu S.S. (2000). Efficient phage display of polypeptides fused to the carboxy-terminus of the M13 gene-3 minor coat protein. FEBS Lett. 480(2-3):231-4
Rakonjac J., Feng J., Model P. (1999). Filamentous phage are released from the bacterial membrane by a two-step mechanism involving a short C-terminal fragment of pIII. J Mol Biol. 289(5):1253-65
Smith, G.P. (1985). Filamentous fusion phage: Novel expression vectors that display cloned antigens on the virus surface. Science 228: 1315-17
Genbank file:
Media: UP_Biobrick_myc_gene III.gb
BioBrick AraC-TEV
Part name: BBa_K627008, BBa_K627009, BBa_K627010
Part type: Device
Short description: Fusion part of arabinose-inducible induction system and the TEV protease
RFC: 10, 12, 23
Full description:
Introduction
This BioBrick is a 2 part fusion of arabinose-inducible induction system and the TEV protease.
General Information of the single sub parts:
TEV protease
TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus (TEV). It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.
At 37°C, the TEV protease forms inclusion body, which leads to an inactive form. Incubated at 30 °C, the protease is expressed as soluble type and is highly active.
Arabinose-inducible induction system
The arabinose-inducible induction system was amplified via PCR from the pBAD_iGEMexpress vector. Based on the inhibitionary funtion of the catabolic activator protein (CAP) it got a very low expression rate without induction by arabinose. For the induction concentration between 2 mM up to 50 mM arabinose can be used to get an high expression rate.
General Function:
When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the TEV protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.
Performance and Summary:
In combination with our created ssTorA_CS-TEV_blaFL device (BBa_K627012, [http://partsregistry.org/Part:BBa_K627012 More Information]) this protease generator was able to mediate the cell death of our transformed E.coli cells at ampicillin concentration up to 100 µg/ml.
Source of the part:
Arabinose inducible operon form pBAD_iGEM_express, TEV protease from Gunther Stier et al.
Design Notes
This BioBrick was built by PCR using the following PCR primers:
Primer used for site directed mutagenesis:
Fragment 1:
- r_TEV_ACCAGC: GGAATGTGCAGCTGGTGTCTGACACC
- f_TEV_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGAGAAAGCTTGTTTAAGGGA
Fragment 2:
- r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
- f_TEV_ACCAGC: GGTGTCAGACACCAGCTGCACATTCC
Primer used for assembly PCR of mutated fragments:
- f_TEV_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGAGAAAGCTTGTTTAAGGGA
- r_TEV_iGEM_BamHI: TATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
Primers used for amplification of pBAD arabinose-inducible induction system:
- f_AraC_iGEM_HindIII: TATAAGCTTGAATTCGCGGCCGCTTCTAGATTATGACAACTTGACGGCTACATCATT
- r_AraC_NgoMIV: ATAGCCGGCCTCCTTCTTAAAGTTAAACAAAATTATTTCTAGCCC
Primers used for amplification and modification of TEV protease:
- f_TEV_AraFusion_NgoMIV: ATATTGCCGGCATGGGAGAAAGCCTGTTTAAGGGA
- r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
Experience:
References:
Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S. P. (2007). Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16: 2360-2367
Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D., and Waugh, D. S. (2001). Tobacco etch virus protease: Mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Prot. Eng. 14: 993-1000.
Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable form of recombinant tobacco etch virus protease. Biotechniques 30: 544-550.
Genbank file
Media:BioBrick_AraC-TEV_comp.gb
Three parts are existing.
BioBrick AraC-14_3C protease
Part name: BBa_K627011
Part type:Device
Short description: Fusion part of arabinose-inducible induction system and the HRV 14_3C protease
RFC: 10, 12, 23
Full description:
Introduction
This BioBrick is a 2 part fusion of arabinose-inducible induction system and the HRV 14_3C protease.
General Information of the single sub parts:
HRV 14_3C
The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln/GlyPro.
The 22 kDa protease got its optimal activity at 4°C but also shows a high cleavage rate at 37°C. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.
Arabinose-inducible induction system
The arabinose-inducible induction system was amplified via PCR from the pBAD_iGEMexpress vector. Based on the inhibitionary funtion of the catabolic activator protein (CAP) it got a very low expression rate without induction by arabinose. For the induction concentration between 2 mM up to 50 mM arabinose can be used to get an high expression rate.
General Function:
When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the HRV 14_3C protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.
Performance and Summary:
In combination with our created ssTorA_CS-14_3C_blaFL device (BBa_K627013, [http://partsregistry.org/Part:BBa_K627013 More Information]) this protease generator was able to mediate the cell death of our transformed E.coli cells at ampicillin concentration up to 100 µg/ml.
Source of the part:
Arabinose inducible operon form pBAD_iGEM_express was ampilied via PCR, 14_3C protease was isolated from the genome of human rhinovirus 14_3C and mutated with primers listed in part description BBa: to remove all iGEM restriction sites inside the protease gene.
Design Notes:
This BioBrick was built by PCR using the following PCR primers:
Site directed mutagenesis of HRV 14 3C protease:
Fragment 1:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_ACCAGC: ACCATTTGCTGGTACATCATCACCAGG
Fragment 2:
- f_14_3C_ACCAGC: CCTGGTGATGATGTACCAGCAAATGGT
- r_14_3C_tm_XbaI208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC
Fragment 3:
- f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG
- r_14_3C_tm_XbaI280_A-T: ATCCACACCTTCGAGATCTTCTGATAT
Fragment 4:
- f_14_3C_tm_Xba280_A-T: ATATCAGAAGATCTCGAAGGTGTGGAT
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primers used for first assembly PCR of mutated fragments:
Fragment 5, containing Fragment 1 and 2:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_tm_Xba208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC
Fragment 6, containing Fragment 3 and 4:
- f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primer used for final assembly PCR of mutated fragments:
Complete mutated 14_3C protease, containing Fragment 5 and 6:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primers used for amplification of the pBAD arabinose-inducible induction system:
- f_AraC_iGEM_HindIII: TATAAGCTTGAATTCGCGGCCGCTTCTAGATTATGACAACTTGACGGCTACATCATT
- r_AraC_NgoMIV: ATAGCCGGCCTCCTTCTTAAAGTTAAACAAAATTATTTCTAGCCC
Primers used for amplification and modification of HRV 14 3C protease:
- f_14_3C_AraFusion_NgoMIV: ATATTGCCGGCATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Experience:
References:
Combination of Two Separate Binding Domains Defines Stoichiometry between Type III Secretion System Chaperone IpgC and Translocator Protein IpaB, Ravi Kumar Lokareddy, Michele Lunelli, Björn Eilers, Vivien Wolter, and Michael Kolbe, December 17, 2010 The Journal of Biological Chemistry, 285, 39965-39975.
Cleavage of Small Peptides In Vitro by Human Rhinovirus 14 3C Protease expressed in Escherichia coli, MICHAEL G. CORDINGLEY,* R. BRUCE REGISTER, PIA L. CALLAHAN, VICTOR M. GARSKY,AND RICHARD J. COLONNO JOURNAL OF VIROLOGY, Dec. 1989, p. 5037-5045
Genbank file:
Media:BioBrick_AraC-14_3C.gb
BioBrick ssTorA_CS-TEV_blaFL
Part name: BBa_K627012
Part type: Composite Part
Short description: Fusion of TorA sig-seq, TEV protease cleavage site and b-lactamase
RFC: 10, 23, 25
Full description:
Introduction
This device is an detector for proteolytic activity of the matching protease (see cleavage site, here for TEV protease) inside the cytoplasm of E.coli. If there is an active protease, than the resistance capacity against ampicillin is lowered and the cells won't survive on higher antibiotic concentrations.
General Information of the single sub parts:
ssTorA
The TorA signal-sequence originate from the enzyme Trimethylamin-N-Oxid-Reduktase, which needs to be folded in the cytoplasm.
Lactamase:
The beta lactamase is an enzyme which cleaves lactam rings and makes bacteria resistant to beta lactam antibiotics like ampicillin and penicillin.
Proteolytic cleavage site for TEV protease:
The cleavage site for the TEV protease was created via oligo hybridization and ligated into the construct with cleavage site for XhoI and NheI, which makes this part modular and easy adaptable for any other protease.
Setting up the right order...
This construct contains the cleavage site for the TEV protease flanked by the sequence of beta-lactamase at the end and the TorA signal sequence for the TAT (twin arginine transporter) pathway infront of the cleavage site.
General Function
This protease activity detector device provides the signal sequence from TorA for the TAT pathway, called ssTorA. These signal sequence is used for the export of folded proteins into the periplasm of gram-negative bacteria like E.coli.
The modular cleavage site, which can be exchanged with the enzymes XhoI and NheI, can be adapted to any protease. This feature allows the user to check for specific proteolytic activity in the cytoplasm.
The beta lactamase, a popular resistance marker, is used to check the activity of the desired protease inside the cytoplasm. By using the TAT pathway to export the lactamase into the periplasm the survival of bacteria cells at high ampicillin concentrations shows a low protease activity. In contrast, the death of all colony forming units indicates a high proteolytic activity inside the cytoplasm, which leads to a reduced export capacity of beta lactamase via the TAT pathway.
Also the modular construction of the cleavage site, which can be exchanged with the restriction enzymes XhoI and NheI, makes this protease activity detector highly adaptable to any desired protease.
Performance and Summary:
With co-transformation of the TEV protease on a second plasmid, we can show the desired proof of principal: when the protease expression was activated, the transformed E.coli cells did not grow at ampicillin concentrations up to 100 µg/ml. The control colonies without induction of the protease survived up to concentrations of 400 µg/ml ampicillin.
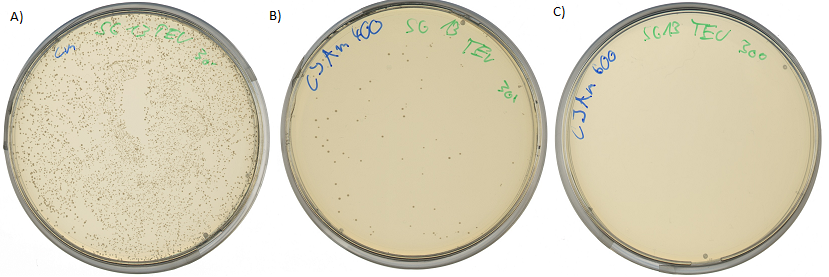
Source
TorA signal sequence and the lactamase was amplified via PCR from the vector pJC354, which originates from two iGEM compatible BioBricks: BBa_K208005 and BBa_I757010. The cleavage site was created, based on the information of the Brenda enzymes database, via 2 oligonucleotides ordered from SigmaAldrich.
Design Notes:
This BioBrick was built by PCR using the following PCR primers:
- p_TorA-f: CTTCTAGATGAACAATAACGATCTCTTTCAGGCATC
- o_Bla_igem_r: CTACTAGTATTAACCGGTCCAATGCTTAATCAGTGAGGCAC
References
ssTorA - http://partsregistry.org/Part:BBa_K208005
blaFL - http://partsregistry.org/Part:BBa_I757010
Genbank file
Media:BioBrick_ssTorA_CS-TEV_blaFL.gb
BioBrick ssTorA_CS-14_3C_blaFL
Part name: BBa_K627013
Part type: Composite Part
Short description: Fusion of ssTorA, a cleavage site for HRV 14_3C protease and beta lactamase
RFC: 10, 23, 25
Full description:
Introduction
This device is an detector for proteolytic activity of the matching protease (see cleavage site, here for HRV 14_3C protease) inside the cytoplasm of E.coli. If there is an active protease, than the resistance capacity against ampicillin is lowered and the cells won't survive on higher antibiotic concentrations.
General Information of the single sub parts:
ssTorA
The TorA signal-sequence originate from the enzyme Trimethylamin-N-Oxid-Reduktase, which needs to be folded in the cytoplasm.
Lactamase:
The beta lactamase is an enzyme which cleaves lactam rings and makes bacteria resistant to beta lactam antibiotics like ampicillin and penicillin.
Proteolytic cleavage site for HRV 14_3C protease:
The cleavage site for the TEV protease was created via oligo hybridization and ligated into the construct with cleavage site for XhoI and NheI, which makes this part modular and easy adaptable for any other protease.
Setting up the right order...
This construct contains the cleavage site for the HRV 14_3C protease flanked by the sequence of beta-lactamase at the end and the TorA signal sequence for the TAT (twin arginine transporter) pathway infront of the cleavage site.
General Function
This protease activity detector device provides the signal sequence from TorA for the TAT pathway, called ssTorA. These signal sequence is used for the export of folded proteins into the periplasm of gram-negative bacteria like E.coli.
The modular cleavage site, which can be exchanged with the enzymes XhoI and NheI, can be adapted to any protease. This feature allows the user to check for specific proteolytic activity in the cytoplasm.
The beta lactamase, a popular resistance marker, is used to check the activity of the desired protease inside the cytoplasm. By using the TAT pathway to export the lactamase into the periplasm the survival of bacteria cells at high ampicillin concentrations shows a low protease activity. In contrast, the death of all colony forming units indicates a high proteolytic activity inside the cytoplasm, which leads to a reduced export capacity of beta lactamase via the TAT pathway.
Also the modular construction of the cleavage site, which can be exchanged with the restriction enzymes XhoI and NheI, makes this protease activity detector highly adaptable to any desired protease.
Performance and Summary:
With co-transformation of the HRV 14_3C protease on a second plasmid, we can show the desired proof of principal: when the protease expression was activated, the transformed E.coli cells did not grow at ampicillin concentrations up to 100 µg/ml. The control colonies without induction of the protease survived up to concentrations of 400 µg/ml ampicillin.
Source
TorA signal sequence and the lactamase was amplified via PCR from the vector pJC354, which originates from two iGEM compatible BioBricks: BBa_K208005 and BBa_I757010. The cleavage site was created, based on the information of the Brenda enzymes database, via 2 oligonucleotides ordered from SigmaAldrich.
Design Notes:
This BioBrick was built by PCR using the following PCR primers:
- p_TorA-f: CTTCTAGATGAACAATAACGATCTCTTTCAGGCATC
- o_Bla_igem_r: CTACTAGTATTAACCGGTCCAATGCTTAATCAGTGAGGCAC
Experience
References
ssTorA - http://partsregistry.org/Part:BBa_K208005
blaFL - http://partsregistry.org/Part:BBa_I757010
Genbank file
Media:BioBrick_ssTorA_CS-14_3C_blaFL.gb
pSB1A3 Ara and pSB1A3 lac
Part name: BBa_K627014
Part type: Plasmid Backbone
Short description: Plasmid backbone pSB1A3 carrying the arabinose induction system
Part name: BBa_K627015
Part type: Plasmid Backbone
Short description: Plasmid backbone pSB1A3 carrying the lac operon induction system
Full description:
In a subtask of our work we wanted to construct auxiliary expression backbones with inducible promotors. In contrast to constitutive systems inducible systems only express the protein of interest after adding an inducer, which can be for example AHL or arabinose.
We decided to construct IPTG- and arabinose-inducible systems. Therefore we amplified the promotor region and fused it to a reporter gene, which is constituted by YFP. Using this we have the option to verify the presence of the inducible promotor and also the function of the induction process by fluorescence. Our idea was to use the reporter gene as a placeholder. Via restriction enzyme digestion using the iGEM restriction enzyme site you will be able to replace the reporter gene with your gene of interest. As vector backbone we made use of the pSB1A3, which has ampicillin resistance (Fig. 8). The constructs using pSB1K3 (kanamycin resistance) and pSB1C3 (chloramphenicol resistance) are still on the anvil.
We successfully tested the IPTG- and arabinose-inducible system (Fig 9). Using fluorescence microscopy we were able to detect the YFP-expression after an induction time of approximately 1.5 hrs. To prove the hypothesis we opposed an induced sample with a non-induced control for each promotor. By use of brightfield combined with differential interference contrast microscopy we traced the E. coli cells and switched then to the YFP detecting channel to investigate the fluorescence of these cells.
We also tested the inducible systems by using fluorescence spectroscopy. For this experiment we induced both systems at a time when the cultures were located in phase of exponential growth. After inducing, the cultures were analyzed by fluorescence spectroscopy. In this analyze the cells were excitated by 500 nm. The resultant emission was measured in a spectrum between 510 and 580 nm. (Fig 10;11)
In contrast to the control of the arabinose inducible system the Lac-control also shows fluorescence. This demonstrates the leaky Lac-promotor, what is also a scientifically proven fact. A leaky promotor means that the promotor is not regulated in a leakproof manner. Even in repressed conditions the transcript can arise.
The lacking of the Lac-I gene on the vector is the reason for expression of YFP in both controls. For further research in this field it is necessary to clone the Lac-I gene inside the constructed vector.
Source
- pGA14mVenus Geneart
- Linearized BioBrick backbones
Design Notes:
We decided to construct IPTG- and arabinose-inducible systems. Therefore we amplified the promotor region and fused it to a reporter gene, which is constituted by YFP. Using this we have the option to verify the presence of the inducible promotor and also the function of the induction process by fluorescence. Our idea was to use the reporter gene as a placeholder. Via restriction enzyme digestion using the iGEM restriction enzyme site you will be able to replace the reporter gene with your gene of interest. As vector backbone we made use of the pSB1A3, which has ampicillin resistance (Fig. 8). The constructs using pSB1K3 (kanamycin resistance) and pSB1C3 (chloramphenicol resistance) are still on the anvil.
BioBrick TEV protease
Part name: BBa_K627016
Part type: Coding part
Short description: mutated TEV protease without iGEM restriction sites
General Informations
TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus (TEV). It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.
At 37°C, the TEV protease forms inclusion body, which leads to an inactive form. Incubated at 30 °C, the protease is expressed as soluble type and is highly active.
Primer used for site directed mutagenesis:
Fragment 1:
- r_TEV_ACCAGC: GGAATGTGCAGCTGGTGTCTGACACC
- f_TEV_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGAGAAAGCTTGTTTAAGGGA
Fragment 2:
- r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
- f_TEV_ACCAGC: GGTGTCAGACACCAGCTGCACATTCC
Primer used for assembly PCR of mutated fragments:
- f_TEV_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGAGAAAGCTTGTTTAAGGGA
- r_TEV_iGEM_BamHI: TATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
Primers used for amplification of pBAD arabinose-inducible induction system:
- f_AraC_iGEM_HindIII: TATAAGCTTGAATTCGCGGCCGCTTCTAGATTATGACAACTTGACGGCTACATCATT
- r_AraC_NgoMIV: ATAGCCGGCCTCCTTCTTAAAGTTAAACAAAATTATTTCTAGCCC
Primers used for amplification and modification of TEV protease:
- f_TEV_AraFusion_NgoMIV: ATATTGCCGGCATGGGAGAAAGCCTGTTTAAGGGA
- r_TEV_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTATTGCGAGTACACCAATTCATTCAT
References:
Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S. P. (2007). Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16: 2360-2367
Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D., and Waugh, D. S. (2001). Tobacco etch virus protease: Mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Prot. Eng. 14: 993-1000.
Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable form of recombinant tobacco etch virus protease. Biotechniques 30: 544-550.
BioBrick HRV14 3C protease
Part name: BBa_K627017
Part type: Coding part
Short description: HRV14 3C protease
HRV 14_3C
The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln/GlyPro.
The 22 kDa protease got its optimal activity at 4°C but also shows a high cleavage rate at 37°C. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.
General Function:
When arabinose is added to the media (2mM up to 50 mM), the CAP lost its inhibitory function and the arabinose-inducible induction system gets activated. The protein, here the HRV 14_3C protease, after the T7 RBS gets express at a very high rate. So we can assume that this system is an effectiv protease generator.
Design Notes:
This BioBrick was built by PCR using the following PCR primers:
Site directed mutagenesis of HRV 14 3C protease:
Fragment 1:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_ACCAGC: ACCATTTGCTGGTACATCATCACCAGG
Fragment 2:
- f_14_3C_ACCAGC: CCTGGTGATGATGTACCAGCAAATGGT
- r_14_3C_tm_XbaI208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC
Fragment 3:
- f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG
- r_14_3C_tm_XbaI280_A-T: ATCCACACCTTCGAGATCTTCTGATAT
Fragment 4:
- f_14_3C_tm_Xba280_A-T: ATATCAGAAGATCTCGAAGGTGTGGAT
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primers used for first assembly PCR of mutated fragments:
Fragment 5, containing Fragment 1 and 2:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_tm_Xba208_A-T: CACTGTAAGCTCAAGATTAATGTTCTC
Fragment 6, containing Fragment 3 and 4:
- f_14_3C_tm_Xba208_A-T: GAGAACATTAATCTTGAGCTTACAGTG
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primer used for final assembly PCR of mutated fragments:
Complete mutated 14_3C protease, containing Fragment 5 and 6:
- f_14_3C_iGEM: ATATAGAATTCGCGGCCGCTTCTAGATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
Primers used for amplification of the pBAD arabinose-inducible induction system:
- f_AraC_iGEM_HindIII: TATAAGCTTGAATTCGCGGCCGCTTCTAGATTATGACAACTTGACGGCTACATCATT
- r_AraC_NgoMIV: ATAGCCGGCCTCCTTCTTAAAGTTAAACAAAATTATTTCTAGCCC
Primers used for amplification and modification of HRV 14 3C protease:
- f_14_3C_AraFusion_NgoMIV: ATATTGCCGGCATGGGACCAAACACAGAATTTGCACTATCC
- r_14_3C_iGEM_BamHI: ATATAGGATCCACTGCAGCGGCCGCTACTAGTTTTGTTTCTCTACAAAATATTGTTTTTTAAGTTGAGCTGA
References:
Combination of Two Separate Binding Domains Defines Stoichiometry between Type III Secretion System Chaperone IpgC and Translocator Protein IpaB, Ravi Kumar Lokareddy, Michele Lunelli, Björn Eilers, Vivien Wolter, and Michael Kolbe, December 17, 2010 The Journal of Biological Chemistry, 285, 39965-39975.
Cleavage of Small Peptides In Vitro by Human Rhinovirus 14 3C Protease expressed in Escherichia coli, MICHAEL G. CORDINGLEY,* R. BRUCE REGISTER, PIA L. CALLAHAN, VICTOR M. GARSKY,AND RICHARD J. COLONNO JOURNAL OF VIROLOGY, Dec. 1989, p. 5037-5045
 "
"