Team:BU Wellesley Software/G-nomeSurferPro
From 2011.igem.org
| (41 intermediate revisions not shown) | |||
| Line 23: | Line 23: | ||
/*actual content styles*/ | /*actual content styles*/ | ||
| - | body {width: | + | body {width: 800px; margin:auto;} |
#bu-wellesley_wiki_content {height:auto; line-height:100%;} | #bu-wellesley_wiki_content {height:auto; line-height:100%;} | ||
| - | #bu-wellesley_wiki_content a {color:#69d01d;} | + | /*#bu-wellesley_wiki_content a {color:#69d01d;}*/ |
| - | #bu-wellesley_wiki_content a:hover {text-decoration:none; color:# | + | #bu-wellesley_wiki_content a:hover {text-decoration:none; color:#3d3f3c;} |
| - | + | .navbar li {color: #ffffff;} | |
| - | + | .navbar li a {color: #ffffff;} | |
| + | .navbar li a:hover {background:#69d01d; color: #ffffff;} | ||
| + | /*only use for current page content header (i.e. Team, G-nomeSurferPro, etc)*/ | ||
| + | H6 { | ||
| + | font-family: Helvetica; | ||
| + | text-transform: uppercase; | ||
| + | text-decoration: none; | ||
| + | text-align: left; | ||
| + | color: #3d3f3c; | ||
| + | font-size: 32pt; | ||
| + | } | ||
| + | </style> | ||
| + | <link rel="stylesheet" type="text/css" href="http://cs.wellesley.edu/~hcilab/iGEM_wiki/css/Team.css"> | ||
</head> | </head> | ||
<body class="basiclayout"> | <body class="basiclayout"> | ||
| - | |||
<div id="bu-wellesley_wiki_content"> | <div id="bu-wellesley_wiki_content"> | ||
| + | |||
<p style="text-align:center;"><a href="https://2011.igem.org/Team:BU_Wellesley_Software"><img src="http://cs.wellesley.edu/~hcilab/iGEM_wiki/images/banner.png" width="800px"></a></p> | <p style="text-align:center;"><a href="https://2011.igem.org/Team:BU_Wellesley_Software"><img src="http://cs.wellesley.edu/~hcilab/iGEM_wiki/images/banner.png" width="800px"></a></p> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | + | <ul id="nav"> | |
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Team">Team</a></li> | ||
| + | <li><a href="#">Project</a> | ||
| + | <ul> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Project_Overview">Overview</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Clotho">Clotho</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/G-nomeSurferPro">G-nome Surfer Pro</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/OptimusPrimer">Optimus Primer</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Trumpet">Trumpet</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Puppetshow">Puppetshow</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/eLabNotebook">eLabNotebook</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Wet_Lab">Wet Lab</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Downloads_and_Tutorials">Downloads and Tutorials</a></li> | ||
| + | </ul> | ||
| + | </li> | ||
| + | <li><a href="#">Process</a> | ||
| + | <ul> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Methodology">Methodology</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Safety">Safety</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Notebook">Notebook</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Outreach">Outreach</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Tips">Tips and Tricks</a></li> | ||
| + | </ul> | ||
| + | </li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Gold">Medal Fulfillment</a></li> | ||
| + | <li><a href="#">Additional Info</a> | ||
| + | <ul> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Acknowledgement">Acknowledgement</a></li> | ||
| + | <li><a href="https://2011.igem.org/Team:BU_Wellesley_Software/Social">Fun</a></li> | ||
| + | </ul> | ||
| + | </li> | ||
| + | </ul> | ||
| - | + | <br> | |
| + | <h6>G-nome Surfer Pro</h6> | ||
| - | + | <div id="tracking_nav"> | |
| + | JUMP TO...<br> | ||
| + | <a href="#bu-wellesley_wiki_content">Top</a><br> | ||
| + | <a href="#overview">Tool Overview</a><br> | ||
| + | <a href="#demo">Demo</a><br> | ||
| + | <a href="#results">Results</a><br> | ||
| + | <a href="#futurework">Future Work</a> | ||
| + | </div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<div id="overview"> | <div id="overview"> | ||
| - | <h1>Overview | + | <h1>Tool Overview</h1> |
| - | < | + | <p> |
| - | + | G-nome Surfer Pro is an integrated environment for viewing prokaryotic genomic data and literature. Users can find genes on the chromosome wheel or search for a gene name or Genbank number, access Genbank notes and publications from Pubmed and view alignment and translation from the chromosome bar. The extended desktop provides distraction free workspace for information processing and hypothesis formation. The Microsoft Surface allows users to collaborate in pairs and teams on design and research tasks. | |
| - | + | <p> | |
| - | < | + | <img style="width:800px" src="http://cs.wellesley.edu/~hcilab/iGEM_wiki/images/System/GSPro.jpg"/> |
| + | G-nome Surfer Pro is written in C# for the <a href="http://www.microsoft.com/surface/en/us/default.aspx">Microsoft Surface</a> using the <a href="http://www.microsoft.com/download/en/details.aspx?displaylang=en&id=15532">Surface SDK</a>. State is managed using the <a href="http://msdn.microsoft.com/en-us/magazine/dd419663.aspx">MVVM Architecture</a> and time consuming processes, like retrieving publications and running BLAST, are run in threads behind the main UI thread. | ||
| + | <p> | ||
| + | After selecting a genome from the tree menu the chromosome bar is populated with genes from the GenBank file. Another genome can be selected at anytime from the pull down search menu. | ||
| + | <p> | ||
| + | The chromosome bar is infinitely scrolling and position on the chromosome bar is tied into the orientation of the chromosome wheel through the MVVM architecture allowing manipulation of one to affect change of the other. GenBank files are parsed for sequence, translation and any notes, all of which are available as information objects, which are accessible from any gene on the bar. Users can take genes on the bar directly into the <a href="https://2011.igem.org/Team:BU_Wellesley_Software/OptimusPrimer">Optimus Primer </a> surface application for primer design. | ||
| + | <p> | ||
| + | Information from any gene’s GenBank file are viewable as “ScatterViewItem” objects which can be moved and re-sized on the main “ScatterView” workspace. Users can view and annotate sequence/translation information by dragging and overlaying one over the other. All information objects can be moved to and from the extended desktop by dragging or tossing or erased by tossing them off the sides of the workspace. <p> | ||
| + | Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data. | ||
| + | <p> | ||
| + | By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view. | ||
| + | <p> | ||
| + | Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
| - | <div id=" | + | |
| - | <h1> | + | <div id="demo"> |
| - | + | <h1>Demo Video</h1> | |
| - | + | <p><a href="https://github.com/igemsoftware/BU_Wellesley_Software_2011/tree/master/G-nome-Surfer-Pro"><img id="download_button" src="http://cs.wellesley.edu/~hcilab/iGEM_wiki/images/temp_download_button.jpg" width="100px"/></a> | |
| - | + | <center><iframe width="560" height="345" src="http://www.youtube.com/embed/URUkqrdFV0k" frameborder="0" allowfullscreen></iframe></center> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</div> | </div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | + | <div id="results"> | |
| - | + | <h1>Results</h1> | |
| - | + | <p> | |
| - | + | We used G-nome Surfer Pro on the Microsoft surface to easily visualize the plasmid of TB myobacterium and choose the transcription factors we would like to use in our basic device. It also is linked to GenBank files to tell us the sequence of the gene of interest and the amino acids it codes for. In addition, we looked up more information about the transcription factors such as finding it in publications from the PubMed database. This information was crucial to learn more about the transcription factors we were going to handle in the lab. | |
| - | + | </div> | |
| + | |||
| + | |||
| + | <div id="futurework"> | ||
| + | <h1>Future Work</h1> | ||
| + | <ul> | ||
| + | <li> Integrate with the Clotho database. | ||
| + | <li> Expand annotations to include GenBank notes and publications. | ||
| + | <li> Create save functionality for sessions and workspaces. | ||
| + | <li> Allow users to export publications and annotations.</ul> | ||
| + | </div> | ||
Latest revision as of 20:01, 28 September 2011
G-nome Surfer Pro
Tool Overview
G-nome Surfer Pro is an integrated environment for viewing prokaryotic genomic data and literature. Users can find genes on the chromosome wheel or search for a gene name or Genbank number, access Genbank notes and publications from Pubmed and view alignment and translation from the chromosome bar. The extended desktop provides distraction free workspace for information processing and hypothesis formation. The Microsoft Surface allows users to collaborate in pairs and teams on design and research tasks.
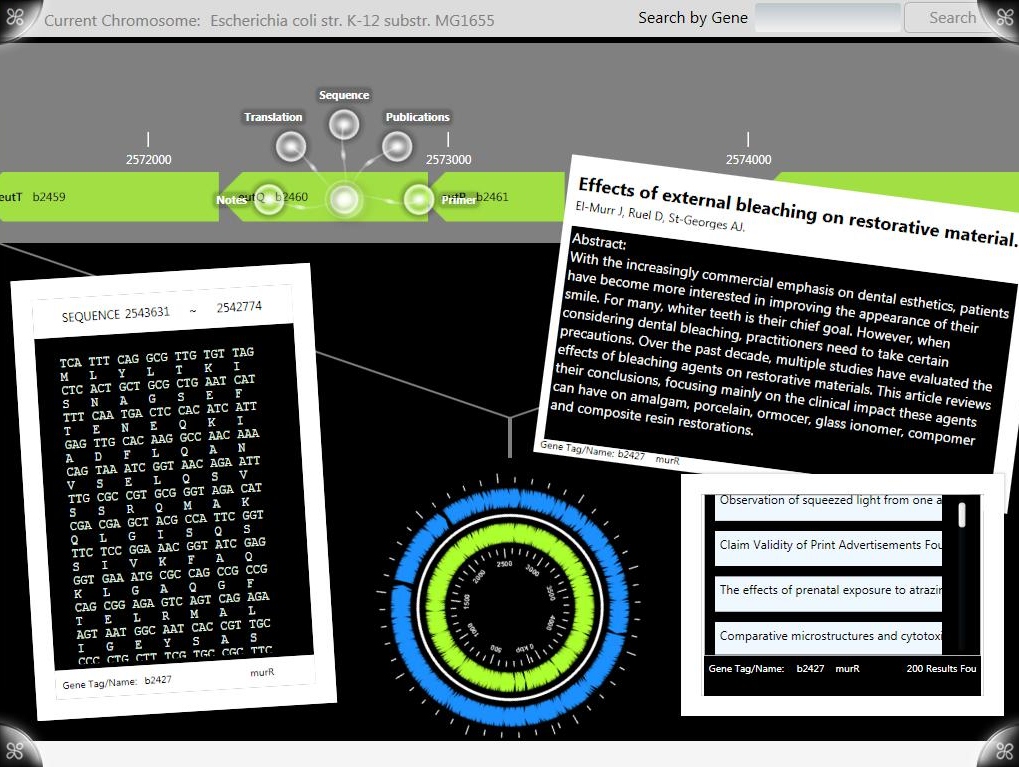 G-nome Surfer Pro is written in C# for the Microsoft Surface using the Surface SDK. State is managed using the MVVM Architecture and time consuming processes, like retrieving publications and running BLAST, are run in threads behind the main UI thread.
G-nome Surfer Pro is written in C# for the Microsoft Surface using the Surface SDK. State is managed using the MVVM Architecture and time consuming processes, like retrieving publications and running BLAST, are run in threads behind the main UI thread.
After selecting a genome from the tree menu the chromosome bar is populated with genes from the GenBank file. Another genome can be selected at anytime from the pull down search menu.
The chromosome bar is infinitely scrolling and position on the chromosome bar is tied into the orientation of the chromosome wheel through the MVVM architecture allowing manipulation of one to affect change of the other. GenBank files are parsed for sequence, translation and any notes, all of which are available as information objects, which are accessible from any gene on the bar. Users can take genes on the bar directly into the Optimus Primer surface application for primer design.
Information from any gene’s GenBank file are viewable as “ScatterViewItem” objects which can be moved and re-sized on the main “ScatterView” workspace. Users can view and annotate sequence/translation information by dragging and overlaying one over the other. All information objects can be moved to and from the extended desktop by dragging or tossing or erased by tossing them off the sides of the workspace.
Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data.
By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view.
Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data.
Results
We used G-nome Surfer Pro on the Microsoft surface to easily visualize the plasmid of TB myobacterium and choose the transcription factors we would like to use in our basic device. It also is linked to GenBank files to tell us the sequence of the gene of interest and the amino acids it codes for. In addition, we looked up more information about the transcription factors such as finding it in publications from the PubMed database. This information was crucial to learn more about the transcription factors we were going to handle in the lab.
Future Work
- Integrate with the Clotho database.
- Expand annotations to include GenBank notes and publications.
- Create save functionality for sessions and workspaces.
- Allow users to export publications and annotations.
 "
"

