Team:Tsinghua/project
From 2011.igem.org
m (→Project Overview) |
(→Project Overview) |
||
| Line 19: | Line 19: | ||
Binding | Binding | ||
| - | Src-homology 3 (SH3) domain has high affinity for proline-rich peptides and together they can form a left-handed poly-Pro type II helix, with the minimal consensus Pro-X-X-Pro. The small size and high affinity are ideal for our carrier design. In our experiment, we used a short peptide with a Kd of 3.67μM (Y. Jacquot, et al., 2007) as the binding motif and planned to transport protein substrates with this peptide sequence. | + | Src-homology 3 (SH3) domain has high affinity for proline-rich peptides and together they can form a left-handed poly-Pro type II helix, with the minimal consensus Pro-X-X-Pro. The small size and high affinity are ideal for our carrier design. In our experiment, we used a short peptide with a Kd of 3.67μM (Y. Jacquot, et al., 2007) as the binding motif and planned to transport protein substrates with this peptide sequence.[[File:SH3_domain.jpg]] |
| - | OmpA-like transmembrane domain is an evolutionarily conserved domain of outer membrane proteins. This domain consists of an eight-stranded beta barrel. OmpA is the predominant cell surface antigen in enterobacteria found in about 100,000 copies per cell. The expression of OmpA is tightly regulated by a variety of mechanisms. | + | OmpA-like transmembrane domain is an evolutionarily conserved domain of outer membrane proteins. This domain consists of an eight-stranded beta barrel. OmpA is the predominant cell surface antigen in enterobacteria found in about 100,000 copies per cell. The expression of OmpA is tightly regulated by a variety of mechanisms.[[File:Ompa-like_transmembrane_domain.jpg]] |
These two domains, as well as the proline-rich peptides are three most important parts if we want to achieve the goal of binding the target protein to the outer membrane of E.coli. First of all, we add proline-rich peptides to the target protein, in this case, mCherry which is easy to detect if the experiment is successful. Then OmpA-SH3 protein is expressed in E.coli and located onto the outer membrane so it can bind with the purified proline-rich containing mCherry protein in the medium. | These two domains, as well as the proline-rich peptides are three most important parts if we want to achieve the goal of binding the target protein to the outer membrane of E.coli. First of all, we add proline-rich peptides to the target protein, in this case, mCherry which is easy to detect if the experiment is successful. Then OmpA-SH3 protein is expressed in E.coli and located onto the outer membrane so it can bind with the purified proline-rich containing mCherry protein in the medium. | ||
| Line 31: | Line 31: | ||
It’s difficult to release the substrate from strong binding, and hence protease is called into play. HIV-protease is readily available and its high efficiency, low molecular weight and high specificity constitutes fine candidate. We only need to add the substrate sequence (HIV-protease site) between OmpA and SH3 sequence and the protease will cut off the binding cassette. In order to segregate the action, we fuse the HIV-protease with OmpA and expressed it in a second E.coli strain which we immobilize at the destination. So, when the E.coli transporter arrives at the destination, the HIV-protease that is located on the outer membrane of the second E.coli strain will help release the substrate (together with SH3 domain) by cut the HIV-protease site. | It’s difficult to release the substrate from strong binding, and hence protease is called into play. HIV-protease is readily available and its high efficiency, low molecular weight and high specificity constitutes fine candidate. We only need to add the substrate sequence (HIV-protease site) between OmpA and SH3 sequence and the protease will cut off the binding cassette. In order to segregate the action, we fuse the HIV-protease with OmpA and expressed it in a second E.coli strain which we immobilize at the destination. So, when the E.coli transporter arrives at the destination, the HIV-protease that is located on the outer membrane of the second E.coli strain will help release the substrate (together with SH3 domain) by cut the HIV-protease site. | ||
| + | [[File:SH3-HIV_protease_site-OmpA.png]] | ||
| + | [[File:SH3-HIV_protease_site-OmpA_final.png]] | ||
Revision as of 13:58, 1 September 2011
| Home | Team | Project | Parts Submitted to the Registry | Modeling | Notebook | Safety |
|---|
Project Overview
'E. Coli Transporter'
Overview
This project is destined to generate an E. Coli strain which can move forth and back to transport any desired protein along a gradient. To be specific, the bacteria can bind with the target protein at one place and then, by sensing the gradient of certain amino acid, move to its destination, where it releases the protein with the help from another E. Coli strain. When this is done, it will move back to transport more target protein. To realize our goal, four modules are necessary: binding, releasing, movement and transition.
Binding
Src-homology 3 (SH3) domain has high affinity for proline-rich peptides and together they can form a left-handed poly-Pro type II helix, with the minimal consensus Pro-X-X-Pro. The small size and high affinity are ideal for our carrier design. In our experiment, we used a short peptide with a Kd of 3.67μM (Y. Jacquot, et al., 2007) as the binding motif and planned to transport protein substrates with this peptide sequence.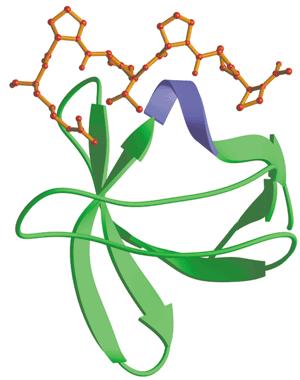
OmpA-like transmembrane domain is an evolutionarily conserved domain of outer membrane proteins. This domain consists of an eight-stranded beta barrel. OmpA is the predominant cell surface antigen in enterobacteria found in about 100,000 copies per cell. The expression of OmpA is tightly regulated by a variety of mechanisms.File:Ompa-like transmembrane domain.jpg
These two domains, as well as the proline-rich peptides are three most important parts if we want to achieve the goal of binding the target protein to the outer membrane of E.coli. First of all, we add proline-rich peptides to the target protein, in this case, mCherry which is easy to detect if the experiment is successful. Then OmpA-SH3 protein is expressed in E.coli and located onto the outer membrane so it can bind with the purified proline-rich containing mCherry protein in the medium.
So, the binding module actually comprises expression of three proteins, namely, OmpA-SH3 protein, which functions as the binding vehicle, proline-rich containing mCherry protein, which functions as the binding substrate and OmpA-mCherry protein, which functions as the positive control.
Releasing
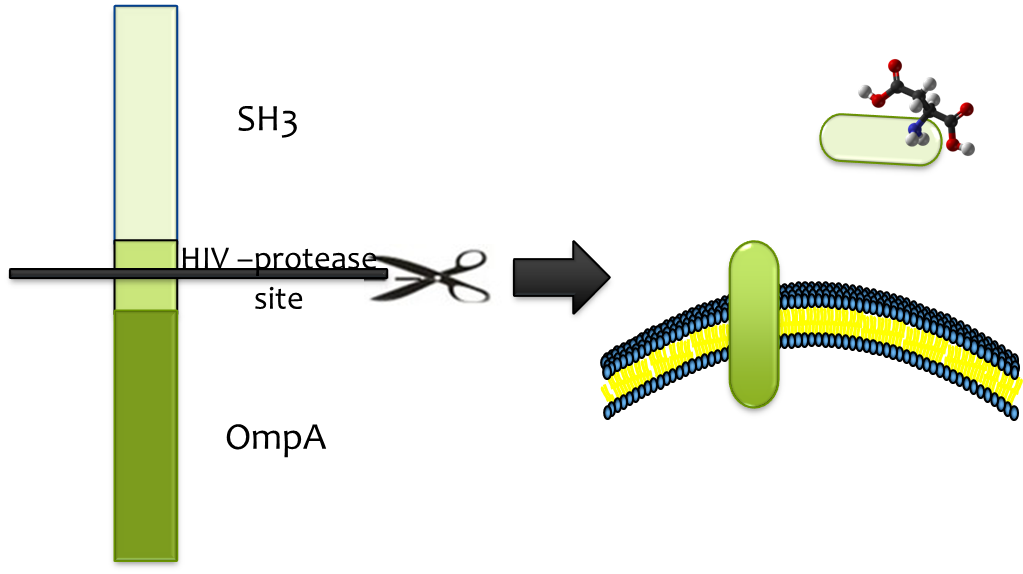
It’s difficult to release the substrate from strong binding, and hence protease is called into play. HIV-protease is readily available and its high efficiency, low molecular weight and high specificity constitutes fine candidate. We only need to add the substrate sequence (HIV-protease site) between OmpA and SH3 sequence and the protease will cut off the binding cassette. In order to segregate the action, we fuse the HIV-protease with OmpA and expressed it in a second E.coli strain which we immobilize at the destination. So, when the E.coli transporter arrives at the destination, the HIV-protease that is located on the outer membrane of the second E.coli strain will help release the substrate (together with SH3 domain) by cut the HIV-protease site.

Movement
Kinase system is well utilized in E. Coli movement control. We’ll again use LVA tagged kinase to drive E. Coli towards the target.
Transition
We need to shift between two states, the binding state and the releasing state. This molecular device is well-known for its lag in phase change, which is well adapted for our application. The lag during the phase transition is appropriate for movement and transport.
 "
"