Team:Freiburg/Notebook/1 August
From 2011.igem.org
(Difference between revisions)
SophieCramer (Talk | contribs) (→NAME OF YOUR EXPERIMENT) |
|||
| (23 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Freiburg/Templates/header}} | {{:Team:Freiburg/Templates/header}} | ||
| + | <html> | ||
| + | <div id="notebook-page-header"> | ||
| + | <div id="notebook-back" width="100px" > | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/29_July">Previous entry</a> | ||
| + | </div> | ||
| + | <div id="notebook-title"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook"> 1 August </a> | ||
| + | </div> | ||
| + | <div id="notebook-next"> | ||
| + | <a href="https://2011.igem.org/Team:Freiburg/Notebook/2_August">Next entry</a> | ||
| + | </div> | ||
| + | </div> | ||
| + | </html> | ||
| + | |||
==<span style="color:green;">green light receptor</span>== | ==<span style="color:green;">green light receptor</span>== | ||
| - | === | + | ===Qickchange PCR of CcaS=== |
| - | '''Investigators: | + | '''Investigators:Julia''' |
| + | <br/> | ||
| + | repeating experiment from___, because we had no positive transformation, probably chosen wrong annealing temperature before. | ||
| + | ==<span style="color:blue;">blue light receptor</span>== | ||
| + | ===PCR=== | ||
| - | + | '''Investigators: Sophie''' | |
| - | |||
| - | |||
| - | |||
As our last PCRs of the LovTAP with the Gibson overhangs didn't work well, we now try a PCR with primers without overhangs for Gibson assembly. | As our last PCRs of the LovTAP with the Gibson overhangs didn't work well, we now try a PCR with primers without overhangs for Gibson assembly. | ||
| Line 82: | Line 97: | ||
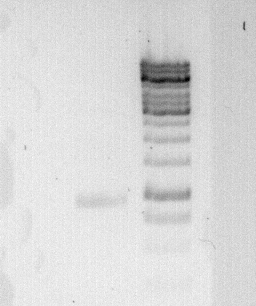
'''To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel.''' | '''To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel.''' | ||
| + | [[File:Freiburg_2011_08_01.jpg|caption|border]] | ||
'''How did you label the PCR-Product, where is it stored and what do you do next? ''' | '''How did you label the PCR-Product, where is it stored and what do you do next? ''' | ||
| Line 89: | Line 105: | ||
==<span style="color:red;">red light receptor</span>== | ==<span style="color:red;">red light receptor</span>== | ||
| - | === | + | ===3A-assembly with pcyA and the terminator BBa_1006=== |
| - | '''Investigators: | + | '''Investigators:Julia''' |
| + | <br/> | ||
| + | ====1.Digestion:<br/>==== | ||
| + | terminator (S21a), 26µl of the miniprep+ 12,1 µl H2O, cut with XbaI&PstI<br/> | ||
| + | pcyA (part BBa_I15009), 3 µl of miniprep + 35µl H2O, cut with EcoRI&SpeI<br/> | ||
| + | Vector-backbone (10 µl) pSB1T3, cut with EcoRI,PstI & DpnI<br/> | ||
| + | <br/> | ||
| + | to each reation 5µl BSA(10x) and 5µl NEB buffer4 were added, digested at 37°C for two hours, inactivated at 80°C for 20min.<br/> | ||
| + | ====2.Ligation<br/>==== | ||
| + | <br/> | ||
| + | 2µl of digested pcyA,terminator and vector were added to 11µl H2O,2µl ligase buffer and 1µl T4 ligase.<br/> | ||
| + | Incubated at room temperature for 35 min, inactivated at 80°C for 20 min.<br/> | ||
| + | ====3.Transformation==== | ||
| + | <br> | ||
| + | <br/> | ||
| + | ==<span style="color:orange;">Lysis cassette</span>== | ||
| + | ===Phage Lysis Cassette (K124017) + RBS (B0034)=== | ||
| - | + | '''Investigators: Theo''' | |
| - | + | <br> | |
| - | ''' | + | ===='''Digestion of 3A Assembly'''==== |
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Theo | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 01.08.2011 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from Experiment : - | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Phage Lysis Cassette (K124017) + RBS (B0034) | ||
| + | |||
| + | |} | ||
| + | Procedure | ||
| + | |||
| + | |||
| + | # add H<sub>2</sub>O (38μl-DNA ) | ||
| + | # 5 μl NEB4 buffer (stored at iGEM’s, -20°C) | ||
| + | # 5 μl 10x BSA (used 1:10 diluted sample stored at iGEM’s, -20°C) | ||
| + | # DNA (500 ng) (Vector:Insert ratio 1:3 in following Ligation) | ||
| + | # 1 μl restriction enzymes (stored at iGEM’s, -20°C) | ||
| + | # heat for 1-2 hours 37°C (6 hours if time) | ||
| + | # heat for 20 minutes 80°C (inactivation of enzymes) | ||
| + | # keep at 4°C if you cannot continue | ||
| + | |||
| + | Restriction enzymes you need to cut the vector, insert1 and insert 2: | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| '''Components''' | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| <center>'''Vector (μl)'''</center> | ||
| + | | colspan="3" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| <center>'''Insert1 -'''B0034-''' and 2 -'''K124017'''-(μl)'''</center> | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| DNA (500ng) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 10 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 9 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 8 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| BSA (10x) (5μl) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| NEB4 Buffer (5μl) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Enzyme 1 (1μl) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| EcoRI + DpnI | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| EcoRI | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| XbaI | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Enzyme 2 (1μl) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| PstI | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| SpeI | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| PstI | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| H<sub>2</sub>O (38 μl- DNA) | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 27 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 29 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 30 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| In total 50 μl | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | |||
| + | |} | ||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Where are the samples stored? Antibiotica resistance, vector used etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| A RBS (ribosome binding site) between the lamba cI regulated Promotor (R0051 of K098995) and the holin ATG of K124017 is absent. So a 3A assembly had to be performed in order to get a functioning Lysis cassette. | ||
| + | |||
| + | Resistance of RBS (B0034)=Cm and of Phage Lysis Cassette (K124017)=Amp, so the Vector is Tet (pSB1T3) | ||
| + | |||
| + | |} | ||
| + | |||
| + | <br> | ||
| + | <br/> | ||
| + | |||
| + | ===='''Ligation'''==== | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Theo | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 01.08.2011 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from: 01.08.2011 Phage Lysis Cassette (K124017) + RBS (B0034) Digestion | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Phage Lysis Cassette (K124017) + RBS (B0034) Ligation | ||
| + | |||
| + | |} | ||
| + | '''Procedure''' | ||
| + | |||
| + | |||
| + | PCR tube: | ||
| + | |||
| + | total volume 20 μl | ||
| + | |||
| + | |||
| + | # add H<sub>2</sub>O (17 μl -X-Y-Z) | ||
| + | # add 2 μl Ligase Buffer 10x | ||
| + | # add Insert 1, Insert 2(when proceeding from 3A digestion use 2 μl of each) | ||
| + | # add Vector (20ng needed. When proceeding from 3A digestion use 2 μl) | ||
| + | # Add 1 μl T4-DNA Ligase | ||
| + | # Incubate 10-30 min at room temperature | ||
| + | # heat for 20 minutes at 80°C | ||
| + | # store at -20°C or directly proceed to transformation | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name of part | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Ratio Insert:Vector | ||
| + | |||
| + | <nowiki>= 3:1 or 1:1</nowiki> | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Volume (μl) | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| X insert 1 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| B0034 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 3:1 (1*3) | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 3 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Y insert 2 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| K124017 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 3:1 (2*3) | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 6 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Z vector | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| pSB1T3 | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 1:3 (1) | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 1 | ||
| + | |||
| + | |- | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| H<sub>2</sub>O | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="background-color:#bfbfbf;border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| 7 | ||
| + | |||
| + | |} | ||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Where are your parts stored? Name the parts for ligation etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Ligation step of 3A assembly. | ||
| + | |||
| + | |||
| + | How to calculate ratios --> e.g. for K124017 | ||
| + | |||
| + | Length of pSB1T3= ca 2200bp | ||
| + | |||
| + | Length of K124017+Vector<nowiki>= ca 4500bp</nowiki> | ||
| + | |||
| + | 4500/2200= ca 2 | ||
| + | |||
| + | So 2*3 µl (since 3:1 is needed) = 6 µl | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |} | ||
| + | |||
| + | <br> | ||
| + | <br/> | ||
| + | |||
| + | ===='''Transformation'''==== | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Name: Theo | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Date: 01.08.2011 | ||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Continue from : 01.08.2011 Phage Lysis Cassette (K124017) + RBS (B0034) Ligation | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |- | ||
| + | | colspan="2" style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| Project Name: Phage Lysis Cassette (K124017) + RBS (B0034) | ||
| + | |||
| + | |} | ||
| + | Procedure | ||
| + | |||
| + | |||
| + | # take cells from -80°C freezer and put them on ice! (every eppi contains about 400 μl cells) | ||
| + | # thaw cells on ice 20 minutes | ||
| + | # pipette 50 μl cells and 2 μl DNA into eppi still on ice! | ||
| + | # Incubate for 30 minutes on ice | ||
| + | # Heat at 42°C for 60 sec | ||
| + | # Incubate on ice for 5 minutes | ||
| + | # Add 200 μl LB Broth | ||
| + | # Incubate for 2 hours at 37°C (cells with lysis cassette at 30°C!!) | ||
| + | # Plate 50 μl and 200μl on two different LB/Agar plates with appropriate antibiotic resistance | ||
| + | |||
| + | '''Documentation:''' | ||
| + | |||
| + | Why are you doing this experiment? Name of the sample? Where are they stored? Name the vector with inserts, antibiotika resistance etc. | ||
| + | |||
| + | |||
| + | {| style="border-spacing:0;" | ||
| + | | style="border:0.0069in solid #00000a;padding-top:0in;padding-bottom:0in;padding-left:0.075in;padding-right:0.075in;"| New Composite Part in pSB1T3 Vector with modified K124017 containing a strong RBS in front of first ATG | ||
| + | |||
| + | RESULT: No colonies!!! :( Probably because of the Tet vector, since other members of the group also had problems. We are trying it again with the pSB1C3 Vector | ||
| + | |} | ||
| + | |||
| + | <br> | ||
| + | <br/> | ||
==<span style="color:grey;">Precipitator</span>== | ==<span style="color:grey;">Precipitator</span>== | ||
| - | + | ===PCR=== | |
| Line 171: | Line 427: | ||
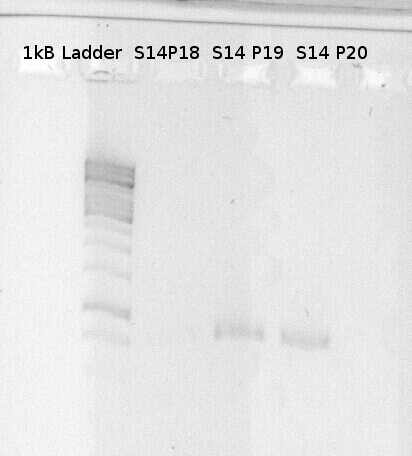
To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel. | To confirm the PCR-Product has the correct size, load 2 µl of the sample onto an agarose-gel. | ||
| - | + | [[File:Freiburg_1.8.11.jpg|capture|border]] | |
How did you label the PCR-Product, where is it stored and what do you do next? | How did you label the PCR-Product, where is it stored and what do you do next? | ||
| + | |||
| + | |||
| + | ===3A-assembly=== | ||
| + | ====1.Digestion==== | ||
| + | |||
| + | '''Investigators: Theo''' | ||
| + | |||
| + | Digestion of PCR product. | ||
| + | |||
| + | *S39 | ||
| + | *S43 | ||
| + | |||
| + | were digested with EcoRI and SpeI | ||
| + | |||
| + | *P18 | ||
| + | *P19 | ||
| + | *P20 | ||
| + | |||
| + | were digested with XbaI and PstI. | ||
| + | |||
| + | |||
| + | ====2.Ligation==== | ||
| + | |||
| + | '''Investigators: Theo''' | ||
| + | |||
| + | Ligation of: | ||
| + | |||
| + | *S39+P18 | ||
| + | *S43+P18 | ||
| + | *S39+P19 | ||
| + | *S43+P19 | ||
| + | *S39+P20 | ||
| + | *S43+P20 | ||
| + | |||
| + | stored in Theo´s box. | ||
Latest revision as of 01:03, 22 September 2011
 "
"
 Contact
Contact