Team:Washington/Protocols/CompDesign
From 2011.igem.org
| Line 15: | Line 15: | ||
[[Team:Washington/Protocols/Software|A more in depth explanation of FoldIt here.]] | [[Team:Washington/Protocols/Software|A more in depth explanation of FoldIt here.]] | ||
| - | + | ||
'''← [[Team:Washington/Protocols|Back to Protocols]]''' | '''← [[Team:Washington/Protocols|Back to Protocols]]''' | ||
Revision as of 18:24, 20 September 2011
Using FoldIt to Make CapD_CP a Better Hydrolase
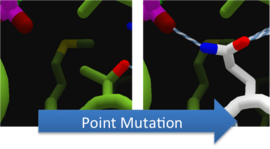
To increase the decarbonylase of Aldehyde Decarbonylase (ADC), we made point mutations to the active site. We focused our attention on two types of mutations.
First type of mutations is to increase hydrolysis by lowering the activation energy. To accomplish this, we created point mutations that can establish hydrogen bondings to a modeled transition state of our substrate. Second type of mutations concerns with the openness and polarity of the active site. To accomplish this, we mutated the active site into a more open and polar area so water molecules can enter and participate in hydrolysis easily.
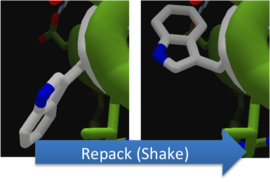
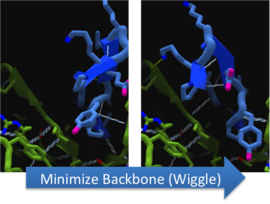
To make these point mutation designs, we used a computer program named FoldIt to predict how changes in protein structure and composition will affect protein stability. FoldIt provides a 3D representation of a protein's crystal structure that can be manipulated. Manipulation functions include point mutations, insertions, deletions, repacking of side chains (rotamer optimization), and backbone movement, which FoldIt then assesses for stability. This allows the user to quickly interact with a protein and easily predict how mutations will affect a protein.
A more in depth explanation of FoldIt here.
 "
"