Team:Washington/Magnetosomes/Results
From 2011.igem.org
| (16 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
| - | === | + | ===About the Magnetosome Toolkit:=== |
| - | + | Using standard synthetic biology protocols and the vectors we created in our Gibson Assembly Toolkit, our team was able to create a '''"Magnetosome Toolkit"''' consisting of the most basic parts required for magnetosome formation. Providing this toolkit allows future iGem teams to manipulate and understand magnetosome formation to one day create magnets in various types of bacteria. | |
| + | |||
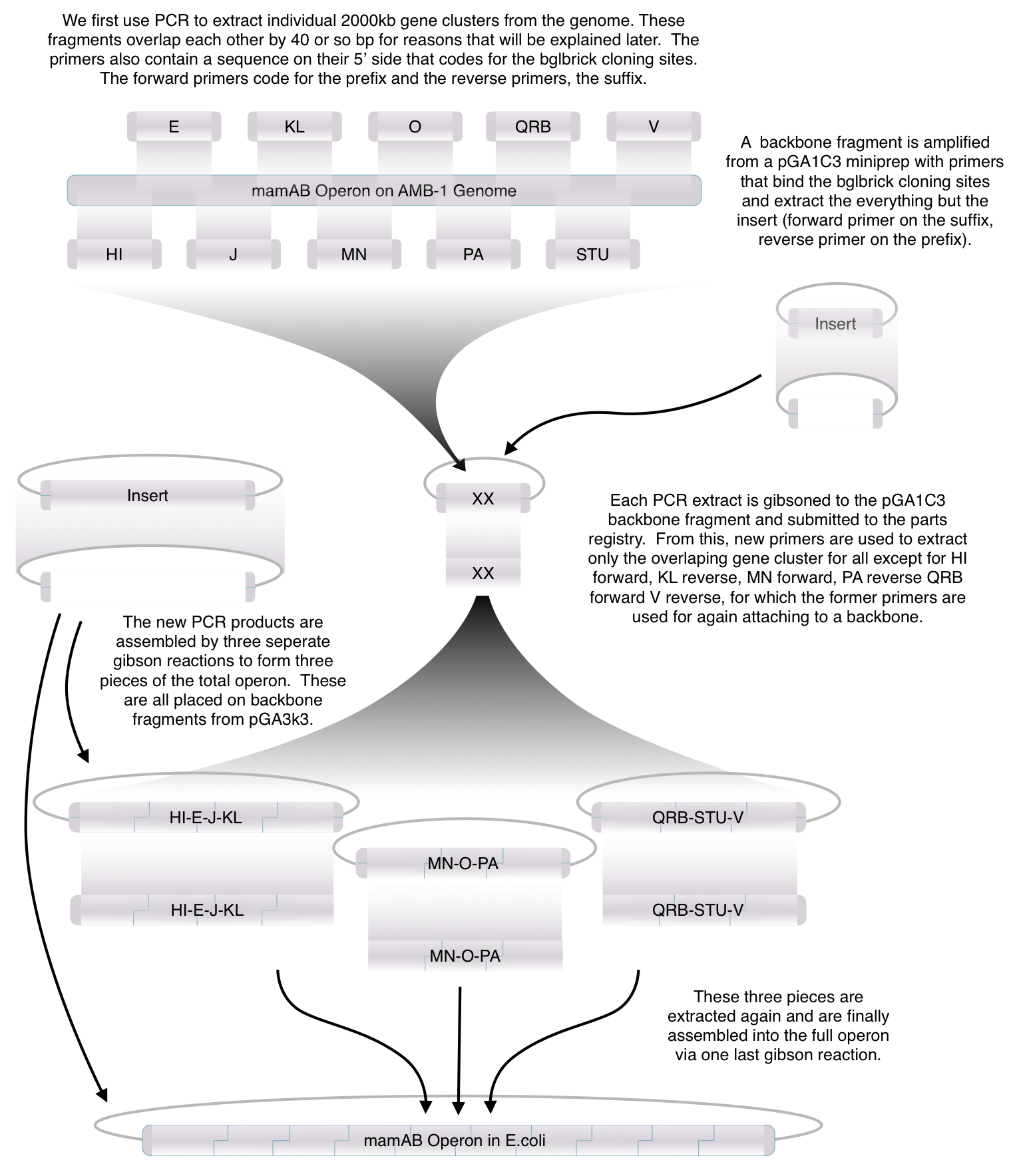
| + | === Toolkit construction and mamAB assembly in E.coli === | ||
| + | [[File:Washington Methode image.jpg|center|955px]] | ||
| + | |||
| + | ===What's in the toolkit=== | ||
| + | |||
| + | |||
| + | Before piecing together the 16 kb genome of the mamAB gene cluster within the magnetosome island (MAI), we extracted out the genes in the following group: | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |- | ||
| + | ! Gene groups | ||
| + | ! Length (bp) | ||
| + | |- | ||
| + | | mamHI | ||
| + | | 1541 | ||
| + | |- | ||
| + | | mamE | ||
| + | | 2172 | ||
| + | |- | ||
| + | | mamJ | ||
| + | | 1538 | ||
| + | |- | ||
| + | | mamKL | ||
| + | | 1336 | ||
| + | |- | ||
| + | | mamMN | ||
| + | | 2323 | ||
| + | |- | ||
| + | | mamO | ||
| + | | 1914 | ||
| + | |- | ||
| + | | mamPA | ||
| + | | 1493 | ||
| + | |- | ||
| + | | mamQRB | ||
| + | | 2029 | ||
| + | |- | ||
| + | | mamSTU | ||
| + | | 2030 | ||
| + | |- | ||
| + | | mamV | ||
| + | | 1002 | ||
| + | |- | ||
| + | |}. | ||
| + | |||
| + | Using standard PCR protocols, these genes were extracted and..... | ||
| + | These genes were visualized on gels and sequence confirmed; therefore, we can be sure that these are some of the many genes required for proper magnetosome formation. | ||
| + | |||
| + | As previously noted, magnetosome formation within the host-organism, ''Magnetospirillium magneticum'', strain AMB-1, is a highly regulated step-wise process. As shown in (link: figure #), genes encode for an invagination in the inner membrane, there are genes which help align the magnetosomes into their characteristics chains, and there are genes which regulate the biomineralization of magnetic particles. Our team chose to focus on genes specifically related to magnetosome scaffolding/alignment, since it has many practical uses in synthetic biology (??? maybe we could make a link to the alkenes future if they wanted to use mamI....) | ||
| + | |||
| + | |||
| + | Our genes of interest were mamK and mamI as they have functions related to localization of the magnetosome. Specifically, mamK is a bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain. mamK is also shown to localize the mamI, which is loss inhibits membrane formation. | ||
| + | For more information, refer to the table below: | ||
| + | |||
| + | {| class="wikitable" | ||
| + | |- | ||
| + | ! Gene | ||
| + | ! AMB Number | ||
| + | ! Cluster Membership | ||
| + | ! Member of 28 genes list? (specific*/related**) | ||
| + | ! Function Summary (Vesicle chain formation, and/or biomineralization) | ||
| + | ! Gene Function | ||
| + | |- | ||
| + | | mamH | ||
| + | | amb0961 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | |- | ||
| + | | mamI | ||
| + | | amb0962 | ||
| + | | mamAB | ||
| + | | Specific | ||
| + | | Vesicle, (Chain Formation?) | ||
| + | | >berkeley 2010: Loss causes no membrane formation, is localized onto chains | ||
| + | |- | ||
| + | | mamE | ||
| + | | amb0963 | ||
| + | | mamAB; mam Islet | ||
| + | | Related | ||
| + | | | ||
| + | | >Membrane-bound serine protease required for magnetite formation; might control the localization of other magnetosome proteins | ||
| + | |- | ||
| + | | mamJ | ||
| + | | amb0964 | ||
| + | | mamAB; mam Islet | ||
| + | | Specific | ||
| + | | Chain Formation | ||
| + | | >Proper magnetosome chain organization/assembly | ||
| + | |- | ||
| + | | mamK | ||
| + | | amb0965 | ||
| + | | mamAB; mam Islet | ||
| + | | Related | ||
| + | | Chain Formation | ||
| + | | >required for proper magnetosome chain organization; *bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain, shown to localize the mamI | ||
| + | |- | ||
| + | | mamL | ||
| + | | amb0966 | ||
| + | | mamAB; mam Islet | ||
| + | | Specific | ||
| + | | Vesicle, biomineralization | ||
| + | | >berkely 2010: Crucial to mangneosome membrane creation, shown to be spread across the cell membrane and sometimes forms lines | ||
| + | |- | ||
| + | | mamM | ||
| + | | amb0967 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | | ||
| + | | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | ||
| + | |- | ||
| + | | mamN | ||
| + | | amb0968 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | | ||
| + | | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | ||
| + | |- | ||
| + | | mamO | ||
| + | | amb0969 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | | ||
| + | | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | ||
| + | |- | ||
| + | | mamP | ||
| + | | amb0970 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | Biomineralization | ||
| + | | >berkeley 2010: loss causes weak magnetic response, with large but fewer crystals | ||
| + | |- | ||
| + | | mamA | ||
| + | | amb0971 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | | ||
| + | | >Required for magnetosome activation; activation of vessicles | ||
| + | |- | ||
| + | | mamQ | ||
| + | | amb0972 | ||
| + | | mamAB; mam Islet | ||
| + | | Related | ||
| + | | | ||
| + | | >ORF; formation/maintenance of magnetosome membranes | ||
| + | |- | ||
| + | | mamR | ||
| + | | amb0973 | ||
| + | | mamAB | ||
| + | | Specific | ||
| + | | Chain formation, Biomineralization | ||
| + | | >ORF; plays a role in controlling both particle number and size of magnetite cyrstals | ||
| + | |- | ||
| + | | mamB | ||
| + | | amb0974 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | Vesicle, Biomineralization | ||
| + | | >indirect role in magnetosome membrane invagination and biomineralization; magnetosome compartment formation | ||
| + | |- | ||
| + | | mamS | ||
| + | | amb0975 | ||
| + | | mamAB | ||
| + | | Specific | ||
| + | | | ||
| + | | | ||
| + | |- | ||
| + | | mamT | ||
| + | | amb0976 | ||
| + | | mamAB | ||
| + | | Specific | ||
| + | | Biomineralization | ||
| + | | >magnetite crystal growth; participates in different steps during magnetite synthesis | ||
| + | |- | ||
| + | | mamU | ||
| + | | amb0977 | ||
| + | | mamAB | ||
| + | | Related | ||
| + | | | ||
| + | | | ||
| + | |- | ||
| + | | mamV | ||
| + | | amb0978 | ||
| + | | mamAB | ||
| + | | N/A | ||
| + | | | ||
| + | | | ||
| + | |- | ||
| + | |}. | ||
| + | <br> | ||
| + | Using our two genes of interest, we wanted to create C-terminal sfGFP fusions so we could track the localization of each gene within E.coli. | ||
| + | |||
| + | To begin, we preformed a Gibson reaction using our genes from the Magnetosome toolkit, sfGFP, and a 1C3 vector with a pLac promoter. After the standard protocols of Gibson, transformation and plating, we imaged the samples and were able to see that the gene localizations in the strain AMB-1 were also shown in ''E.coli.'' | ||
| + | |||
| + | {| border="1" | ||
| + | |+ sfGFP fusions of mamK and MamI in both AMB-1 and ''E.coli''. | ||
| + | ! | ||
| + | ! scope="col" | Strain AMB-1 | ||
| + | ! scope="col" | ''E.coli'' | ||
| + | |- | ||
| + | ! scope="row" | mamK-sfGFP | ||
| + | | Cell 2 || Cell 3 | ||
| + | |- | ||
| + | ! scope="row" | mamI-sfGFP | ||
| + | | Cell B | ||
| + | | Cell C | ||
| + | |} | ||
Latest revision as of 19:50, 19 September 2011
About the Magnetosome Toolkit:
Using standard synthetic biology protocols and the vectors we created in our Gibson Assembly Toolkit, our team was able to create a "Magnetosome Toolkit" consisting of the most basic parts required for magnetosome formation. Providing this toolkit allows future iGem teams to manipulate and understand magnetosome formation to one day create magnets in various types of bacteria.
Toolkit construction and mamAB assembly in E.coli
What's in the toolkit
Before piecing together the 16 kb genome of the mamAB gene cluster within the magnetosome island (MAI), we extracted out the genes in the following group:
| Gene groups | Length (bp) |
|---|---|
| mamHI | 1541 |
| mamE | 2172 |
| mamJ | 1538 |
| mamKL | 1336 |
| mamMN | 2323 |
| mamO | 1914 |
| mamPA | 1493 |
| mamQRB | 2029 |
| mamSTU | 2030 |
| mamV | 1002 |
Using standard PCR protocols, these genes were extracted and..... These genes were visualized on gels and sequence confirmed; therefore, we can be sure that these are some of the many genes required for proper magnetosome formation.
As previously noted, magnetosome formation within the host-organism, Magnetospirillium magneticum, strain AMB-1, is a highly regulated step-wise process. As shown in (link: figure #), genes encode for an invagination in the inner membrane, there are genes which help align the magnetosomes into their characteristics chains, and there are genes which regulate the biomineralization of magnetic particles. Our team chose to focus on genes specifically related to magnetosome scaffolding/alignment, since it has many practical uses in synthetic biology (??? maybe we could make a link to the alkenes future if they wanted to use mamI....)
Our genes of interest were mamK and mamI as they have functions related to localization of the magnetosome. Specifically, mamK is a bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain. mamK is also shown to localize the mamI, which is loss inhibits membrane formation.
For more information, refer to the table below:
| Gene | AMB Number | Cluster Membership | Member of 28 genes list? (specific*/related**) | Function Summary (Vesicle chain formation, and/or biomineralization) | Gene Function |
|---|---|---|---|---|---|
| mamH | amb0961 | mamAB | Related | ||
| mamI | amb0962 | mamAB | Specific | Vesicle, (Chain Formation?) | >berkeley 2010: Loss causes no membrane formation, is localized onto chains |
| mamE | amb0963 | mamAB; mam Islet | Related | >Membrane-bound serine protease required for magnetite formation; might control the localization of other magnetosome proteins | |
| mamJ | amb0964 | mamAB; mam Islet | Specific | Chain Formation | >Proper magnetosome chain organization/assembly |
| mamK | amb0965 | mamAB; mam Islet | Related | Chain Formation | >required for proper magnetosome chain organization; *bacterial actin-like cytoskeleton protein required for proper alignment of the magnetosomes in a chain, shown to localize the mamI |
| mamL | amb0966 | mamAB; mam Islet | Specific | Vesicle, biomineralization | >berkely 2010: Crucial to mangneosome membrane creation, shown to be spread across the cell membrane and sometimes forms lines |
| mamM | amb0967 | mamAB | Related | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | |
| mamN | amb0968 | mamAB | Related | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | |
| mamO | amb0969 | mamAB | Related | >biomineralization, involved in iron transport, magnetite nucleation, or establishement of the proper chemical enviornment for magnetite synthesis in the magnetosome | |
| mamP | amb0970 | mamAB | Related | Biomineralization | >berkeley 2010: loss causes weak magnetic response, with large but fewer crystals |
| mamA | amb0971 | mamAB | Related | >Required for magnetosome activation; activation of vessicles | |
| mamQ | amb0972 | mamAB; mam Islet | Related | >ORF; formation/maintenance of magnetosome membranes | |
| mamR | amb0973 | mamAB | Specific | Chain formation, Biomineralization | >ORF; plays a role in controlling both particle number and size of magnetite cyrstals |
| mamB | amb0974 | mamAB | Related | Vesicle, Biomineralization | >indirect role in magnetosome membrane invagination and biomineralization; magnetosome compartment formation |
| mamS | amb0975 | mamAB | Specific | ||
| mamT | amb0976 | mamAB | Specific | Biomineralization | >magnetite crystal growth; participates in different steps during magnetite synthesis |
| mamU | amb0977 | mamAB | Related | ||
| mamV | amb0978 | mamAB | N/A |
Using our two genes of interest, we wanted to create C-terminal sfGFP fusions so we could track the localization of each gene within E.coli.
To begin, we preformed a Gibson reaction using our genes from the Magnetosome toolkit, sfGFP, and a 1C3 vector with a pLac promoter. After the standard protocols of Gibson, transformation and plating, we imaged the samples and were able to see that the gene localizations in the strain AMB-1 were also shown in E.coli.
| Strain AMB-1 | E.coli | |
|---|---|---|
| mamK-sfGFP | Cell 2 | Cell 3 |
| mamI-sfGFP | Cell B | Cell C |
 "
"