Team:Washington/Celiacs/Methods
From 2011.igem.org
(→Using Foldit to Design Mutations) |
(→Using Foldit to Design Mutations) |
||
| Line 6: | Line 6: | ||
==Using Foldit to Design Mutations== | ==Using Foldit to Design Mutations== | ||
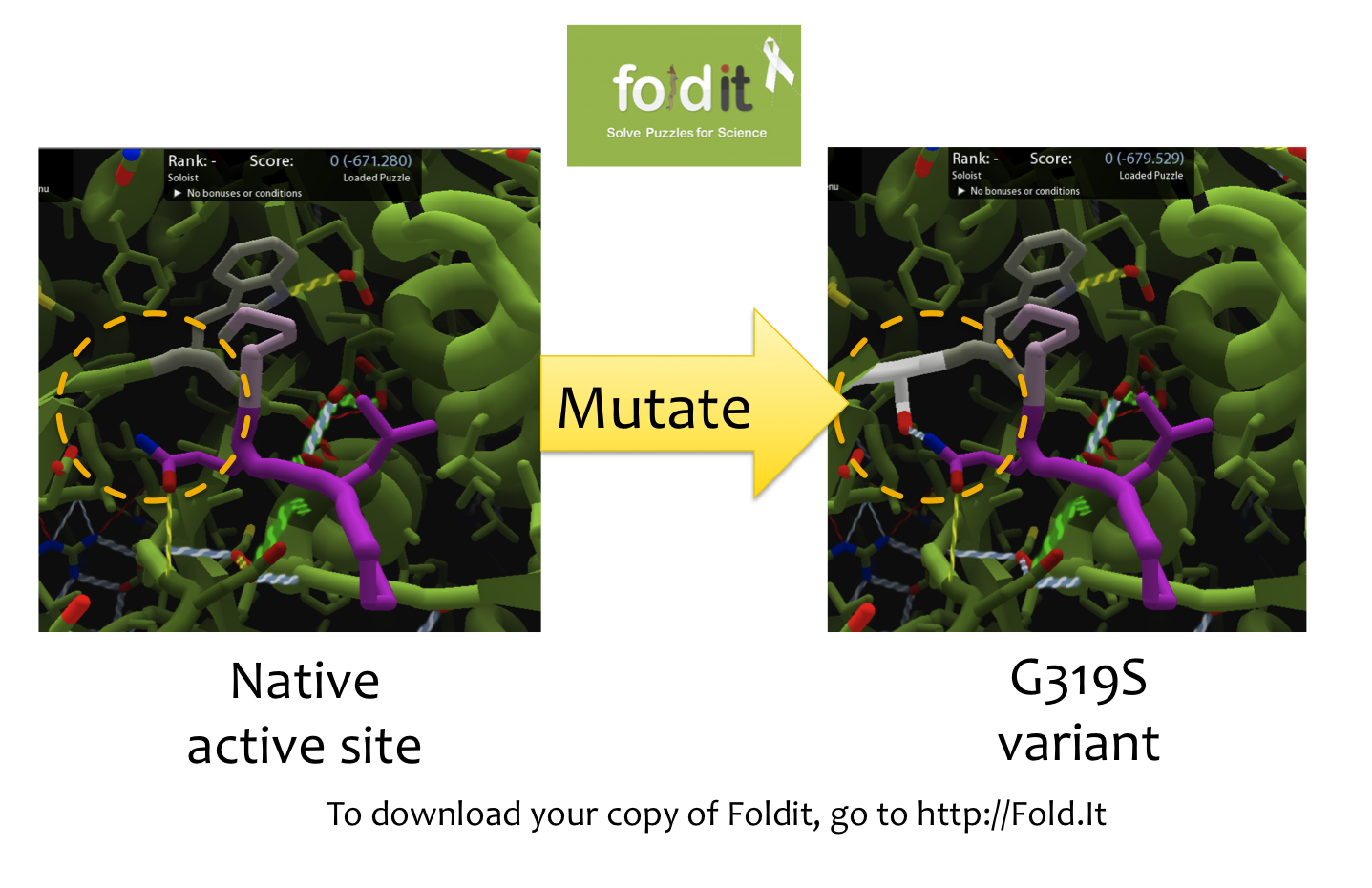
| - | In order to design mutations to wild-type Kumamolisin that would increase the enzyme’s proteolytic activity on gluten, we used a computational enzyme editing program called Foldit, which allows the user to hypothetically modify the amino acid sequence of a protein by creating point mutations at any location within the protein’s crystal structure. Within Foldit, we loaded Kumamolisin’s crystal structure in complex with a model PQLP peptide that recurs frequently in gluten, thus mimicking gluten as a substrate. We then modified the amino acid residues around the active site of Kumamolisin in the crystal structure, attempting to decrease the free energy of, and thus stabilize, the system | + | In order to design mutations to wild-type Kumamolisin that would increase the enzyme’s proteolytic activity on gluten, we used a computational enzyme editing program called Foldit, which allows the user to hypothetically modify the amino acid sequence of a protein by creating point mutations at any location within the protein’s crystal structure. Within Foldit, we loaded Kumamolisin’s crystal structure in complex with a model PQLP peptide that recurs frequently in gluten, thus mimicking gluten as a substrate. We then modified the amino acid residues around the active site of Kumamolisin in the crystal structure, attempting to decrease the free energy of, and thus stabilize, the system. Estimations of free energy were based on algorithms run by Foldit. |
Using this method, we designed over 100 novel mutants, each of which could potentially increase Kumamolisin’s proteolytic activity on gluten. | Using this method, we designed over 100 novel mutants, each of which could potentially increase Kumamolisin’s proteolytic activity on gluten. | ||
Revision as of 21:01, 14 September 2011
Redesigning Kumamolisin to Have Higher Activity at Low pH
Using Foldit to Design Mutations
In order to design mutations to wild-type Kumamolisin that would increase the enzyme’s proteolytic activity on gluten, we used a computational enzyme editing program called Foldit, which allows the user to hypothetically modify the amino acid sequence of a protein by creating point mutations at any location within the protein’s crystal structure. Within Foldit, we loaded Kumamolisin’s crystal structure in complex with a model PQLP peptide that recurs frequently in gluten, thus mimicking gluten as a substrate. We then modified the amino acid residues around the active site of Kumamolisin in the crystal structure, attempting to decrease the free energy of, and thus stabilize, the system. Estimations of free energy were based on algorithms run by Foldit.
Using this method, we designed over 100 novel mutants, each of which could potentially increase Kumamolisin’s proteolytic activity on gluten.
Mutagenizing Kumamolisin
Producing ssDNA
Once we had obtained feasible mutants from the Foldit program, we ordered these mutations in oligonucleotide primers to be used in Kunkel Mutagenesis. Kunkel mutagenesis is a short and simple procedure, so it was ideal for producing many mutants at a time To perform the mutagenesis, we first transformed the plaasmid containing wild-type kumamolisin into chemically competent cells of Escherichia coli CJ236. These cells were then grown and harvested for their single stranded DNA (ssDNA). We then determined the concentration of the DNA using a nanodrop machine. This was done to ensure that the DNA was at a high enough concentration to be used for mutagenesis.
Kunkel Mutagenesis
Using ssDNA as a template, we annealed oligo nucleotides in a PCR block to increase specific binding. The primer was then extended on the ssDNA using T7 DNA polymerase. We then dialyzed the DNA to remove salts introduced in previous steps. We then added the DNA to cuvettes containing electrically competent E. coli BL21* cells. These cells were then pulsed with electricity to allow the uptake of the newly mutagenized DNA. After growing the cells for an hour in TB, we plated the cells on LB agar containing kanamycin.
Using a Whole Cell Lysate Assay to Test Activity of Mutants
After the cells had been allowed to grow overnight, colonies were picked and used to inoculate a 96 well plate containing LB and kanamycin. This step allowed us to grow a representative amount of cells containing each mutation. After growing overnight at 37 degrees celcius cells from each well were transferred to another 96 well plate containing TB and kanamycin. These plates were incubated at 37 degrees celcius and later induced using IPTG. After induction, we incubated the plates at 18 degrees celcius overnight. We then lysed the cells and tested the supernatant for proteolytic activity towards PQLP in an assay which measured PQLP degradation over a period of 30 minutes. The assay was done at pH 4 in accordance with the assays done to test ScPEP according to the literature. The mutants were tested against wild-type kumamolisin and ScPEP, an enzyme currently used for the treatment of gluten intolerance via proteolysis. The assay we used was not highly accurate in terms of actual activity. However, what the assay allowed us to do was determine activity relative to our controls. This allowed us to determine which mutants were worth purifying to get more accurate activity data. File:Washington Assay.png
Testing Purified Mutants to Accurately Assess Activity
Purification
After compiling a set of mutants which showed a relative increase in activity we proceeded to purify our mutant proteins. This step is crucial because it allows us to determine how our mutant compares with the wild-type on a quantitative level. For instance, if the whole cell lysate assay showed one mutant to have ten times the activity level as wild-type kumamolisin we cannot assume that there has been an increase in activity because there could simply be ten times the amount of protein with the same level of activity. To purify our mutants, we first grew them in TB and kanamycin with a single colony of the mutant. This inoculation was grown over 24 hours at 37 degrees celcius and then expanded to 50 mililiters (TB+kanamycin). We then allowed the culture to grow to a specific optical density of cells before inducing the culture using IPTG. We then allowed the cells to grow again to express kumamolisin. We then lysed the cells using a lysis buffer and centrifugation. This allowed the enzymes to be released from the cells. The proteins were then collected on a column, washed with buffer, and eluted off the column. Finally, wwe dialyzed the protein in a sodium acetate buffer (pH 4).
Assay
Once we had pure protein, we determined the concentration of each using a NanoDrop machine. With the concentration of each, we then diluted the proteins to the same concentration. For the assay, we also used purified wild-type kumamolisin and ScPEP. The assay was again run for 30 minutes at pH 4. Using the data from the purified assays, we were able to deermine which mutants had higher activity than kumamolisin and by how much their activity was greater.
 "
"