|
Stop-Codon Switch
Background
The Rare Codon Switch can only regulate the amount of target protein production. To make our device a strict molecular switch that turns on/off protein biosynthesis, we need to eliminate the background.
In recent years, stop codons (codonST) and stop codon suppressor tRNAs (tRNASS) are used to incorporate unnatural amino acids in protein biosynthesis. Researchers like Peter Schultz and Kiga have made a lot of contributions in this field. The advantage of codonST and tRNASS is that they can incorporate target amino acids without background noise. We make use of codonST and tRNASS as the controlling element for protein biosynthesis.
Introduction
We design a Stop-Codon Switch to turn on/off protein translation.
We can control the translation process by controlling whether the ribosome can get through the stop codon placed in the target protein's mRNA.
This process can be achieved by controlling the existence of charged tRNASS that recognizes the stop codon. This process is controlled by two elements:
The existence of tRNASS
aminoacyl tRNA synthetases (aaRS) that can charge tRNASS
Reporter: a stop codon is put immediately after the initial codon ATG in the target protein' s mRNA.
Design
- tRNAAsp-TAG: tRNAAsp-TAG with its anticodon mutated to CUA can base pair with stop codon UAG.
- aaRS: the modified AspRS without anticodon recognition domain. This modified enzyme can charge Asp to tRNAAsp-TAG.
Click here to see Modeling.
- Luciferase-TAG: A stop codon TAG is put immediately after the start codon ATG in the luciferase gene.
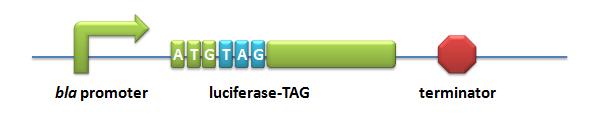
Action: If the ribosome can get through the stop codon with the help of Stop-Codon Switch, luciferase can be expressed. Otherwise, luciferase cannot be expressed.
Results
We have used Pbla-Luc-TAG ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567003 BBa_K567003]) as our Reporter. The amount of luciferase produced is reflected using the bioluminescence emitted during the luciferin reaction. Our results demonstrate that TAG insertion into luciferase blocks luciferase production, which was shown in the control group. In the experimental group, with the help of PT7-TDRS([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567011 BBa_K567011]) and tRNAAsp-TAG([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567013 BBa_K567013]),luciferase was produced and bioluminescence was emitted during the luciferin reaction. These results proved that Stop-Codon Switch can turn on protein expression. Yet further work is needed to optimize the device.
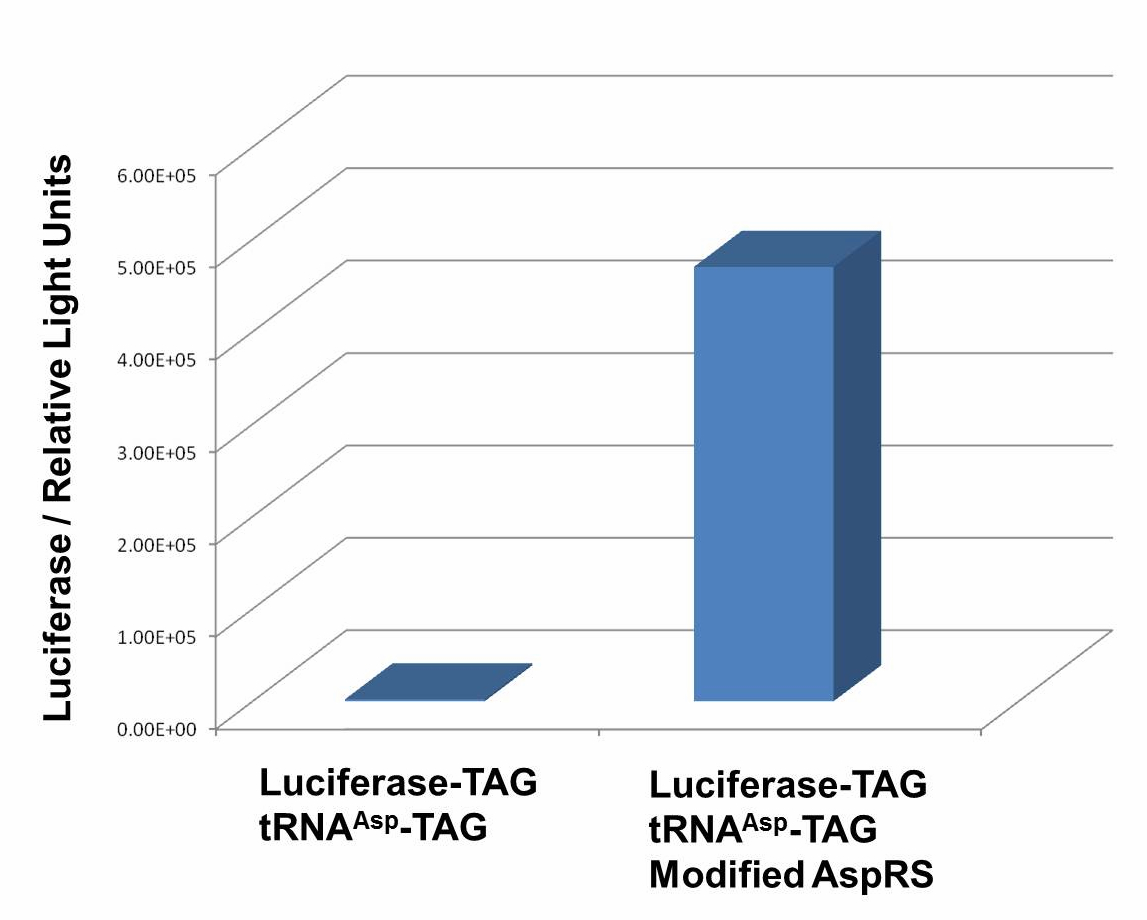 Fig 1. Functional Analysis of Stop-Codon Switch. ER2566 cannot produce luciferase with Pbla-Luc-TAG ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567003 BBa_K567003]) only. When PT7-TDRS ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567011 BBa_K567011]) and tRNA Asp-TAG ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K567013 BBa_K567013]) are also transformed into the cell, luciferase is produced. The results proved Stop-Codon Switch as a strict molecular switch without background noise. Conclusions
We have successfully constructed the Stop-Codon Switch and have tested the device. Results proved Stop-Codon Switch as a strict molecular switch without background noise. tRNAAsp-TAG recognizes stop codon TAG and can be charged with Asp by modified AspRS.
Referance
1.Cropp, T.A. and P.G. Schultz, An expanding genetic code. Trends Genet, 2004. 20(12): p. 625-30.
2.Liu, C.C., et al., Efficient expression of tyrosine-sulfated proteins in E. coli using an expanded genetic code. Nat Protoc, 2009. 4(12): p. 1784-9.
3.Liu, C.C. and P.G. Schultz, Adding new chemistries to the genetic code. Annu Rev Biochem. 79: p. 413-44.
4.Liu, D.R., T.J. Magliery, and P.G. Schultz, Characterization of an 'orthogonal' suppressor tRNA derived from E. coli tRNA2(Gln). Chem Biol, 1997. 4(9): p. 685-91.
5.Ryu, Y. and P.G. Schultz, Efficient incorporation of unnatural amino acids into proteins in Escherichia coli. Nat Methods, 2006. 3(4): p. 263-5.
6.Wang, F., et al., Genetic incorporation of unnatural amino acids into proteins in Mycobacterium tuberculosis. PLoS One. 5(2): p. e9354.
7.Wang, L., et al., Expanding the genetic code of Escherichia coli. Science, 2001. 292(5516): p. 498-500.
8.Wang, L. and P.G. Schultz, A general approach for the generation of orthogonal tRNAs. Chem Biol, 2001. 8(9): p. 883-90.
9.Wang, L. and P.G. Schultz, Expanding the genetic code. Chem Commun (Camb), 2002(1): p. 1-11.
10.Wang, L., J. Xie, and P.G. Schultz, Expanding the genetic code. Annu Rev Biophys Biomol Struct, 2006. 35: p. 225-49.
11.Xie, J. and P.G. Schultz, Adding amino acids to the genetic repertoire. Curr Opin Chem Biol, 2005. 9(6): p. 548-54.
12.Xie, J. and P.G. Schultz, An expanding genetic code. Methods, 2005. 36(3): p. 227-38.
13.Young, T.S. and P.G. Schultz, Beyond the canonical 20 amino acids: expanding the genetic lexicon. J Biol Chem. 285(15): p. 11039-44.
14.Zhang, Y., et al., Crystal structures of apo wild-type M. jannaschii tyrosyl-tRNA synthetase (TyrRS) and an engineered TyrRS specific for O-methyl-L-tyrosine. Protein Sci, 2005. 14(5): p. 1340-9.
|