|
|
| (6 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| | <div id="614" style="display:none"> | | <div id="614" style="display:none"> |
| | == June 14 == | | == June 14 == |
| | + | ===Wet Lab=== |
| | *Made four LB-based media solutions, and later created glycerol stocks from these and placed in -80⁰C freezer | | *Made four LB-based media solutions, and later created glycerol stocks from these and placed in -80⁰C freezer |
| | **Selection strain (ΔHis3ΔPyrFΔrpoZ) in 3 mL LB and 3µL of 1000x Tet solution stock | | **Selection strain (ΔHis3ΔPyrFΔrpoZ) in 3 mL LB and 3µL of 1000x Tet solution stock |
| Line 54: |
Line 55: |
| | We finished writing the generate function, and now have a working sequence generator. We also began more in-depth research into the 2011 work by Persikov which deals with how the zinc finger binds to DNA. He predicts several relations which we should be able to test. | | We finished writing the generate function, and now have a working sequence generator. We also began more in-depth research into the 2011 work by Persikov which deals with how the zinc finger binds to DNA. He predicts several relations which we should be able to test. |
| | | | |
| - | [[File:HARVInteraction Map.png|frame|right|Proposed novel interactions between helical zinc finger residues and base pairs of the target DNA sequence (based on Persikov 2011 <cite>Persikov2011</cite>]] | + | [[File:HARVInteraction Map.png|frame|right|Proposed novel interactions between helical zinc finger residues and base pairs of the target DNA sequence (based on [http://www.ncbi.nlm.nih.gov/pubmed/21572177 Persikov 2011])]] |
| | | | |
| | Persikov sent us his SVM code (used to calculate the probability of a sequence binding to given DNA), so we also worked on adapting this to use when narrowing our sequences to those most likely to work. | | Persikov sent us his SVM code (used to calculate the probability of a sequence binding to given DNA), so we also worked on adapting this to use when narrowing our sequences to those most likely to work. |
| Line 68: |
Line 69: |
| | <div id="615" style="display:none"> | | <div id="615" style="display:none"> |
| | | | |
| - | ==June 15th== | + | ==June 15th - Wet Lab== |
| | '''Gradient PCR:''' 5µL of each PCR product were run on a 1% gel. No bands appeared: the PCR appears to not have worked. | | '''Gradient PCR:''' 5µL of each PCR product were run on a 1% gel. No bands appeared: the PCR appears to not have worked. |
| | | | |
| Line 147: |
Line 148: |
| | | | |
| | *Justin continued work on a sequence-finding program, the most up to date version can be found in the Dropbox under code/zfsitefinder.html. | | *Justin continued work on a sequence-finding program, the most up to date version can be found in the Dropbox under code/zfsitefinder.html. |
| - | *Justin and Will found 10 candidate sequences across 4 diseases that hopefully should encompass a good amount of diversity in terms of expanding the ZF library. These sequences can be found in the table below, with more details [[File:HARVZF_Binding_Sequence_Candidates.xls|here]]. | + | *Justin and Will found 10 candidate sequences across 4 diseases that hopefully should encompass a good amount of diversity in terms of expanding the ZF library. These sequences can be found in the table below, with more details [https://2011.igem.org/File:HARVZF_Binding_Sequence_Candidates.xls here]. |
| | | | |
| | {| class="wikitable" cellpadding="5" | | {| class="wikitable" cellpadding="5" |
| Line 242: |
Line 243: |
| | <div id="616" style="display:none"> | | <div id="616" style="display:none"> |
| | | | |
| - | ==June 16th== | + | ==June 16th - Wet Lab== |
| | | | |
| - | *So there was totally a crazy bee hive outside today!! | + | *There was totally a crazy bee hive outside today!! |
| | | | |
| | '''Glycerol Stock pKD42''' | | '''Glycerol Stock pKD42''' |
| Line 388: |
Line 389: |
| | ***We can now create lists of the zinc fingers that bind to any triplet, and create interaction matrices and frequency tables for any triplet input.</div> | | ***We can now create lists of the zinc fingers that bind to any triplet, and create interaction matrices and frequency tables for any triplet input.</div> |
| | <div id="617" style="display:none"> | | <div id="617" style="display:none"> |
| - | ==June 17th== | + | ==June 17th - Wet Lab== |
| | '''Update on selection strain knockout status:''' We are trying to reach Addgene to check how His3 was knocked out---instead of deleting the gene, they may have simply introduced an early stop codon. If that's the case, our gel would have the correct bands because the primers we designed can only show whether a deletion or insertion was in that locus. | | '''Update on selection strain knockout status:''' We are trying to reach Addgene to check how His3 was knocked out---instead of deleting the gene, they may have simply introduced an early stop codon. If that's the case, our gel would have the correct bands because the primers we designed can only show whether a deletion or insertion was in that locus. |
| | | | |
| Line 418: |
Line 419: |
| | ***Graph: # of var (# of tries by the computer) vs. % space covered | | ***Graph: # of var (# of tries by the computer) vs. % space covered |
| | | | |
| - | [[File:HARVInteraction Map.png|frame|right|Proposed interactions between helical zinc finger residues and base pairs of the target DNA sequence (based on Persikov 2011 <cite>Persikov2011</cite>]] | + | [[File:HARVInteraction Map.png|frame|right|Proposed interactions between helical zinc finger residues and base pairs of the target DNA sequence (based on [http://www.ncbi.nlm.nih.gov/pubmed/21572177 Persikov 2011])]] |
| | | | |
| | ====Options for Target DNA Sequences / ZF Helices==== | | ====Options for Target DNA Sequences / ZF Helices==== |
| Line 440: |
Line 441: |
| | ---- | | ---- |
| | ====References==== | | ====References==== |
| - | <biblio>
| |
| | #Persikov2011 pmid=21572177 | | #Persikov2011 pmid=21572177 |
| | #CodonUsage http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/in-vitro-genetics/codon-usage.html | | #CodonUsage http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/in-vitro-genetics/codon-usage.html |
| | #OpenWetWareCodonUsage http://openwetware.org/wiki/Escherichia_coli/Codon_usage | | #OpenWetWareCodonUsage http://openwetware.org/wiki/Escherichia_coli/Codon_usage |
| | #NIHRareCodonCalculator http://nihserver.mbi.ucla.edu/RACC/ | | #NIHRareCodonCalculator http://nihserver.mbi.ucla.edu/RACC/ |
| - | </biblio></div>
| + | </div> |
| | <div id="620" style="display:none"> | | <div id="620" style="display:none"> |
| - | ==June 20th== | + | ==June 20th - Wet Lab== |
| | | | |
| | *Grew up colony of the selection strain with pKD46 in an attempt to reach mid-log and create glycerol stock | | *Grew up colony of the selection strain with pKD46 in an attempt to reach mid-log and create glycerol stock |
June 15th - Wet Lab
Gradient PCR: 5µL of each PCR product were run on a 1% gel. No bands appeared: the PCR appears to not have worked.
Selection construct: bacteria containing the selection construct (plasmid containing ZF, omega subunit, ZFB, His3, URA3, etc.) was made into a glycerol stock (see 6/14 and 6/9), miniprepped, and used for PCR:
- Miniprep: used Qiagen kit
- 82.5ng/µL, 260/280=1.99
- 91.1ng/µL, 260/280=2.01
- 77.9ng/µL, 260/280=1.98
- PCR:
- PCR used to amplify section of plasmid containing zinc finger binding site, weak promoter, His3, and URA3 (with homology to join it to kan cassette)
- Reagents
- zinc finger binding site and weak promoter, selection construct plasmid:
- 1µL ZFB-wp-f (5µM) (made by 1:20 dilution of 100µM stock)
- 1µL ZFB-wp-hisura-r (5µM) (made by 1:20 dilution of 100µM stock)
- 1µL selection construct (1:100 dilution of overnight culture)
- 22.5µL of invitrogen's Platinum PCR SuperMix
- PCR used to amplify the kan cassette
- Reagents
- KAN cassette, pZE22g plasmid:
- 1µL hisura-kan-f (5µM) (made by 1:20 dilution of 100µM stock)
- 1µL kan-r (5µM) (made by 1:20 dilution of 100µM stock)
- 1µL pZE22g (1:100 dilution of glycerol stock culture)
- 22.5µL of invitrogen's Platinum PCR SuperMix
- Parameters:
- 1) 94°C for 2 min (denature template, activate enzyme)
- 2) 94°C for 30 sec (denature)
- 3) 53°C for 30 sec (anneal)
- 4) 72°C for 2 min (extend)
- 5) Repeat 2-4 for 25 cycles total
- 6) 72⁰C for 5 min
- 7) 4°C forever
PCR Purification:
- Used the Qiagen PCR purification kit and instructions in order to purify the Kan cassette and selections construct PCR products
- Nanodrop the purification results and observed 3 ng/µL for Kan cassette and 29.8 ng/µL for ZFB-wp-His3: purification did not work well, especially for the kan
Selection strain (ΔHis3ΔPyrFΔrpoZ):
- saturated overnight culture was inoculated again: 3mL LB, 3µL tet, 30µL of overnight culture, at 37C until mid-log
- glycerol stock
- For transformation tomorrow we grew up pKD42 in 3 mL of LB, 1.5 µL of ampicillin(2000x) and one colony at 30 C
- Also grew up more of the selection strain so it will be ready for electroporation transformation
Gel
- Ran gel with Kan cassette and selection construct (Binding site, His3, and URA3)
- Observations successful and image below
- Used 1 kb plus ladder

Kan and ZFB-wp-his constructs 6/15/11
PCR Overlap
- since the purification was not very successful, we used 3µL saved from the original PCR product
- Procedure
- 25µL of 2x Phusion Master Mix
- 1 µL of ZFB-wp-HisURA-R (100µM)
- 1µL of HisURA-Kan-F (100µM)
- 21 µL of water
- 1 µL Kan template and 1 µL of ZFB-wp-His3-URA3
- 4 tubes
- Both undiluted
- Both 1:10 dilution
- Both 1:100 dilution
- Both 1:1000 dilution
- Protocol
- 98 C for 30 s
- 98 C for 10 s
- 53 C for 30 s
- 72 C for 3 min
- Repeat steps 2-4 for 24 more cycles
- 72 C for 5 min
- 4 C 4EVA!!!
June 15th - Bioinformatics
- We continued research into Persikov's and others' work on binding.
- We worked on using the OPEN data to test Persikov's binding-predicting program: SVM
- Justin continued work on a sequence-finding program, the most up to date version can be found in the Dropbox under code/zfsitefinder.html.
- Justin and Will found 10 candidate sequences across 4 diseases that hopefully should encompass a good amount of diversity in terms of expanding the ZF library. These sequences can be found in the table below, with more details here.
| Disease
| Target Range
| Binding Site Location
| Bottom Finger
| Top Finger
| Bottom AA (F3 to F1)
| Top AA (F3 to F1)
|
| Colorblindness | chrX:153,403,001-153,407,000 | 370 | GTATTTGTT | GGGCCTGCT | N/A | N/A
|
| Colorblindness | chrX:153,403,001-153,407,000 | 3627 | GCTGGCTGG | GCGGTAATG | EGSGLKR.EAHHLSR.####### | RRDDLTR.QRSSLVR.#######
|
| Cystic Fibrosis | chr7:117,074,084-117,089,556 | 14767 | GCAGGTGAT | AAAGAGCCC | QNGTLGR.EAHHLSR.####### | N/A
|
| Familial Hypercholesterolemia | chr19:11,175,000-11,195,000 | 14001 | GGCTGAGAC | GGAGTCCTG | ESGHLKR.QREHLTT.####### | QTTHLSR.DHSSLKR.#######
|
| Tay-Sachs | chr15:72,674,944-72,688,031 | 5888 | GTCTGGTCA | TCAAACTCC | DRSSLRR.RREHLTI.####### | N/A
|
| Pancreatic Cancer | chr7:117,074,084-117,089,556 | 1739 | GATCAAGCT | GTTTCAGTG | N/A | N/A
|
- We collected 15 alternative zinc finger backbones (different from zif268 backbone) and their corresponding base sequences. Many of these were from Persikov 2011 and all binding sequences were confirmed on the Protein Data Bank website. The zinc finger PDB ID's and related links are:
| PDB ID
| Binding Sequence
| Link
|
| 1F2I
| ATGGGCGCGCCCAT
| [1]
|
| 1G2D
| GACGCTATAAAAGGAG
| [2]
|
| 1G2F
| TCCTTTTATAGCGTCC
| [3]
|
| 1MEY
| ATGAGGCAGAACT
| [4]
|
| 1TF6
| ACGGGCCTGGTTAGTACCTGGATGGGAGACC
| [5]
|
| 1UBD
| AGGGTCTCCATTTTGAAGCG
| [6]
|
| 1TF6
| ACGGGCCTGGTTAGTACCTGGATGGGAGACC
| [7]
|
| 1YUI
| GCCGAGAGTAC
| [8]
|
| 2DRP
| CTAATAAGGATAACGTCCG
| [9]
|
| 2GLI
| TTTCGTCTTGGGTGGTCCACG
| [10]
|
| 2I13
| CAGATGTAGGGAAAAGCCCGGG
| [11]
|
| 2KMK
| CATAAATCACTGCCTA
| [12]
|
| 2PRT
| CGCGGGGGCGTCTG
| [13]
|
| 2WBS
| GAGGCGC
| [14]
|
| 2WBU
| GAGGCGTGGC
| [15]
|
June 17th - Wet Lab
Update on selection strain knockout status: We are trying to reach Addgene to check how His3 was knocked out---instead of deleting the gene, they may have simply introduced an early stop codon. If that's the case, our gel would have the correct bands because the primers we designed can only show whether a deletion or insertion was in that locus.
Transformation results: successful!!
- The only plate with colonies was the one plated with 700 µL of cells transformed with 45 ng of pKD42
- Chose a colony to grow in 3mL LB, 1.5µL amp, 30C; make glycerol stock with mid-log cells
- Plate with colonies at 4C
Miniprep of pZE22G: (to have the plasmid containing the kan cassette on hand)
- used 2 tubes of 1.5mL overnight culture, followed Qiagen kit instructions
- 38.0 ng/µL, 260/280=1.99
- 27.8ng/µL, 260/280=2.02
Overlap PCR gel and extraction: 1%, 150V
- Results: adding the primers in after 10 cycles was much more successful than adding them at the beginning, and all three dilutions showed the expected product band (about 2.5kb). The rest of the 1:10 dilution will be run on a gel and extracted.
- 11.2ng/µL, 260/280=2.10
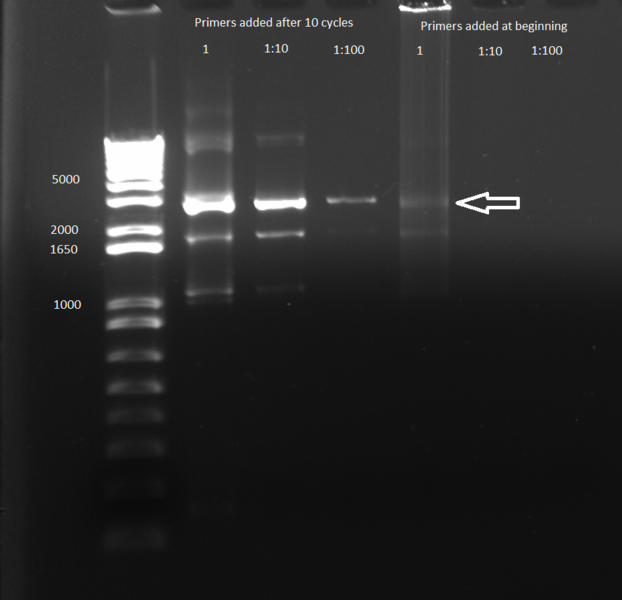
Successful Overlap of Kan Cassette and ZFB Gel 6/17/11
June 17 - Bioinformatics
Goals
- Make BB Database in program-readable format ✓
- Edit out BB with incomplete helices ✓
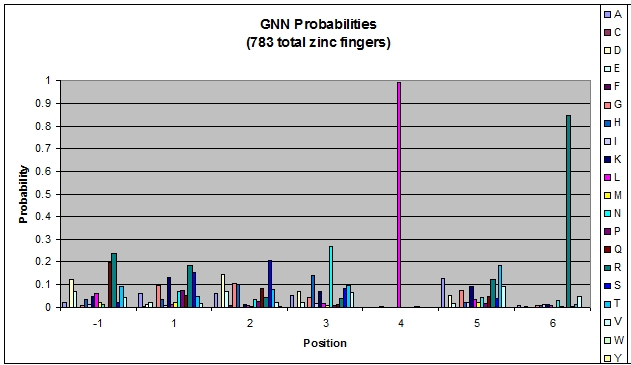
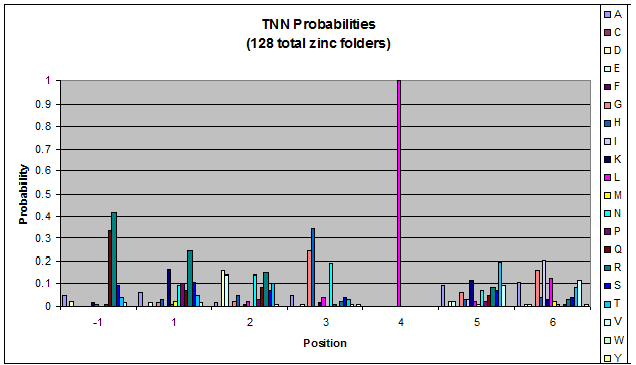
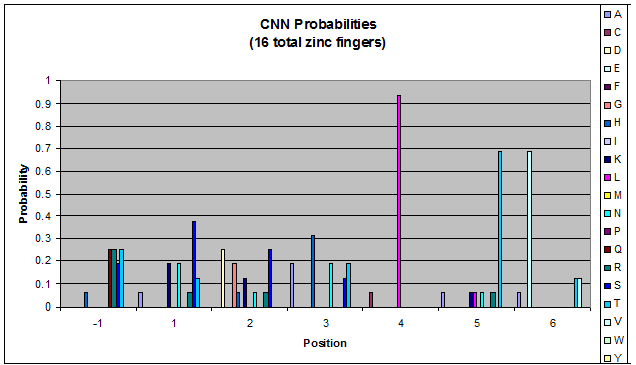
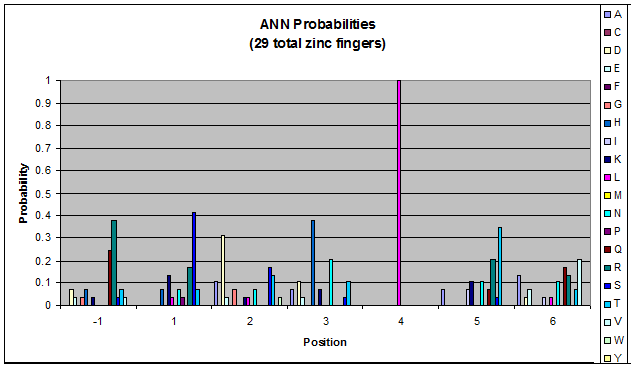
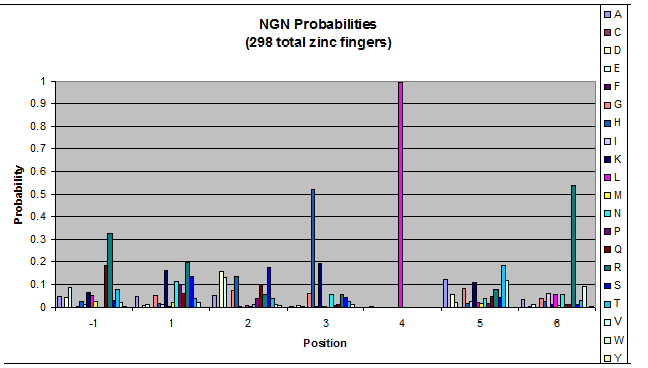
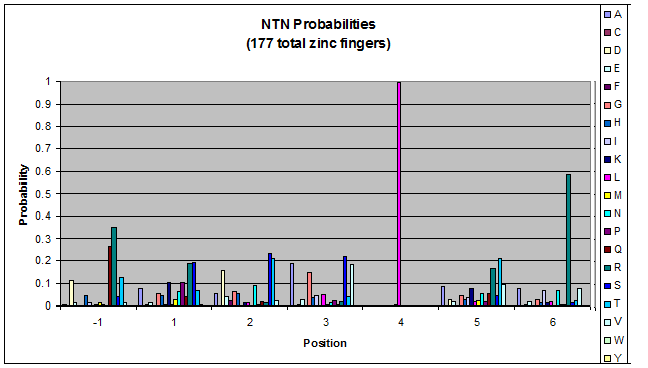
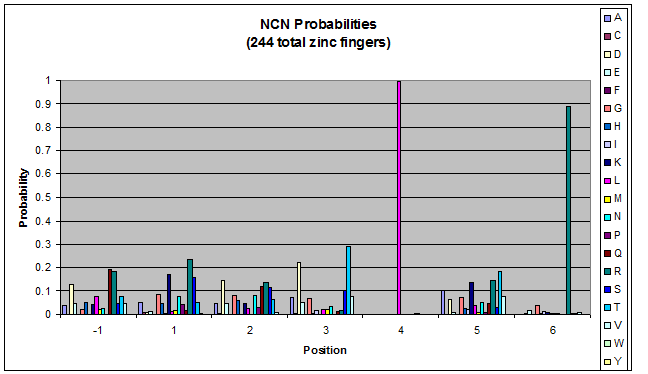
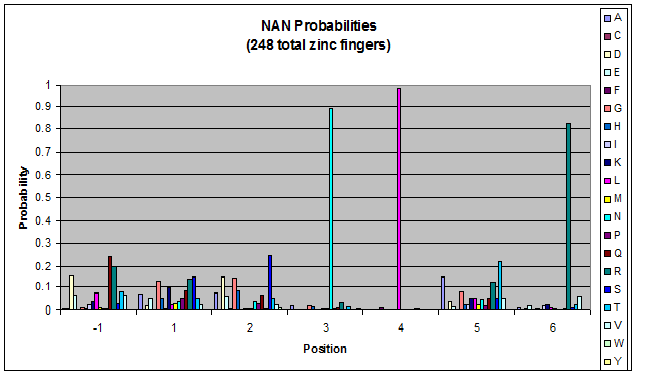
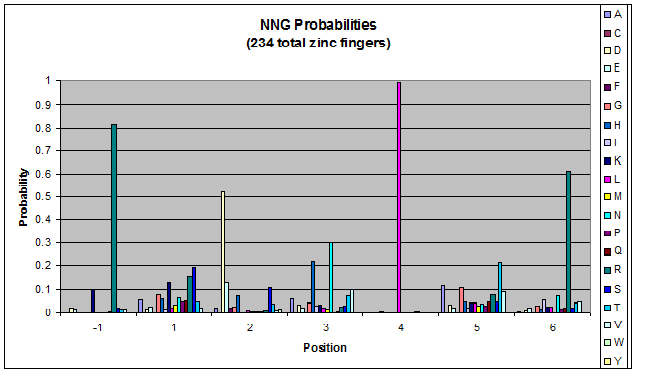
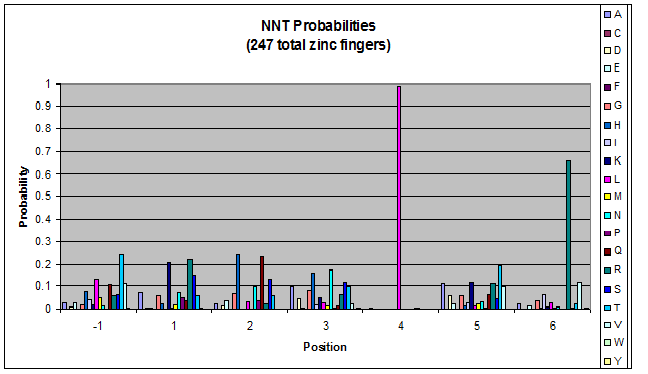
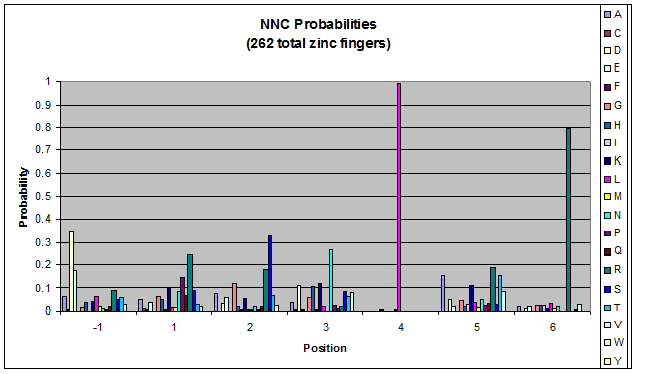
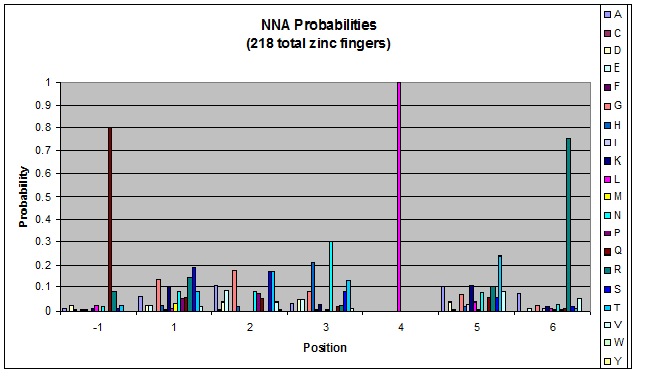
- GNN, TNN, CNN, ANN frequencies
- Targets (5-10; 8) x Backbones (???) x Helices (≥500)=55,000
- Backbones: similar, but not too similar to zif268; more than 1-2 aa changes, but <10
- Helices fixed based on our program-- eventually saturates and levels out
- Graph: # of var (# of tries by the computer) vs. % space covered
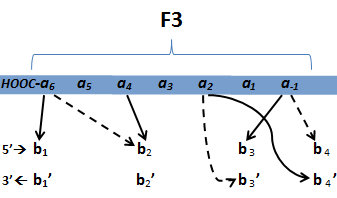
Proposed interactions between helical zinc finger residues and base pairs of the target DNA sequence (based on
Persikov 2011)
Options for Target DNA Sequences / ZF Helices
- F3(known) / F2(known) / F1(novel)
- F3(known) / F2(SNP in b1 position) / F1(known)
- F3(unknown) / F2(unknown) / F2(unknown)
- Excluded Rare Codons (for E. coli)CodonUsage OpenWetWareCodonUsage NIHRareCodonCalculator:
- CTA
- ATA
- CCC
- CGA
- CGG
- AGA
- AGG
- GGA
- GGG
References
- Persikov2011 pmid=21572177
- CodonUsage http://www.sci.sdsu.edu/~smaloy/MicrobialGenetics/topics/in-vitro-genetics/codon-usage.html
- OpenWetWareCodonUsage http://openwetware.org/wiki/Escherichia_coli/Codon_usage
- NIHRareCodonCalculator http://nihserver.mbi.ucla.edu/RACC/
 "
"