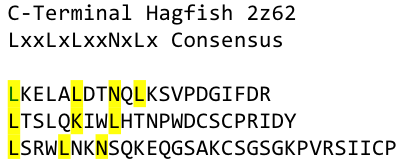Team:Freiburg/Modelling
From 2011.igem.org
(Difference between revisions)
(→Modelling: Rational protein design) |
(→Modelling: Rational protein design) |
||
| Line 6: | Line 6: | ||
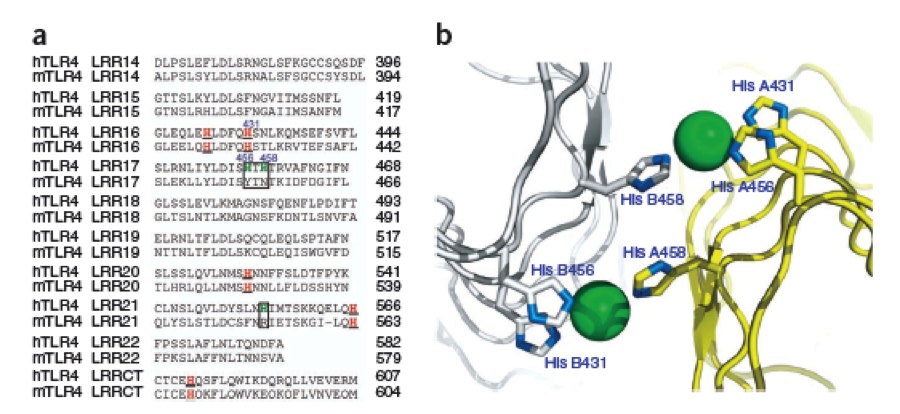
The second milestone in the generation of the idea came up in a lecture of Prof. Martin, about Nickel allergy. He found a mutation in the TLR-4 receptor that indroduces Histidines in the LRR motif of the protein, which are capable of binding nickel and thus forming complexes of receptors on the cell surface of immune cells, triggering the inflammatory reaction. | The second milestone in the generation of the idea came up in a lecture of Prof. Martin, about Nickel allergy. He found a mutation in the TLR-4 receptor that indroduces Histidines in the LRR motif of the protein, which are capable of binding nickel and thus forming complexes of receptors on the cell surface of immune cells, triggering the inflammatory reaction. | ||
| - | [[File:Freiburg11Modelling1.png| | + | [[File:Freiburg11Modelling1.png|thumb|750px]] |
This was the necessary clue that gave answer to the question. | This was the necessary clue that gave answer to the question. | ||
| Line 13: | Line 13: | ||
The Construction of a new protein | The Construction of a new protein | ||
After a long and detailed search, we found out that it is most reasonable to use bacterial LRR motifs, since they seemed very well conserved in sequence, and they are the shortest – with only 21 aminoacids per LRR repeat. (Wei 2008, Kajava 1998). We wanted to have the protein as simple and as predictable in behavior and structure as possible. | After a long and detailed search, we found out that it is most reasonable to use bacterial LRR motifs, since they seemed very well conserved in sequence, and they are the shortest – with only 21 aminoacids per LRR repeat. (Wei 2008, Kajava 1998). We wanted to have the protein as simple and as predictable in behavior and structure as possible. | ||
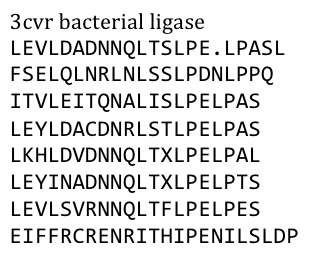
| - | Several search inquiries led to the most conserved and shortest of all bacterial LRR (PDB: NO), unluckily this protein is a toxin derived from Yersinia pestis. We of course did not want to mess around with toxins,[[File:Freiburg11_Seq1.png| | + | Several search inquiries led to the most conserved and shortest of all bacterial LRR (PDB: NO), unluckily this protein is a toxin derived from Yersinia pestis. We of course did not want to mess around with toxins,[[File:Freiburg11_Seq1.png|thumb|250px]] especially not with one of the black plague. We also did not know what amino acid pattern on the LRR motif would cause the toxic effect. There probably would have been no harm from this protein alone, even if we did clone and express it, but why should we do that when we have a huge choice for free at hand anyway. Finally we settled on the bacterial ligase PDB:3CVR. A ligase would be definitively harmless, especially since we planned to restructure it completely. |
To do this we needed to understand how the structure of the Ligase was made. | To do this we needed to understand how the structure of the Ligase was made. | ||
| Line 22: | Line 22: | ||
[[File:Freiburg11Modelling2.png|thumb|750px]] | [[File:Freiburg11Modelling2.png|thumb|750px]] | ||
| - | By analyzing the logo it was obvious which positions of the LRR motif were conserved and which not – AND: which of the non conserved aminoacids appeared in what kind of patterns. Were there positions in the protein that required a polar aminoacid? Or non polar, hydrophobic /-philic, charged, non-charged?. We compared the consensus sequence with the 3D structure, using PYMOL, to extract as much information as possible and then came up with this ideal consensus sequence:[[File:Freiburg11_Seq2. | + | By analyzing the logo it was obvious which positions of the LRR motif were conserved and which not – AND: which of the non conserved aminoacids appeared in what kind of patterns. Were there positions in the protein that required a polar aminoacid? Or non polar, hydrophobic /-philic, charged, non-charged?. We compared the consensus sequence with the 3D structure, using PYMOL, to extract as much information as possible and then came up with this ideal consensus sequence: |
| + | [[File:Freiburg11_Seq2.png|thumb|300px]] | ||
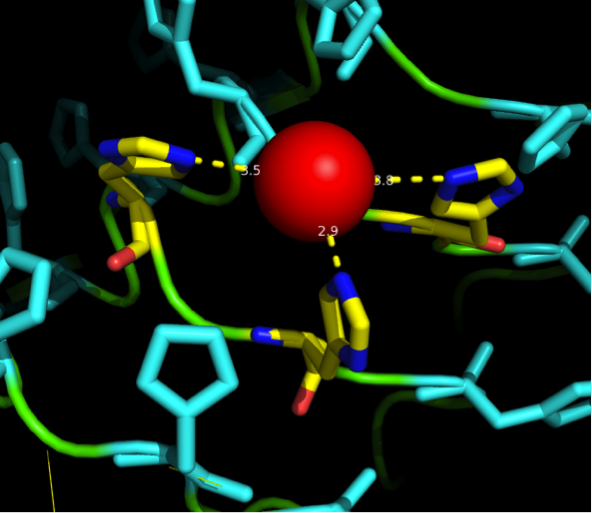
The next consideration we had to do, was: how many Nickel do we need on the surface of our ideal Nickel binding protein, in what pattern, with what distances between, and at what angles towards each other to allow proper ion complexation? | The next consideration we had to do, was: how many Nickel do we need on the surface of our ideal Nickel binding protein, in what pattern, with what distances between, and at what angles towards each other to allow proper ion complexation? | ||
| Line 32: | Line 33: | ||
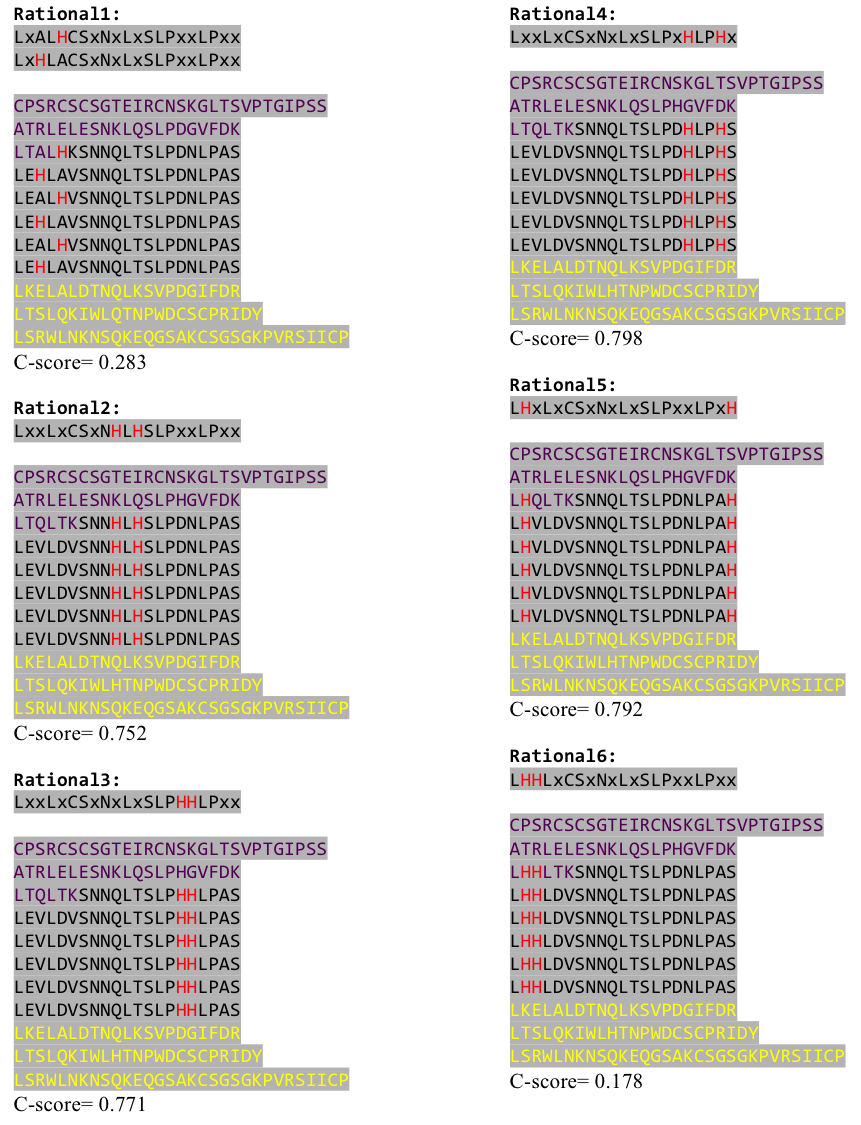
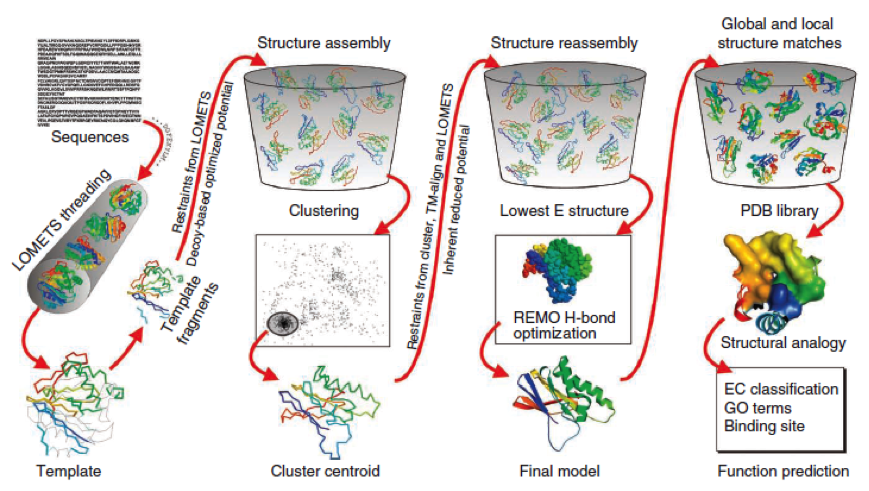
To get a first impression how Histidines are positioned in a bacterial Ligase, we replaced all non conserved aminoacids by Histidines and sent the sequence for a 3D sequence prediction to I-TASSER, an online structure prediction software tool. | To get a first impression how Histidines are positioned in a bacterial Ligase, we replaced all non conserved aminoacids by Histidines and sent the sequence for a 3D sequence prediction to I-TASSER, an online structure prediction software tool. | ||
| - | [[File:Freiburg11_Seq7.png]] | + | [[File:Freiburg11_Seq7.png|thumb|300px]] |
| Line 70: | Line 71: | ||
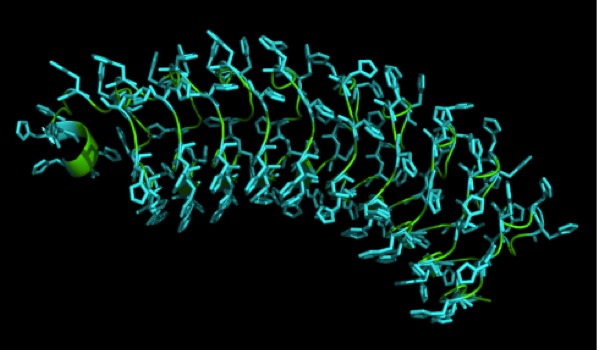
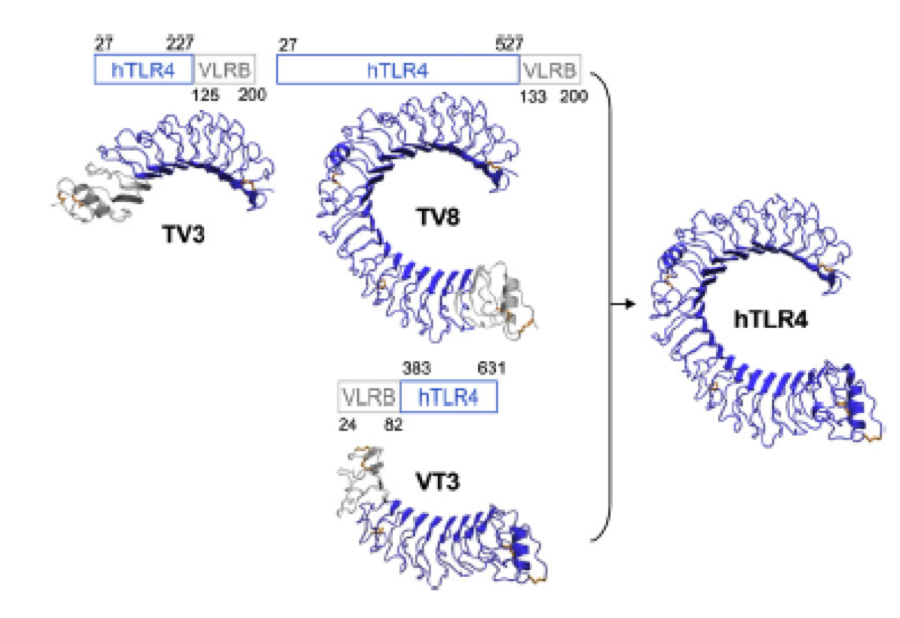
What we further realized from the structure file was, that the end of the LRR segments were “open”, that means, the hydrophobic core of the protein was exposed and as it is visible in the prediction, curled in on one end into a sort of helix. This means the protein folding is not reliable and the structure needs some caps on both ends to stabilize the LRR core motif. A solution to this was shown by Schmidt et al 2010, who crystallized the TLR-4 receptor. | What we further realized from the structure file was, that the end of the LRR segments were “open”, that means, the hydrophobic core of the protein was exposed and as it is visible in the prediction, curled in on one end into a sort of helix. This means the protein folding is not reliable and the structure needs some caps on both ends to stabilize the LRR core motif. A solution to this was shown by Schmidt et al 2010, who crystallized the TLR-4 receptor. | ||
| - | [[File:Freiburg11Modelling6.png| | + | [[File:Freiburg11Modelling6.png|thumb|750px]] |
In this very nice piece of work he dissected the TLR4(PDB: 3FXI) into 3 parts, since it was to large and unhandy to be crystallized at once. To overcome this problem of an exposed hydrophobic core, he used the N- and C-terminal protein fragments of a LRR protein derived from hagfish. They tried a variety of different versions of how to glue together his fragments and these N- and C-terminal caps until he found a working one. | In this very nice piece of work he dissected the TLR4(PDB: 3FXI) into 3 parts, since it was to large and unhandy to be crystallized at once. To overcome this problem of an exposed hydrophobic core, he used the N- and C-terminal protein fragments of a LRR protein derived from hagfish. They tried a variety of different versions of how to glue together his fragments and these N- and C-terminal caps until he found a working one. | ||
We used this knowledge for our purpose and took the same sequences and attached them to our protein sequence. These caps partially still show the typical LRR consensus sequences (which luckily is highly conserved in all kingdoms of life!), which made it possible to fit them onto our stack of LRR loops in the right position. | We used this knowledge for our purpose and took the same sequences and attached them to our protein sequence. These caps partially still show the typical LRR consensus sequences (which luckily is highly conserved in all kingdoms of life!), which made it possible to fit them onto our stack of LRR loops in the right position. | ||
| - | [[File:Freiburg11_Seq3.png]] | + | [[File:Freiburg11_Seq3.png|thumb|300px]] |
| Line 93: | Line 94: | ||
{|style="color:black; border="1" width="85%" | {|style="color:black; border="1" width="85%" | ||
| style="width: 100%;background-color:lightgrey;" | | | style="width: 100%;background-color:lightgrey;" | | ||
| - | ;[[File:Freiburg11_Seq6.png]] | + | ;[[File:Freiburg11_Seq6.png|thumb|750px]] |
|} | |} | ||
{{:Team:Freiburg/Templates/footer}} | {{:Team:Freiburg/Templates/footer}} | ||
Revision as of 15:23, 20 September 2011
 "
"
 Contact
Contact