Team:BU Wellesley Software/G-nomeSurferPro
From 2011.igem.org
(Difference between revisions)
m |
|||
| Line 56: | Line 56: | ||
The chromosome bar is infinitely scrolling and position on the chromosome bar is tied into the orientation of the chromosome wheel through the view-model system allowing manipulation of one to affect change of the other. GenBank files are parsed for sequence, translation and any notes, all of which are available as information objects, which are accessible from any gene on the bar. | The chromosome bar is infinitely scrolling and position on the chromosome bar is tied into the orientation of the chromosome wheel through the view-model system allowing manipulation of one to affect change of the other. GenBank files are parsed for sequence, translation and any notes, all of which are available as information objects, which are accessible from any gene on the bar. | ||
<br><br> | <br><br> | ||
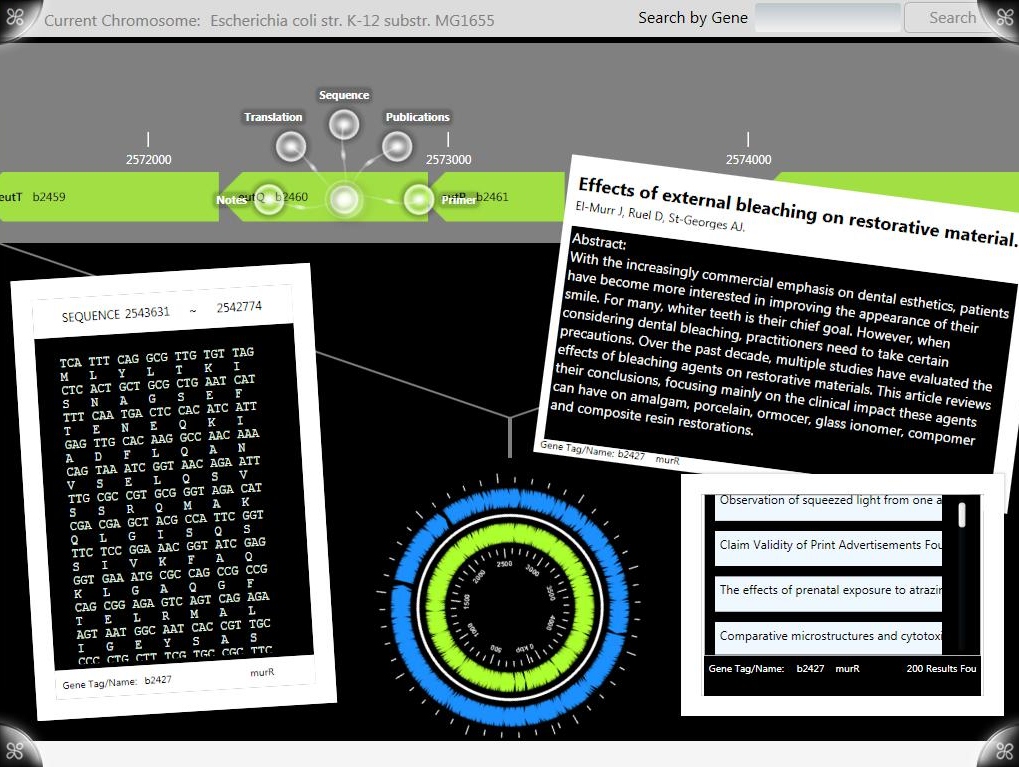
| - | Information from any gene’s GenBank file are viewable as “ScatterViewItem” objects which can be moved and re-sized on the main “ScatterView” workspace. Users can view and annotate sequence/translation information by dragging and overlaying one over the other. All information objects can be moved to and from the extended desktop by dragging or tossing or erased by tossing them off the sides of the workspace. By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view. | + | Information from any gene’s GenBank file are viewable as “ScatterViewItem” objects which can be moved and re-sized on the main “ScatterView” workspace. Users can view and annotate sequence/translation information by dragging and overlaying one over the other. All information objects can be moved to and from the extended desktop by dragging or tossing or erased by tossing them off the sides of the workspace. |
| + | |||
| + | <img style="float:right; width:280px; height:200px" src="http://cs.wellesley.edu/~hcilab/iGEM_wiki/images/System/SurfacePrimer.jpg"/> | ||
| + | By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view. | ||
<br><br> | <br><br> | ||
Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data. | Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data. | ||
<br><br> | <br><br> | ||
| - | |||
<br> | <br> | ||
In the Primer Genie workspace, alignment results are generated using an algorithm .jar written by the software team, and BLAST results are generated using the NCBI BLAST software. Both are accessed from the command line, with BLAST results being written to file and parsed out for display and alignment results being pulled from common output for further calculation for hetero-dimer and self-dimer tests. | In the Primer Genie workspace, alignment results are generated using an algorithm .jar written by the software team, and BLAST results are generated using the NCBI BLAST software. Both are accessed from the command line, with BLAST results being written to file and parsed out for display and alignment results being pulled from common output for further calculation for hetero-dimer and self-dimer tests. | ||
Revision as of 16:23, 10 September 2011
Computational Team
G-nome Surfer Pro is written in C# for the Microsoft Surface using the Surface SDK. State is managed using the ViewModel system and time consuming processes, like retrieving publications and running BLAST, are run in threads behind the main UI thread.
After selecting a genome from the tree menu the chromosome bar is populated with genes from the GenBank file. Another genome can be selected at anytime from the pull down search menu.
The chromosome bar is infinitely scrolling and position on the chromosome bar is tied into the orientation of the chromosome wheel through the view-model system allowing manipulation of one to affect change of the other. GenBank files are parsed for sequence, translation and any notes, all of which are available as information objects, which are accessible from any gene on the bar.
Information from any gene’s GenBank file are viewable as “ScatterViewItem” objects which can be moved and re-sized on the main “ScatterView” workspace. Users can view and annotate sequence/translation information by dragging and overlaying one over the other. All information objects can be moved to and from the extended desktop by dragging or tossing or erased by tossing them off the sides of the workspace.
 By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view.
By moving objects between ScatterViews, users can view and organize data in a distraction free environment which can be pulled up over the main workspace or hidden from view.
Publications are populated in a scrollable list and can be dragged and dropped from the listBox to the main workspace or extended desktop through manipulation of the objects template. The PubMed search results for the gene are fetched and parsed for abstracts, authors, and publication data.
In the Primer Genie workspace, alignment results are generated using an algorithm .jar written by the software team, and BLAST results are generated using the NCBI BLAST software. Both are accessed from the command line, with BLAST results being written to file and parsed out for display and alignment results being pulled from common output for further calculation for hetero-dimer and self-dimer tests.
Results:
Ethical User Study practices:
Overview: G-nome Surfer Pro
Hypothesis Forming
G-nome Surfer Pro is an integrated environment for viewing prokaryotic genomic data and literature. Users can find genes on the chromosome wheel or search for a gene name or Genbank number(?), access Genbank notes and publications from Pubmed and view alignment and translation from the chromosome bar. The extended desktop provides distraction free workspace for information processing and hypothesis formation. The Microsoft Surface allows users to collaborate in pairs and teams on design and research tasks
Genes can be taken directly into the Primer Genie designer for primer design. In the Primer Genie workspace users can perform alignment related tests, BLAST and save designs as BioBricks. The system is designed for integration with the Clotho database.

Demo Video
Wetlab
Lorem ipsum dolor sit amet, consectetur adipiscing elit. In et dictum leo. Maecenas porttitor augue nec arcu lacinia ultricies. Maecenas at dictum augue. Proin eget odio ac mi tristique scelerisque. Integer at
nisl nec purus laoreet condimentum sed semper dui. Duis feugiat, ligula
eu vehicula vulputate, sem nisl ornare neque, eu dictum lacus sapien
eu neque.
Integer at
nisl nec purus laoreet condimentum sed semper dui. Duis feugiat, ligula
eu vehicula vulputate, sem nisl ornare neque, eu dictum lacus sapien
eu neque.
Vestibulum gravida, turpis tempus suscipit euismod, ipsum mi tristique nibh, sed sagittis ante felis at urna. Suspendisse eu neque vitae lorem elementum elementum sit amet ac ligula. Mauris vestibulum Donec ac sapien erat. Proin id enim sed dolor suscipit laoreet vitae sit amet ligula. Nam ultricies orci vitae mauris egestas tincidunt. Vestibulum sit amet est dolor.
Results:
Ut eu nunc eget ante egestas egestas at eu metus. Integer quam justo, vehicula non sodales id, consequat vitae eros.Aenean egestas, ipsum sed fringilla porta, erat mi facilisis lectus, quis elementum quam felis nec elit.
 "
"
