Team:Kyoto/Lab Work
From 2011.igem.org
Lab Work
Week1: Monday 1st - Sunday 7th August
Monday
行った実験名:
使った試薬名、容量:
用いた機械:
行った人:
Tuesday
Wednesday
Thursday
Friday
Saturday
Sunday
Week2: Monday 8th - Sunday 14th August
Monday
Tuesday
Wednesday
Thursday
Friday
Saturday
Sunday
Week3: Monday 15th - Sunday 21th August
Monday
Tuesday
Wednesday
Thursday
Nutritional Signal(Sugiura,Shimosaka & Okumura)
PCR Amplification of glnL and glnG from gDNA of E.coli
--Primers--
glnL
Left primer: tctagaggagactgctttatggcaac
Right primer: actagtaggaactatcgtcatcgactac
glnG
Left primer: tctagaggtgacgtttatgcaacga
Right primer: actagtacacacaagctgtgaatcactc
annealing temperature was 55 degrees.
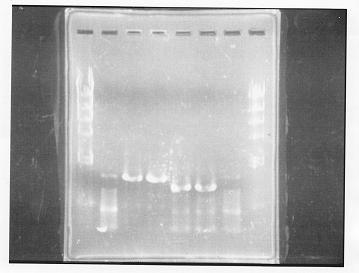
lane 1,2 &7,8: DNA ladder(λDNA digested Hindlll,100bp), lane 3,4: glnG1,glnG2, lane 5,6: glnL1,glnL2
After purification, the concentration of DNA are
glnG1: 127.8ng/ul
glnG2: 118.1ng/ul
glnL1: 137.4ng/ul
glnL2: 124.2ng/ul
Friday
Nutritional Signal(Shimosaka)
Restriction of glnG1 and glnG2
Cut them with Xbal and Spel for 2 hours at 37 degrees.
Then, gel extraction of digested.
Saturday
Sunday
Week4: Monday 22th - Sunday 28th August
Monday
Tuesday
Wednesday
Thursday
Luminescence(Kusaba, Terada, Hara):ハエの走行性実験① ♂、紫外線×2回、緑×2回 ♀、紫外線×2回、緑×2回
Friday
Luminescence(Kusaba):ハエの走行性実験① ♂、赤外線×2回 ♀、赤外線×2回
Nutritional Signal(Hashiya):
Transformation of bellow parts.
4-17M:BBa_K325909(lux operon)
1-12M:BBa_E0240
2-17F:BBa_120260(low copy vector)
PCR amplification of
Saturday
Luminescence(Kusaba):ハエの走行性実験① ♂、赤×2回、青×2回 ♀、赤×2回、青×2回
Sunday
Week5: Monday 29th August - Sunday 4th September
Monday
Tuesday
Nutritional Signal(Hashiya)
・PCR amplification of glnL and glnG from PCR products,glnL1 and glnG1.
--Primers--
glnL
Left primer: ggaattcgcggccgcttctagaggagactgctttatggcaac
Right primer: ggactagtaggaactatcgtcatcgactac
glnG
Left primer: ggaattcgcggccgcttctagaggtgacgtttatgcaacga
Right primer: ggactagtacacacaagctgtgaatcactc
annealing temperature was 55 degrees.
・PCR amplification of glnL+G and rpoN from gDNA of E.coli.
--Primers--
glnL+G
Left primer: ggaattcgcggccgcttctagaggagactgctttatggcaac
Right primer: ggactagtacacacaagctgtgaatcactc
rpoN
Left primer: ggaattcgcggccgcttctagaggttctgaacatgaagcaa
Right primer: ggactagtatccttatcggttgggtca
annealing temperature was 56 degrees.
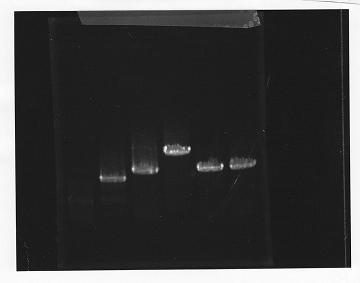
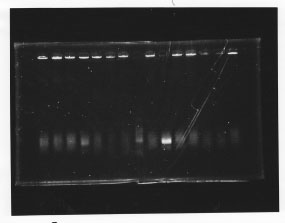
lane1: 100bp DNA ladder, lane2:glnL, lane3:glnG, lane4:glnG+L, lane5:rpoN from gDNA, lane6:rpoN from ASKA clone
After purification, the concentration of DNA are
glnL: 122.3 ng/ul
glnG: 64.7 ng/ul
glnL+G: 106.7 ng/ul
rpoN from gDNA: 111.4 ng/ul
Wednesday
Nutritional Signal(Hashiya)
・Screening PCR of σ54 promoter + pSB1A3
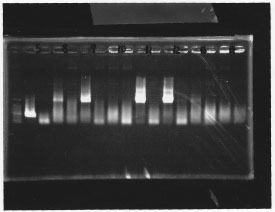
We cultured σ54 promoter5
Friday
Nutritional Signal(Hashiya)
・Restriction of σ54 promoter5,glnL, glnG, glnL+G and rpoN
Cut them with EcoRl and Spel
After purification, the concentration of DNA were
σ54 promoter5: 23.6 ng/ul
glnL: 28.1 ng/ul
glnG: 26.3 ng/ul
glnL+G: 15.3 ng/ul
rpoN: 20.8 ng/ul
・Ligation reaction
Ligated glnL, glnG and rpoN to pSB1K3.
Thursday
Nutritional Signal(Hashiya)
・Screening PCR of glnL, glnG, glnL+G and rpoN
glnL
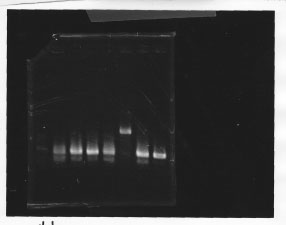
glnG

glnL+G & rpoN
We cultured glnL5, glnG4
Saturday
Nutritional Signal(Hashiya)
・Mini prep of glnL5 and glnG4
glnL5: 43.9 ng/ul
glnG4: 38.1 ng/ul
Predation(Hashiya)
・PCR amplification of glmS
--Primers--
left primer:ggaattcgcggccgcttctagagcaggttgaccgacaacgata
right primer:ggactagtacgcagggcatccatttat
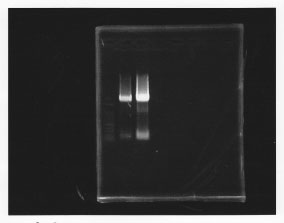
lane1: 100bp DNA ladder, lane2: glmS from gDNA, lane3: glmS from ASKA clone
・TA cloning of glmS
Ligated glmS from gDNA to pTA vector.
Sunday
Nutritional Signal(Hashiya)
・Screening PCR of rpoN

・Transformation of bellow parts
1-23L: BBa_B0015 (double terminator)
1-18E: BBa_J23101 (constitutive promoter)
1-18C: BBa_J23100 (constitutive promoter)
Week6: Monday 5th September - Sunday 11th September
Monday
Nutritional Signal(Hashiya)
・Screening PCR of glnL+G+double terminator
Tuesday
Luminescence(Kusaba, Hara):ハエの走行性実験②(②は改良版) ♂、緑×2回 ♀、青×1回
Wednesday
Luminescence(Hara):ハエの走行性実験② ♂、青×2回 ♀、緑×2回、青×1回
Thursday
Luminescence(Hara):ハエの走行性実験② ♂、紫外線×3回 ♀、紫外線×3回
Friday
Saturday
Luminescence(Kusaba, Hara):ハエの走行性実験② ♂、青×2回、赤×2回 ♀、青×2回、赤×2回
Sunday
Luminescence(Kusaba, Hara):ハエの走行性実験② ♂、紫外線×2回、
Week7: Monday 12th September - Sunday 18th September
Monday
Luminescence:大腸菌の形質転換(Hashiya) ハエの走行性実験②(Kusaba, Hara)
Tuesday
Luminescence:大腸菌はじめて光る。しかし光量は少ない。
Wednesday
Thursday
Friday
Digestion(Kajita)
PCR amplification of SAM-P20 and ChiA
- We performed colony direct PCRs from a S. albogriseolus colony and a S. avermitilis colony.
Reaction mixture Component Volume(μl) 2x Buffer 25 2mM dNTPs 10 Primer 1 1.5 Primer 2 1.5 Template X KOD FX 1 ddH2O up to 50
PCR condition Predenature 94C 2m Denature 98C 10s 30cycles Annealing 56C 30s Extension 68C 1m30s
- We prepared two kinds of templates:
- Picked a colony, suspended in 50ul of water and incubated 1min at 95 degree. Added 1ul to the reaction mixture.
- Picked a colony and dipped in the reaction mixture.
- Gel electrophoresis indicated that the PCR amplifications were successful for a sample, ChiA, but it was a faint band. We decided to retry direct PCR of SAM-P20 and PCR of ChiA.
Saturday
Digestion(Kajita)
Retry of PCR amplification of SAM-P20 and ChiA.
- We amplified SAM-P20 by colony direct PCR and ChiA by PCR using the PCR product that performed yesterday.
- Reaction mixture: The same components and volume as before.
- PCR condition: The same PCR condition as before.
- PCR for SAM-P20
- We prepared two kinds of templates:
- Picked a colony, suspended in 50ul of TE buffer (pH8.0) and incubated 1min at 95 degrees. Added 5ul to the reaction mixture.
- Picked a colony and dipped in the reaction mixture.
- PCR for ChiA
- 1μl of PCR product was added to the reaction mixture as template.
- Gel electrophoresis indicated that the PCR amplifications were successful for all samples. However, an unexpected faint band, about 1000bp, was also observed in the sample of ChiA.
PCR purification of SAM-P20 and gel extraction of ChiA.
- SAM-P20: 43.9 ng/μl
- ChiA: 30.0 ng/μl
Restriction enzyme digestion
- We performed restriction digestions for:
- SAM-P20 with EcoRI and SpeI
- ChiA with EcoRI and SpeI
- Incubated overnight at 37 degrees.
Sunday
Digestion (Kajita)
Purification of digested products
- SAM-P20 : 10.2 ng/μl
- ChiA : 22.5 ng/μl
Ligation
Name Vector Insert 1 pSB1C3 SAM-P20 2 pSB1C3 ChiA 3 BBa_B0015 SAM-P20 4 BBa_B0015 ChiA
- Incubated overnight at 16 degrees.
PCR amplification of BBa_R0011, lactose promoter
- We done the amplification of lactose promoter. After that, we digested the product with DpnI, incubated 1 hour at 37 degrees.
- Gel electrophoresis indicated that the PCR amplification was successful, but DpnI didn't work at all. So, we done the gel extraction.
Restriction enzyme digestion for pSB4K5 with EcoRI and PstI
Week8: Monday 19th September - Sunday 25th September
Monday
Tuesday
Wednesday
Thursday
Friday
Saturday
Sunday
Week9: Monday 26th September - Sunday 2nd October
Monday
Tuesday
Wednesday
Thursday
Friday
Saturday
Sunday
Protocol
Medium for drosophila
[http://www.biol.se.tmu.ac.jp/fly/www/standard-medium.html Medium for drosophila] Materials Methods - water : 500mL
- dry yeast : 20g
- corn flour : 45g
- glucose : 50g
- agarose : 3.5~5g
- propionic acid : 1.5mL
- 10% p-hydroxybenzoate in 70% Eternol : 5g
- Stir dry yeast and agarose with about two-thirds of water. Then, autoclave it.
- Stir corn flour and glucose with the remaining water.
- Stir 1 and 2, then autoclave it again.
- after autoclave, add propionic acid and 10% p-hydroxybenzoate in 70% Eternol into it.■
 "
"






