Team:Potsdam Bioware/Project/Details Selection
From 2011.igem.org
Contents |
In Vivo Selection
Introduction
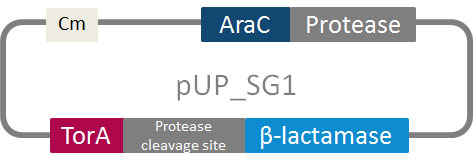
In addition to the phage display we developed a novel selection system. The design aimed for a cheap and time-saving alternative in contrast to an in vitro screen of protease inhibition kinetics. The assay allows us to select effective inhibitors for any protease, among the billions of randomly generated mutants of the Microviridin. For this purpose we designed a plasmid containing two devices, first a protease activity detector and second a protease generator. For the protease activity detector we modified the TAT-hitchhiker system developed by Delisa et al.
We used the BioBrick <partinfo>I757010</partinfo> (β-lactamase) as well as <partinfo>K208005</partinfo> (ssTorA) and fused them together via a linker peptide. For the protease generator we cloned the Arabinose induction system in front of a protease in iGEM standard.
The system works in a combined manner of the two devices. In order to confer β-lactam antibiotic resistance to cells the β-lactamase has to be exported in the periplasm. This transport is mediated by the TorA export sequence via the Twin-Arginine Translocation (TAT) system (DeLisa, 2008). The linker peptide between the TorA export sequence and the β-lactamase displays the corresponding protease cleavage site. In addition the linker peptide is chosen as short as possible to imitate folding because the TAT pathway only allows transport of correctly folded substrates. The protease as well as the linker peptide are designed as exchangeable parts.
If the protease device is functional, it will cleave the linker peptide between TorA and the β-lactamase construct which leaves the cell without any antibiotic resistance. The construct was tested with increasing Ampicillin concentrations. The number of surviving colonies was depended on the export rate of the β-lactamase into the periplasm. A high cleavage rate of the linker peptide leads to a reduced Ampicillin resistance. With expressed protease a dramatically drop of the number of surviving colonies could be observed.
The survival assay was carried out with and without expressed protease. By increasing of the Ampicillin concentration we could detect a cutoff Ampicillin concentration. The colony forming units (CFU) were counted and compared. To detected whether the our constructed library is able to block the protease we co-transformed the library into E. coli carrying our pUP_SG1 plasmid.
Site-directed mutagenesis of TEV and 14_3C proteases in iGEM standard
We chose two model proteases, the Tobacco Etch Virus protease, further along called TEV, and the 14_3C protease from the human rhinovirus, also known as PreScission. Both proteases contained iGEM restriction sites, one in case of the TEV and three for 14 3C, respectively.
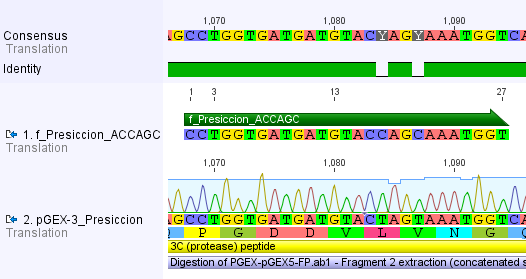
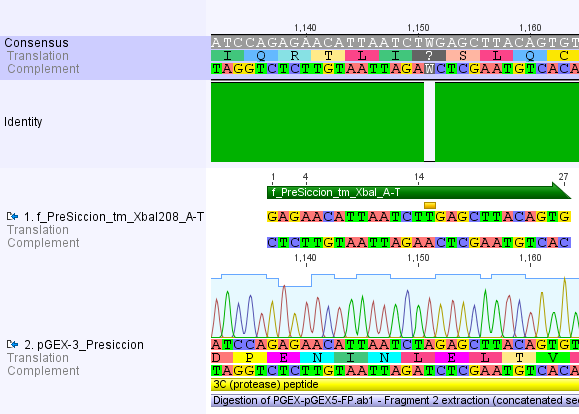
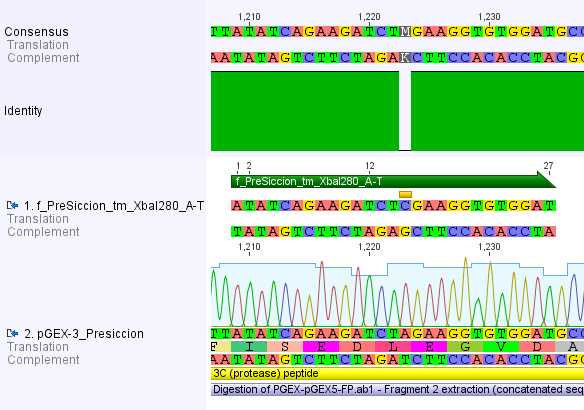
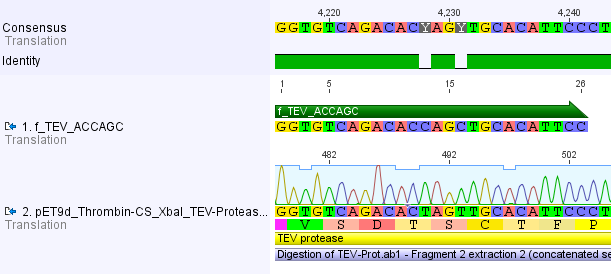
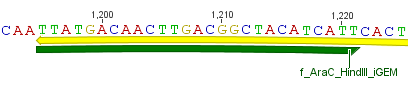
Thus, site-directed mutagenesis was applied to mutate each restriction site in E. coli. For every mutation a forward and a reverse primer had to be designed. The following pictures show the forward primers for the side directed mutagenesis, the reverse primers are the reverse complement sequence of the forward primers.
For the 14_3C protease:
For the TEV protease:
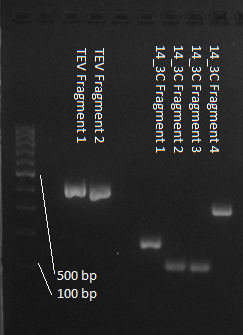
We used assembly PCR to gain the final product. In case of PreScission two assembly PCRs had to be done. The expected size of the fragments are shown in the gel picture. In the following picture all fragments contain the correct sizes.
Construction of two component for the in vivo selection assay
The in vivo assay is based on two devices. On the one hand the protease activity detector device, on the other a protease generator device. The detector device is based on the enzyme β-lactamase, which is only active inside the periplasm and provides the resistance against lactam antibiotics like ampicilin and penicilin. For the protease generator device we needed a second induction system on our plasmid, which is quite uncommon for any plasmid. So several problems had to solved to set up an effective in vivo selection assay:
- a second, very thight induction system has to be found
- a way to control the export rate of β-lactamase into the periplasm, which can be reduce by the protease
The solution to these problems are simple: the protease detector device is under the controll of the lac operon and induced by IPTG, so the choice is easy and we selected the arabinose inducible induction system of the pBAD_iGEMexpress vector. The solution for the export problem of β-lactamase is inspired by Hitchhiker assay of De'Lias et al. Using the TAT pathway for the export of β-lactamase into the periplasm gives us the ability to introduce a protease specific cleavage site infront of the signal sequence to regulate the export.
Introducing the specific protease cleavage site
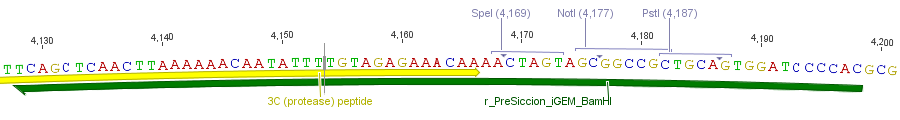
To adapt the protease activity detector device to any protease we want to screen, we had to modularize it. Our detector device was set up on a vector for the Hitchhiker selection. The plasmid offers two unique restriction sites on the plasmid, XhoI and NheI. Both were used to introduce the cleavage site in between the TorA signal sequence and the β-lactamase. By designing two complementary oligonucleotides containing the cleavage site flanked by two short linker regions. After the digest of the vector with XhoI and NheI the hybridized oligos can be ligated into the vector.
Adding the arabinose inducible induction system to control the proteases, TEV and 14_3C
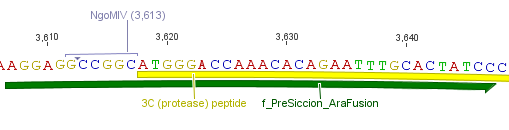
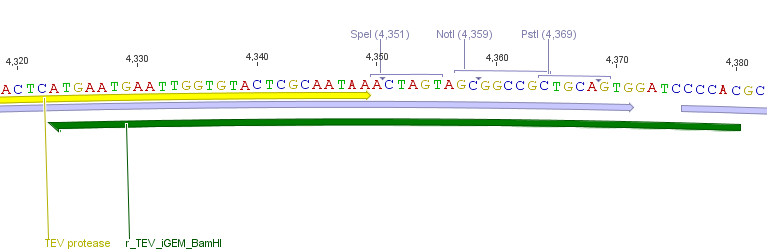
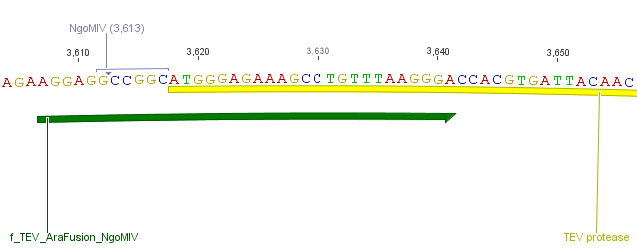
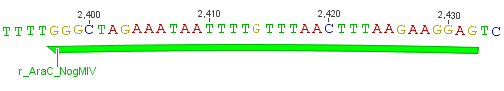
We needed a time independent induction of the ssTorA_CS-Protease_β-lactamase device and the protease. Therefore we cloned the lac-operon in front of the β-lactamase device and the arabionse induction system (araC) in front of the protease device used the Lac-operon for the we amplified the arabinose inducible induction system out of the pBAD_iGEMexpress vector. The fusion of gene construct and induction system was always performed via an NgoMIV restriction site. This restriction site was introduced with our amplification primers.
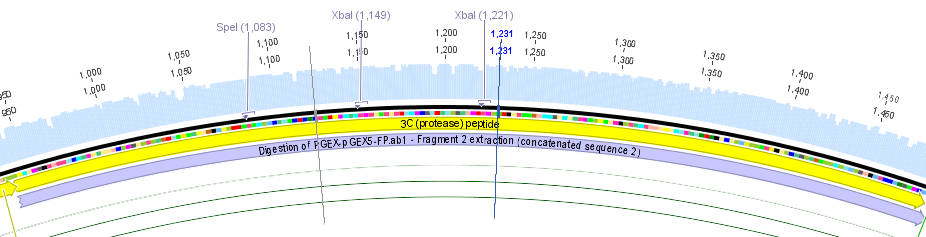
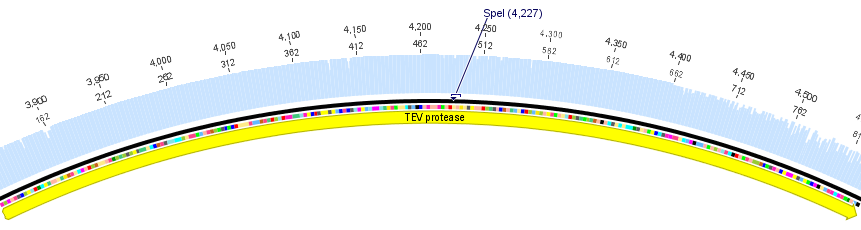
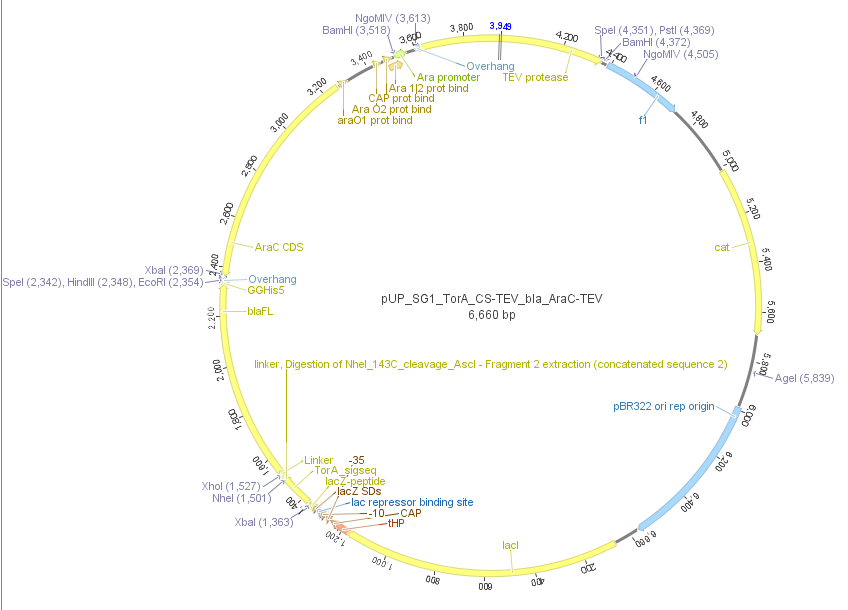
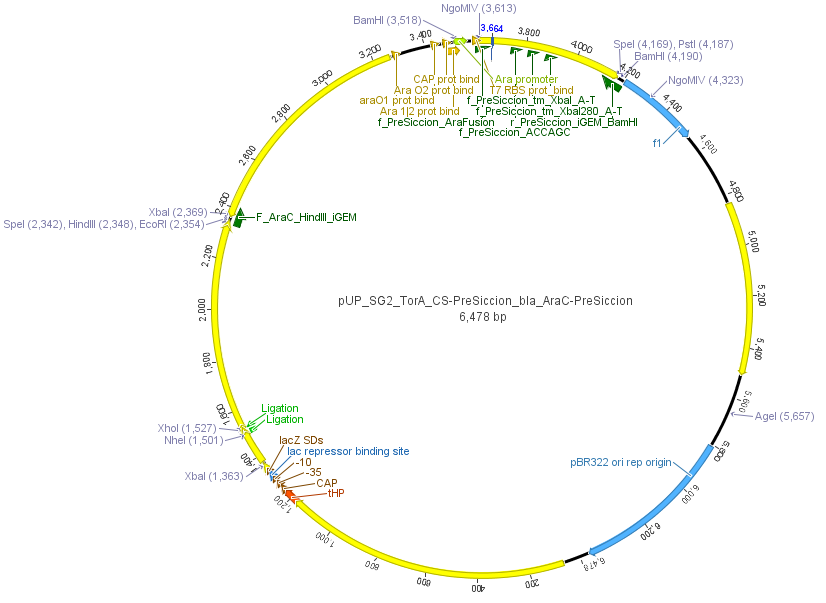
We used the RFC 23 cloning strategy to get a fusion protein of our desired protease and the induction system, which allows us to induce the protease. Both genes were amplified with PCR using two special primers generating a NgoMIV restrictioin site at the end of the induction system and in front of the protease. The amplified protease were digested with BamHI and NgoMIV, the arabinose inducible induction system with NgoMIV and HindIII. The vector, which contains the ssTorA_CS-Protease_blaFL device was digested with HindIII and BamHI. A triple ligation yielded the final construct with both devices: the protease detector and the protease generator. The two pictures below show the final vector constructs , which where used for the survival screening.
Investigation of the in vivo selection assay
Two major points have to be demonstrated: First, the export of β-lactamase over the TAT pathway has to be efficient enough to provide a certain resistance against Ampicillin and second, the activated protease is able to intimately reduce the resistance against Ampicillin by cleavage of the TorA signal sequence from the β-lactamase.
Testing the export efficiency of the protease activity detector
To test the export of the β-lactamse construct we grew an E. coli culture up to an OD600 of 0.6. The cells were then diluted to an OD of 0.002 and 100 µL were plated in triplicates under increasing Ampicillin concentrations.
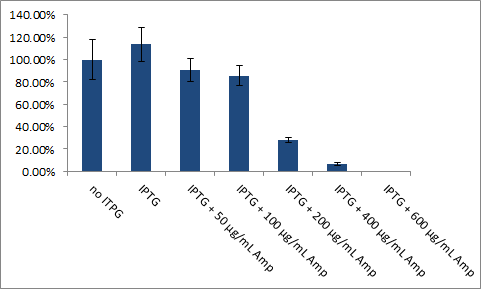
In order to test whether our idea work in vivo we investigated if the induced β-lactamse is able to confer Ampicillin resistance to E.coli. The graph below shows the amount of survived cells in percentage.
transformed two plasmids into E. coli. The first plasmid contained the ssTorA-CS-Protease-β-lactamse construct the second one the appropriate protease.
Pictures below show the surviving rate of transformed cells with constructed vector:
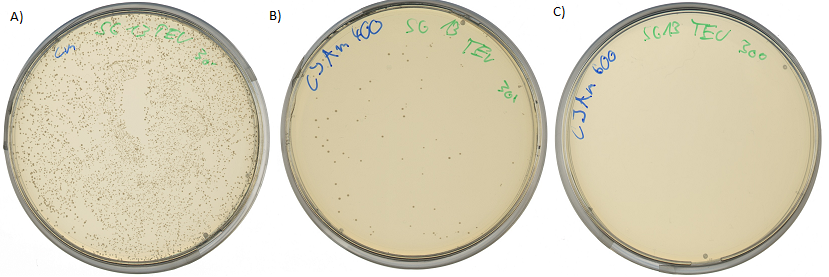
30 Grad assay wichtig!!!!!!
Testing the "Kill Switch" by activating the protease
The second major point which needs to be demonstrated is that we are able to kill the E.coli cells by activating the "Kill Switch". In the final assay we want to have the ability to induce the Protease activity detector device and the protease generate independent from each other.
Transfering and testing the library
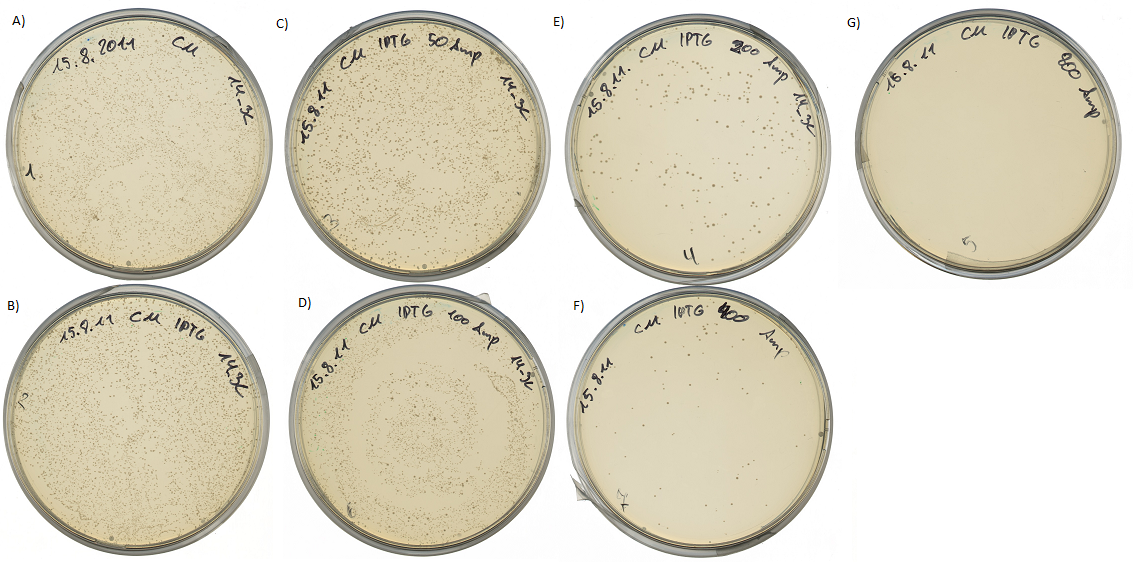
After the successful test of the protease activity we transferred the generated mdnA library into our cells. The following pictures show the amazing results. Cell growth could be detected up to 400 µg Ampicillin/mL.
Background
Twin Arginine Translocon (TAT)
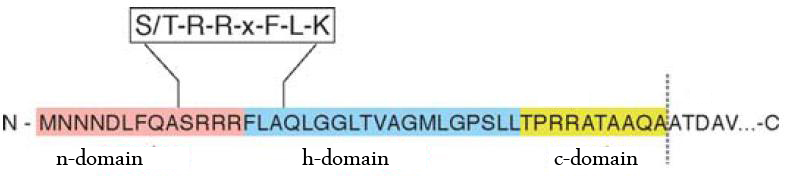
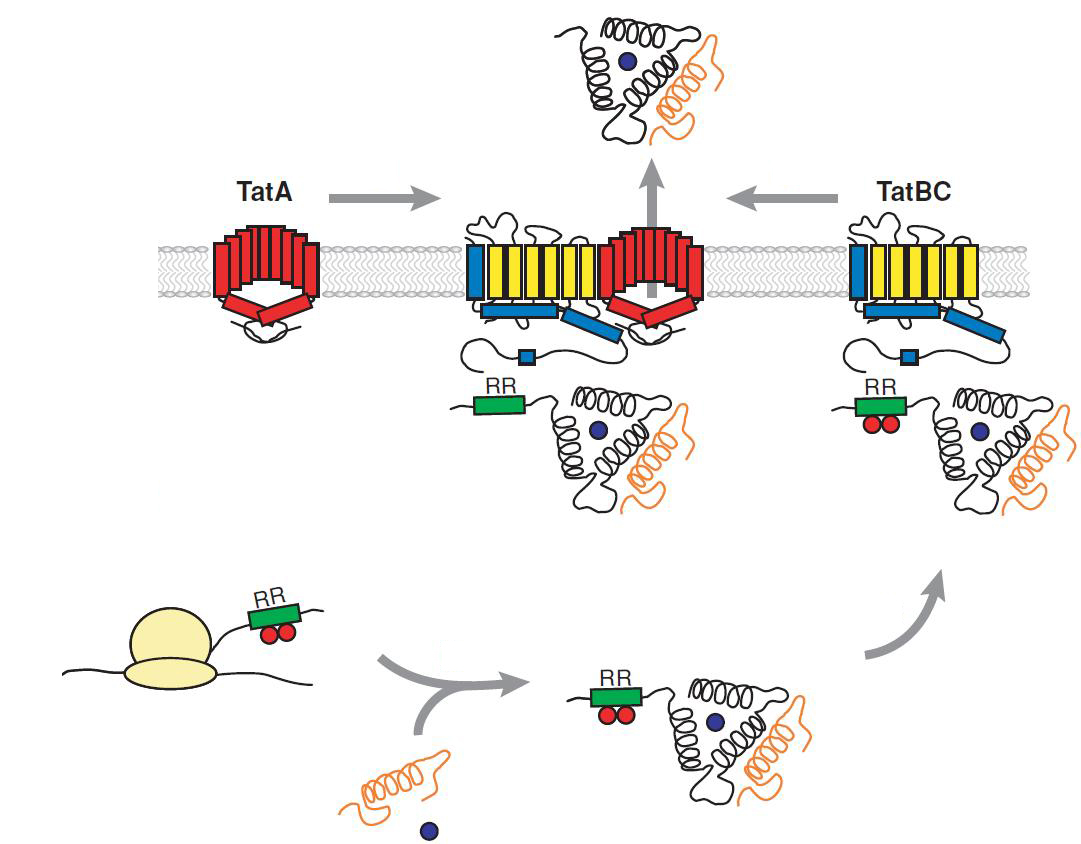
The TAT pathway is responsible for the transport of folded proteins across energy-transducing membranes and it is able of discriminating of unfolded proteins. This transport pathway is common for bacteria, archea and plants and translocates circa 6% of E.coli produced secreted proteins. The translocated proteins must have a signal peptide for targeting of the TAT-transporter, such proteins are e.g. hydrogenases, dehydrogenases and reductases. The signal sequence itself is composed of an N-terminal positive charged domain, a hydrophobic domain and a C-terminal domain.
The translocon is composed of three parts: TatA, TatB and TatC. The transport-pore is proposed to be formed during substrate binding by these three parts. The TatA, TatB and TatC proteins may form complexes of different sizes, which on the other hand form pores matching the size of the folded substrate. The transport of proteins through the TAT pathway depends on the proton motive force. Calculations by Adler & Theg showed that the transport of one folded substrate molecule requires the release of approximately 7.9x104 protons, which equals 10.000 ATP molecules. The signal sequence is cleaved off after translocation.
The used signal sequence originates from the TorA protein (Trimethylamin-N-Oxid-Reductase). It is the main respiratory enzyme which reduces TMAO under anaerobic conditions in the periplasm, where it is transported by the TAT pathway.
Tobacco Etch Virus (TEV) protease
TEV protease is the common name for the 27 kDa catalytic domain of the Nuclear Inclusion a endopeptidase (NIa) encoded by the tobacco etch virus. TEV protease is a useful reagent for cleaving fusion proteins. It recognizes a linear epitope of the general form E-Xaa-Xaa-Y -Xaa-Q-(G/S), with cleavage occurring between Q and G or Q and S. In TEV protease the serine nucleophile of the conventional Ser-Asp-His triad is a cysteine instead. This probably explains why TEV protease is resistant to many commonly used protease inhibitors.
14_3C-Protease
The 14_3C protease originates from the human rhinovirus. Rhinoviruses are the most frequent reason for infections of the upper and lower respiratory tract, also known as the cold. Because the 3C protease of human rhinovirus is necessary for the cleavage of the polyprotein translated from viral RNA it may serve as a potential target for development of antiviral targets. The recombinant type 14_3C protease from human rhinovirus (HRV 3C) recognizes the same cleavage site as the native enzyme: LeuGluValLeuPheGln↓GlyPro. The small, 22-kDa size of the protease got its optimal activity at 4°C but is still very active at 37 °C. It is commonly used for an easy tag removal after the purification of recombinant proteins carrying his-tag. The 14_3C works with a catalytic triade, containing the amino acid residues Ser-Asp-His at its active site.
References
- Cabrita, L. D., Gilis, D., Robertson, A. L., Dehouck, Y., Rooman, M. and Bottomley, S.P. (2007) Enhancing the stability and solubility of TEV protease using in silico design. Protein Sci. 16: 2360-2367
- Genest O., Ilbert M., Méjean V., Iobbi-Nivol C.,(2005) TorD, an Essential Chaperone for TorA Molybdoenzyme Maturation at High Temperature. J. Biol. Chem. 280: 15644-15648.
- Kapust R. B., Tözsér J., Fox J. D., Anderson D. E., Cherry S., Copeland T. D., Waugh D. S. (2001) Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 12:993-1000.
- Lee P. A., Tullman-Ercekand D., Georgiou G., (2006) The bacterial twin-arginine translocation pathway. Annu. Rev. Microbiol. 60:373–95
- Lucast, L. J., Batey, R. T., and Doudna, J. A. (2001). Large-scale purification of a stable form of recombinant tobacco etch virus protease. Biotechniques 30:544-550.
- Shih S. R., Chen S. J., Hakimelahi G. H., Liu H. J., Tseng C. T., Shia K. S., (2004) Selective human enterovirus and rhinovirus inhibitors: An overview of capsid-binding and protease-inhibiting molecules. Med Res Rev 24(4):449-74.
- Wanga QM, Chen SH., (2007) Human rhinovirus 3C protease as a potential target for the development of antiviral agents. Curr Protein Pept Sci. 8(1):19-27.
- Waraho D., DeLisa M. P., (2009). Versatile selection technology for intracellular protein-protein interactions mediated by a unique bacterial hitchhiker transport mechanism. PNAS 106(10):3692-7
 "
"