Team:DTU-Denmark/Project
From 2011.igem.org
(→Natural system) |
(→The natural system) |
||
| Line 7: | Line 7: | ||
== The natural system == | == The natural system == | ||
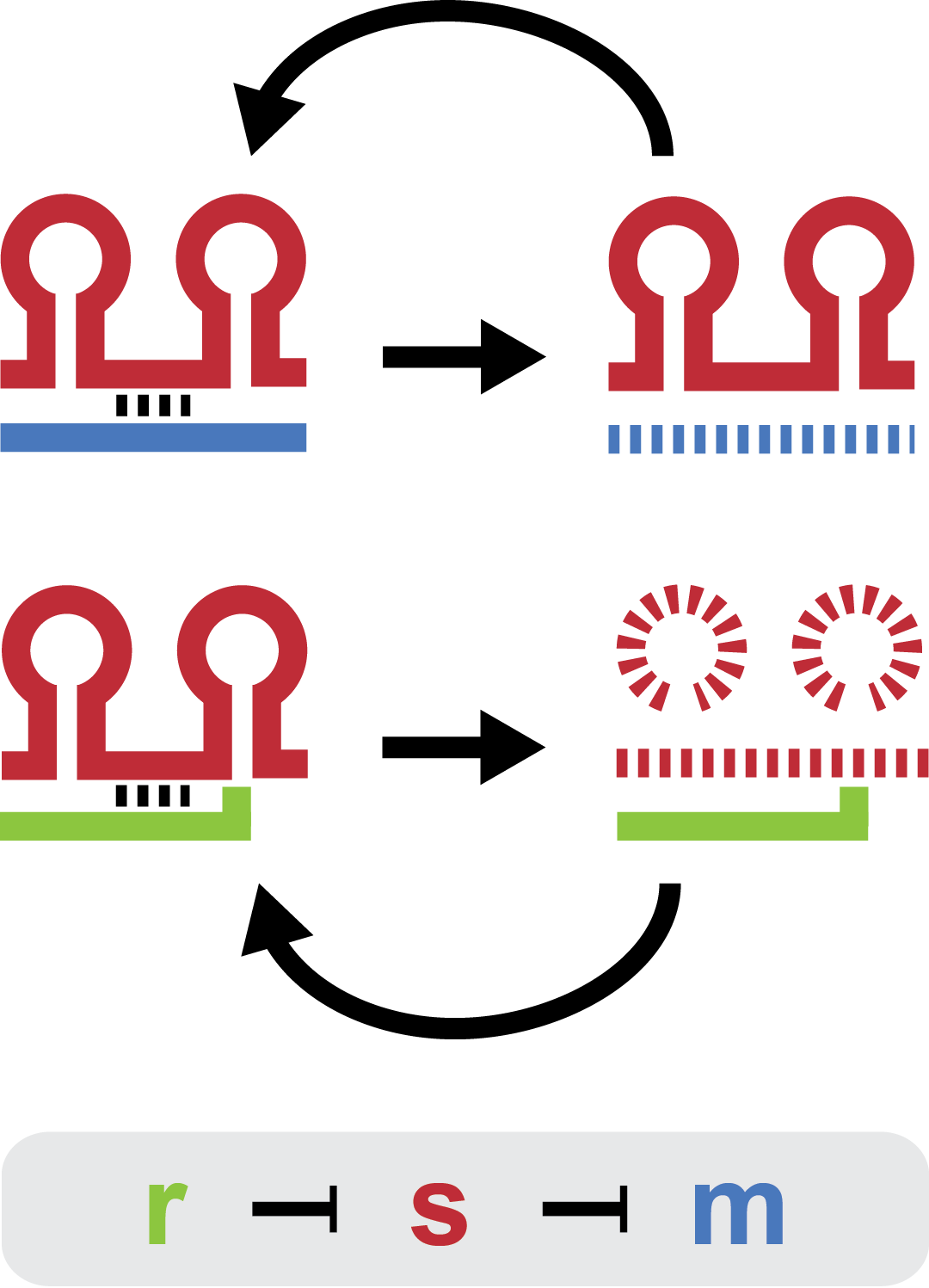
There are three elements in the natural trap RNA system that regulate chitobiose metabolism in E. coli: (1) chitoporin ''chiP'' (alias ''ybfM'') that facilitates uptake of chitosugars from the medium into the cell; (2) sRNA ''chiX'' (alias ''sroB'', ''micM'') which post-transcriptionally regulates ''chiP''; and (3) ''chiXR'' sRNA which is transcribed from intergenic region in ''chbBCARG'' operon and regulates ''chiX'' sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the ''chiP'' mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the ''chiXR'' be transcribed and subsequently its transcript can bind to ''chiX'' sRNA, releasing ''chiP'' repression. | There are three elements in the natural trap RNA system that regulate chitobiose metabolism in E. coli: (1) chitoporin ''chiP'' (alias ''ybfM'') that facilitates uptake of chitosugars from the medium into the cell; (2) sRNA ''chiX'' (alias ''sroB'', ''micM'') which post-transcriptionally regulates ''chiP''; and (3) ''chiXR'' sRNA which is transcribed from intergenic region in ''chbBCARG'' operon and regulates ''chiX'' sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the ''chiP'' mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the ''chiXR'' be transcribed and subsequently its transcript can bind to ''chiX'' sRNA, releasing ''chiP'' repression. | ||
| + | |||
| + | == The idea == | ||
| + | bla bla... | ||
== Experiments == | == Experiments == | ||
Revision as of 09:57, 20 September 2011
Project
Contents |
Abstract
Small regulatory RNA is an active area of research with untapped possibilities for application in biotechnology. Such applications include convenient gene silencing and fine-tuning of synthetic biological circuits, which are currently cumbersome processes. We have investigated a novel type of RNA regulation[2][5], where the inhibition caused by a small regulatory RNA is relieved by another RNA called trap-RNA. The system displays a large dynamic range and can uniquely target and repress any gene of interest providing unprecedented flexibility. We suspect that any level of repression is achievable by simply altering the sequences of the involved RNAs. Multiple such systems can coexist without interfering and are thus compatible with more complex designs. Furthermore the trap-RNA can be fused to any transcript in effect allowing any gene to act as an activator.
The natural system
There are three elements in the natural trap RNA system that regulate chitobiose metabolism in E. coli: (1) chitoporin chiP (alias ybfM) that facilitates uptake of chitosugars from the medium into the cell; (2) sRNA chiX (alias sroB, micM) which post-transcriptionally regulates chiP; and (3) chiXR sRNA which is transcribed from intergenic region in chbBCARG operon and regulates chiX sRNA. ChiX sRNA regulates chiP expression through binding to the Shine-Dalgarno sequence on the chiP mRNA and thus inhibits recruitment of 30S ribosome subunit and translation cannot occur. Only when chitobiose is present in the medium can the chiXR be transcribed and subsequently its transcript can bind to chiX sRNA, releasing chiP repression.
The idea
bla bla...
Experiments
Our project is a proof of concept project, showing that the E. coli sRNA sroB and the intergenic sRNA in the chbBCARG gene can be used to control gene expression by targeting the ybfM Shine-Dalgarno, and that these sRNAs can be rationally designed to target other Shine-Dalgarnos.
The experimental part of the project can be broken into 3 distinct parts:
Construction of plasmids necessary for testing our system involves taking the native system from E. coli, as well as a slightly modified system and putting them on plasmids that let us both control the expression of these components and measure the output of the system.
Strain construction involves deleting the original genes from the chromosome of a E. coli W3110 strain. Since we use the original genes, these need to be deleted from the chromosome to prevent them from interfering with our measurements.
Improving the araBAD promoter entails expanding the dynamic range of this promoter by modifying the -10 and -35 sequence of the promoter, as well as randomly changing the nucleotide sequence around and in between these sequences. Since the araBAD promoter is used in our project improving this promoter could lead to even finer control of our system.
For full description go to experiments.
Modeling
Models are developed to provide a framework for characterization and the means to incorporate the system into larger models.
A steady state analysis revealed that each system has a characteristic fold repression. The influence of parameters on the fold repression was investigated to help guide the design of the trap-RNA system. Temporal simulation provides additional tools for characterization and design. For more information got to modeling.
Bioinformatic
A bioinformatic study was performed to investigate the flexibility when engineering sRNA regulation genetically. For full analysis go to bioinformatic.
References
[1] Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proceedings of the National Academy of Sciences 97, 6640-6645(2000).
[2] Figueroa-Bossi, Nara, Martina Valentini, Laurette Malleret, and Lionello Bossi. “Caught at its own game: regulatory small RNA inactivated by an inducible transcript mimicking its target.” Genes & Development 23, no. 17 (2009): 2004 -2015.
[3] Hayashi, K. et al. Highly accurate genome sequences of Escherichia coli K-12 strains MG1655 and W3110. Molecular Systems Biology 2, 2006.0007(2006).
[4] Lambert, J.M., Bongers, R.S. & Kleerebezem, M. Cre-lox-based system for multiple gene deletions and selectable-marker removal in Lactobacillus plantarum. Applied and environmental microbiology 73, 1126-35(2007).
[5] Overgaard, Martin, Jesper Johansen, Jakob Møller‐Jensen, and Poul Valentin‐Hansen. “Switching off small RNA regulation with trap‐mRNA.” Molecular Microbiology 73, no. 5 (September 2009): 790-800.
 "
"