Team:USTC-China/Project/results
From 2011.igem.org
(→Verification of ΔcheZ strain (protocol)) |
(→Verification of the function of the constructed Aptamer-cheZ part(protocol)) |
||
| Line 21: | Line 21: | ||
<p> From the results shown in Figure2. We can be sure that the Aptamer-cheZ part actually works effectively, especially on 0.3% Semi-solid medium.</p> | <p> From the results shown in Figure2. We can be sure that the Aptamer-cheZ part actually works effectively, especially on 0.3% Semi-solid medium.</p> | ||
[[File:X().jpg|center|650px| ]] | [[File:X().jpg|center|650px| ]] | ||
| - | <p align=center>Figure2. The growing state of the reprogrammed bacteria with Aptamer-cheZ part(Left:0.3%agar with 0mM Thephylline, Right:0.3%agar with 0.25mM Thephylline)</p> | + | <p align=center class="ppp">Figure2. The growing state of the reprogrammed bacteria with Aptamer-cheZ part(Left:0.3%agar with 0mM Thephylline, Right:0.3%agar with 0.25mM Thephylline)</p> |
<html><a name="Verification of the original Toggle-switch from PKU" id=""></a></html> | <html><a name="Verification of the original Toggle-switch from PKU" id=""></a></html> | ||
Revision as of 19:53, 5 October 2011

Experiment results of the Basic Design
and Discussion
and Discussion
1.Verification of ΔcheZ strain
2.Verification of the function of the constructed Aptamer-cheZ part
3.Verification of the original Toggle-switch from PKU
4.Verification of the modified Toggle-switch
5.Test of Incorporated Aptamer-cheZ part into one side of the original Toggle-switch
6.Test of Incorporated Aptamer-cheZ part into one side of the modified Toggle-switch
Verification of ΔcheZ strain (protocol)
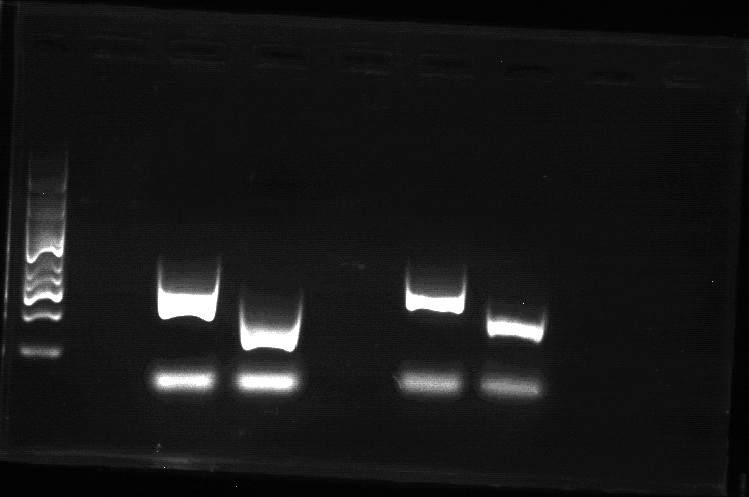
The size of colonies of E.coli strain RP1616 was much smaller than that E.coli steain RP437 under the same circumstance and after same period of incubation time(about 10h), and the result is shown in Figure1. In colony PCR using the primers of cheZ gene following , RP437 absolutely has much more outcomes than RP1616 and we conclude that RP1616 is actually a ΔcheZ strain.
Figure1.The result of colony PCR(From left to right, the first lane is the marker, the third and the forth lane is the PCR outcome of strain RP437 and strain RP1616, the sixth and the seventh lane is the PCR outcome of strain RP437 and strain RP1616.)
Verification of the function of the constructed Aptamer-cheZ part(protocol)
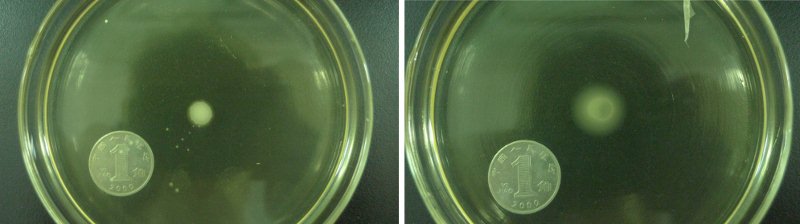
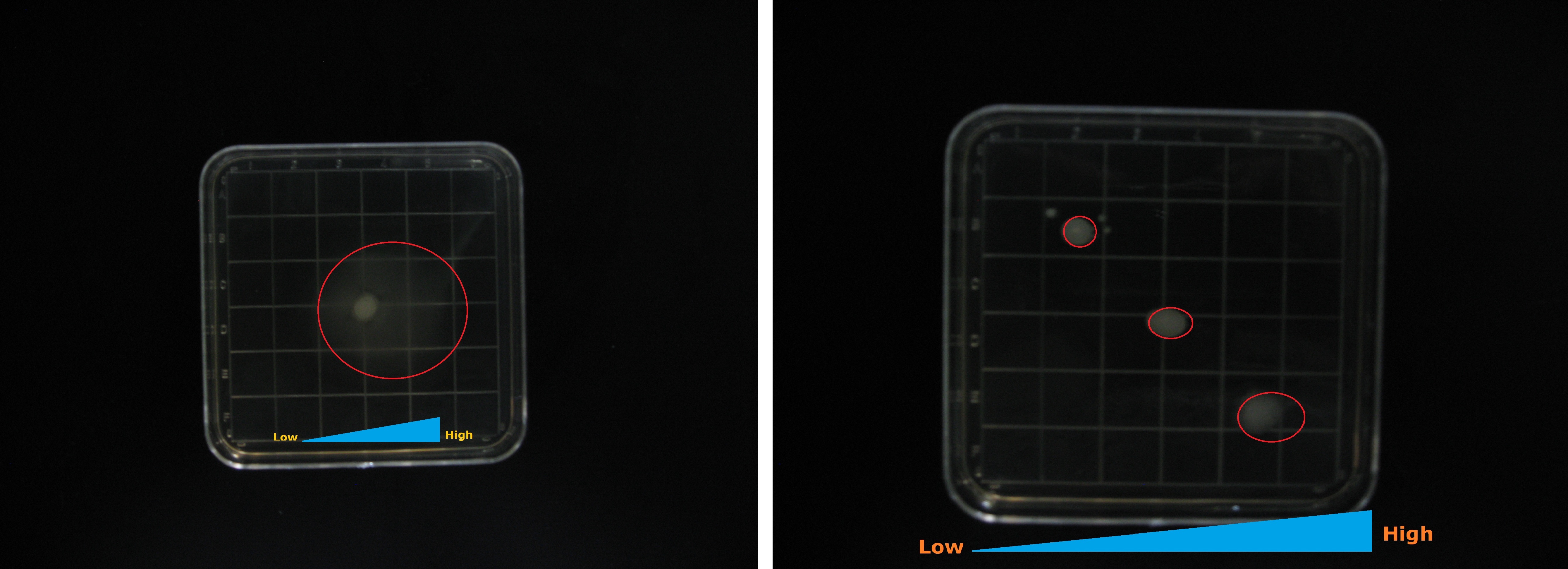
From the results shown in Figure2. We can be sure that the Aptamer-cheZ part actually works effectively, especially on 0.3% Semi-solid medium.
Figure2. The growing state of the reprogrammed bacteria with Aptamer-cheZ part(Left:0.3%agar with 0mM Thephylline, Right:0.3%agar with 0.25mM Thephylline)
Verification of the original Toggle-switch from PKU(protocol)
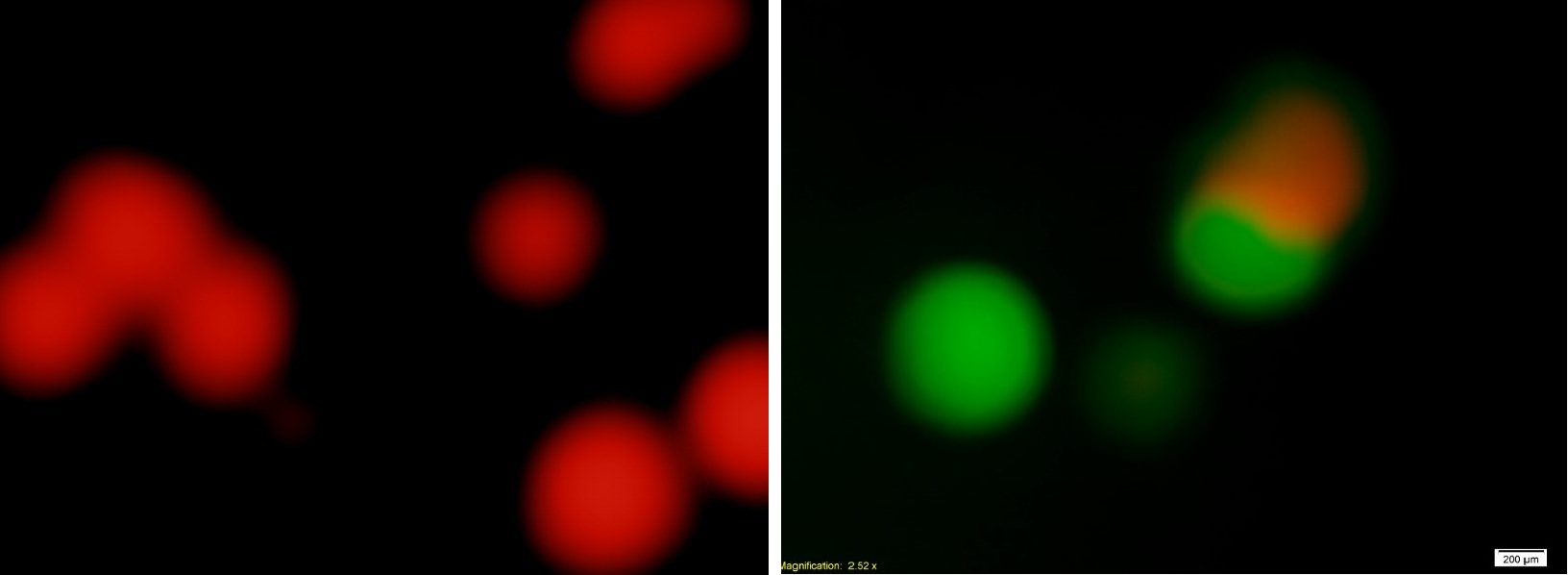
Figure3.A conlony of the bacteria with the original Toggle Switch exhibit two different states(4X Objective)
By calculating with the help of the fluorescence microscope, the ratio between the numbers of colonies with RFP and colonies with GFP ≈ 8:25. It means the original Toggle Switch is not fit for our purpose, so we modify the original by luxPR-cI device.
Verification of the modified Toggle-switch (protocol)
Figure4. The conlonies of the bacteria with the modified Toggle Switch exhibit two different states or just one state(4X Objective)
By calculating with the help of the fluorescence microscope, the ratio between the numbers of colonies with RFP and colonies with GFP ≈ 6:1. It is useful to our project design.
Test of Incorporated Aptamer-cheZ part into one side of the original Toggle-switch (protocol)
Figure5.The growing state of the reprogrammed bacteria with original Toggle-switch-Aptamer-cheZ Device(Red circle:the range of motion, Blue Triangle:the gradient of thepphylline)
Figure6.The fluorescent Photo(4X Objective) of the colony shown in Figure5 Left.
According to the results above, by using this device we can not make the reprogrammed bacteria be divided into two different states effectively, and because of the bias to the green fluorescence and moving towards the high concentration of the theophylline, the bacteria trending to stay are very rare.
Test of Incorporated Aptamer-cheZ part into one side of the modified Toggle-switch (protocol)
Figure7.
Figure7 shows the fluctuation of expression of the modified Toggle Switch device, the number of bacteria with red fluorescent decreases from right to left.
Figure8.
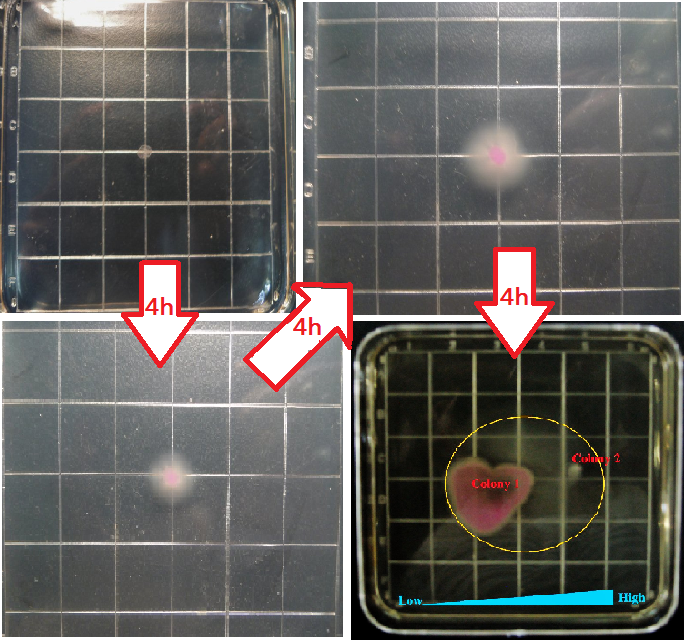
Figure8 shows four different growing states of the reprogrammed bacteria, in which the yellow circle represent the range of motion of reprogrammed bacteria(from the second tube(from right to left) shown in Figure7), and from left to right the concentration of the theophylline increases.
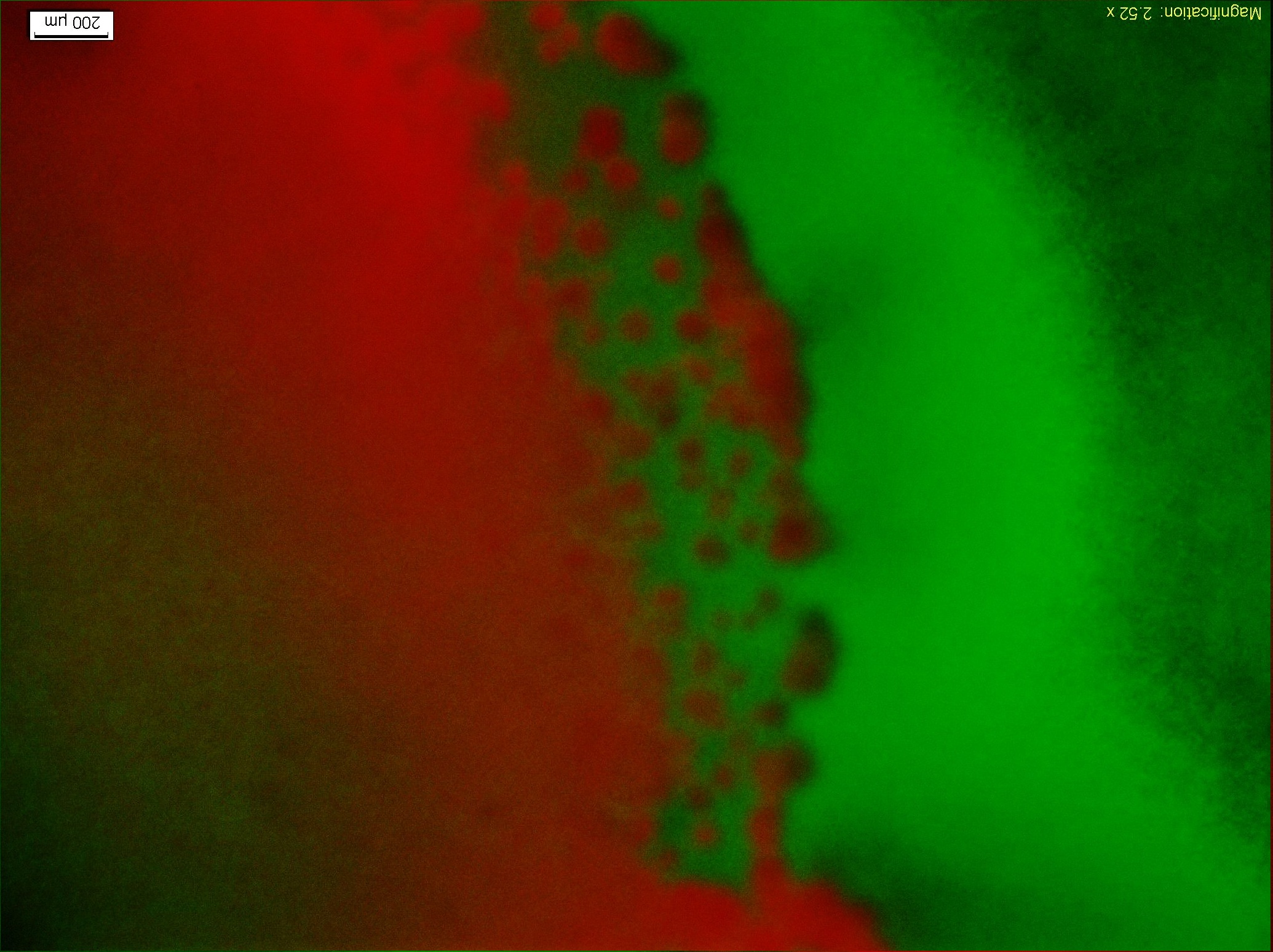
Figure9.The fluorescent Photo(4X Objective)of the colony1 in the lower right of Figure8, and from left to right the concentration of the theophylline increases.
Figure9 clearly display the dividing line between the reprogrammed bacteria at two different states. This means the modified Toggle-switch-Aptamer-cheZ actually work as our design.
 "
"