Team:Tokyo Metropolitan/Project/Visualization
From 2011.igem.org
Visualization
Visualize device is system to check for conjugative transfer function. In this project, we use this device to check for function of killer device.
Background
We referred "Hop-count" of 2007 Peking in basic structure. Hop-count system is showing fluorescence with two conjugations. (fig.1) We use not only designed device also F plasmid and R plasmid. These F, R plasmid loses oriT and TraJ. This device is composed of promoter, oriT and TraJ of F plasmid, oriT and TraJ of R plasmid and GFP (fig. 2) The device which never conjugate expresses Traf F and complement lost TraJ function of F plasmid.
Then conjugative transfer starts from oriT F. The device which once transferred cut between oriT F-oriT F and constitutive expresses TraJ R. (fig.3) The same as, conjugative transfer starts from R. This is the device lost oriT R-oriT R which twice conjugate. Therefore, the device which twice conjugates expresses GFP. (fig.4)
See also 2007 Peking "Hop-count".
Construction
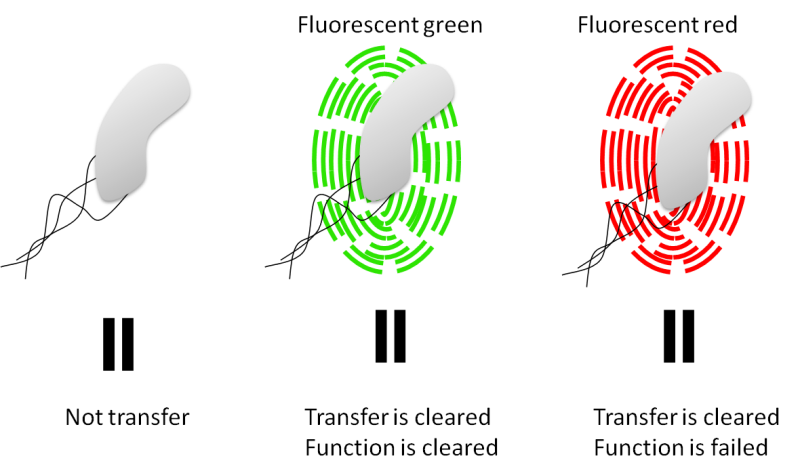
"Hop-count" system can check only 2nd conjugation. Therefore, we could not use this system to check for killer device. The reason is that if killer device works regularly donors are dying. Donor BeE.coli sends killer gene to target E.coli when 1st conjugation, thus target E.coli cannot do 2nd conjugation. Important points are that we can check 1st conjugation and 2nd conjugation in case that killer gene works irregularly. Therefore we can check visibly three cases: never conjugate, once conjugate and twice conjugate. (fig.5)
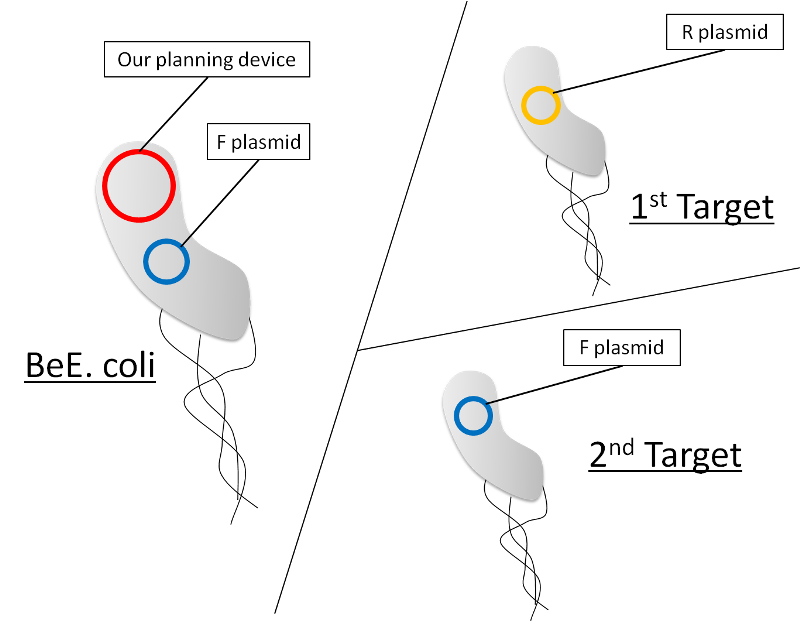
In this time, we make a difference between host E.coli and target E.coli: thus we can make easily device.
We do not use TraJ. We put decrease expression part between oriT-F and put GFP between oriT-R. (fig.6) Cut down decrease expression system when 1st conjugation and GFP when 2nd conjugation then expresses RFP. F plasmid and R plasmid loses each oriT.
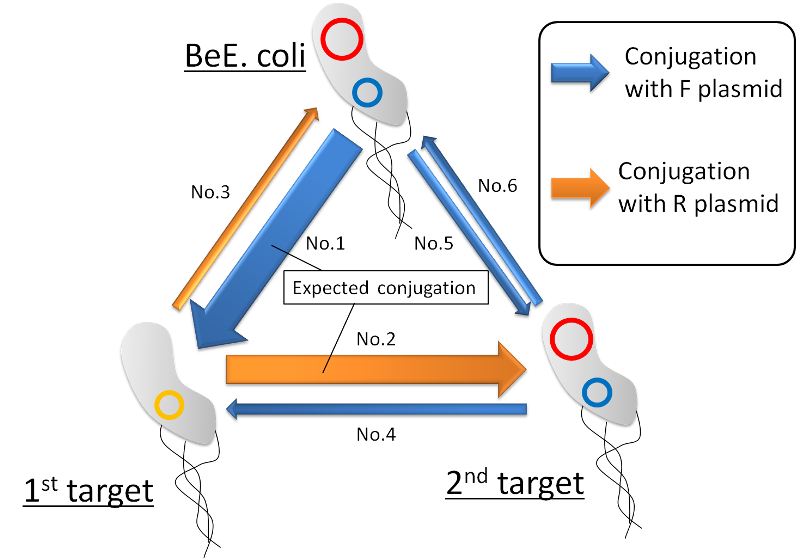
Making BeE.coli, 1st target and 2nd target In theory, we think 6 transfer ways. At first no.1 or no.5 occurs but mating efficiency of no.5 decreases than pair of F+ and F- because no.5 is conjugation both of F+. Therefore no.1 occurs. After transferring to 1st target, no.2 or no.3 occurs. In case no.2 occurs, GFP between oriT R falls out by R transfer. However if no.3 occurs, plasmid of no.3 do not save because the back born is same as.
Finally, no.4 and no.6 eliminates the same reason for no.3. Considering the circumstances, following BeE.coli -> 1st target -> 2nd target is most expected conjugation. (fig.7)
 "
"