Team:Bielefeld-Germany/Project/Description
From 2011.igem.org


Contents |
Project description
The development of sensitive and selective biosensors is an important topic in synthetic biology. Biosensors can be used in a wide range of applications - from the detection of toxic compounds in the environment up to clinical diagnostics. Because cells have to sense their surroundings, there are a lot of natural systems that can be used as biosensors. Unfortunately, cellular biosensors often have interfering properties that preclude any practical application. Common problems are the limited use outside the laboratory due to the use of genetically engineered cells, the low durability because of the usage of living cells and the appearance of undesired signals induced by endogenous metabolic pathways.
To solve these problems, the iGEM-Team Bielefeld 2011 aims at developing a cell-free bisphenol A (BPA) biosensor based on a coupled enzyme reaction fused to S-layer proteins for everyday use. Bisphenol A is a harmful substance which is used in the production of polycarbonate. To detect BPA, it is degraded by a fusion protein under formation of NAD+ which is detected by an NAD+-dependent enzymatic reaction with a molecular beacon. Both enzymes are fused to S-layer proteins which build up well-defined nanosurfaces and are attached to the surface of beads. By providing these nanobiotechnological building blocks the system is expandable to other applications.
An overview of our project is shown in Figure 1. The background and the state of the art of each subproject is described below this figure. To have a quick insight of what is happining in our lab take a look in our labjournal.
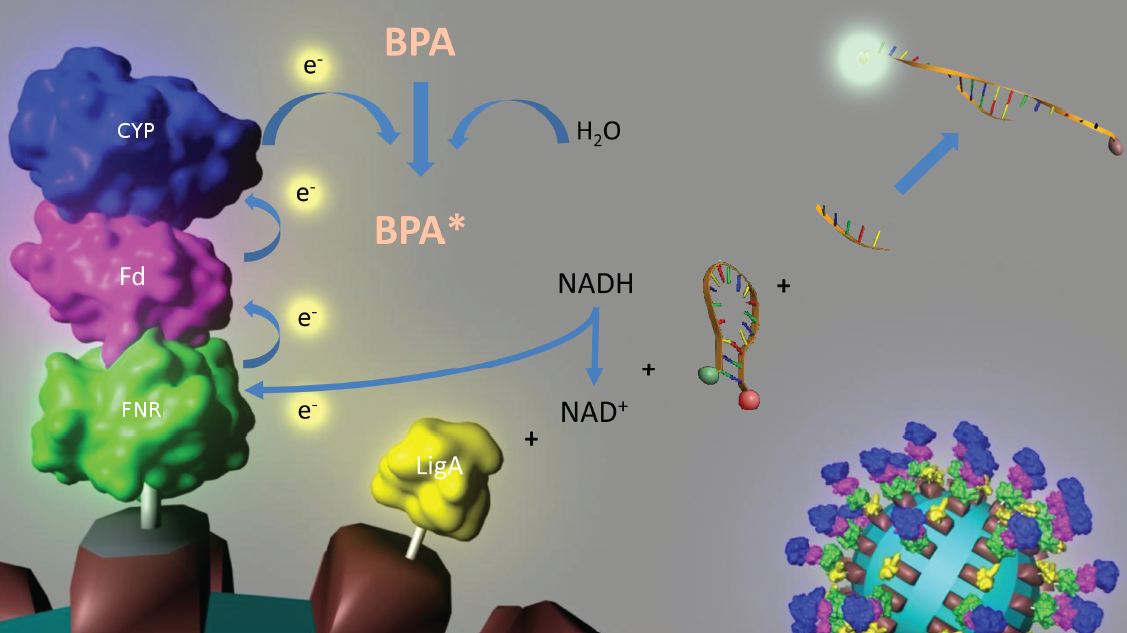
S-layer
S-layers (crystalline bacterial surface layer) are crystal-like layers consisting of multiple protein monomers and can be found in various (archae-)bacteria. They constitute the outermost part of the cell wall. Especially their ability for self-assembly into distinct geometries is of scientific interest. At phase boundaries, in solutions and on a variety of surfaces they form different lattice structures. The geometry and arrangement is determined by the C-terminal self assembly-domain, which is specific for each S-layer protein. The most common lattice geometries are oblique, square and hexagonal. By modifying the characteristics of the S-layer through combination with functional groups and protein domains as well as their defined position and orientation to each other (determined by the S-layer geometry) it is possible to realize various practical applications (Sleytr et al., 2007). The usability of such well defined nano-lattice structures is far-reaching from ultrafiltration membranes to the development of immobilized biosensors.
Especially for the production of cell-free biosensors, functional fusion proteins are of great importance. Kainz et al. (2010) fused fluorescent proteins with an S-layer glycoprotein from Geobacillus stearothermophilus. They demonstrated that the properties of the fusion protein were similar to the native fluorescent protein. The intensity of the fluorescence, the lifetime and the adsorption spectra showed comparable behavior at different pH-values. Enzymes fused to immobilized S-layers showed a significantly longer durability and were more stable against physical and chemical treatment.(Schäffer et al., 2007, Tschiggerl et al., 2008).
The iGEM-Team Bielefeld aims at the assembly, production and immobilization of S-layer fusion proteins for the detection of BPA by a coupled enzymatic reaction. S-layers from five different organisms are employed. The provision of various S-layers with different geometries offers the possibility for the scientific community to create functional nanobiotechnological surfaces with simple and standardized methods, quasi do it yourself nanobiotechnology. First, different fusion proteins with fluorescent proteins and a luciferase are created. The functionality and efficiency of the immobilization to various materials such as silicon dioxide or cellulose is then characterized by measuring the fluorescence and luminescence, respectively.
Bisphenol A degradation
In 2005, Sasaki et al. isolated a soil bacterium from the Sphingomonas genus which is able to degrade the environmental poison bisphenol A (BPA) with a unique rate and efficiency compared to other BPA degrading organisms. This strain was called Sphingomonas bisphenolicum AO1 and is able to completely decompose 120 mg BPA L-1 in about 6 hours. Three genes which are responsible for the first step of the effective BPA degradation by S. bisphenolicum AO1 were identified: a cytochrome P450 (bisdB), a ferredoxin (bisdA) and a ferredoxin-NADP+ oxidoreductase (FNR) (Sasaki et al., 2005b). The bisdAB genes from S. bisphenolicum AO1 were isolated, transformed into and expressed in E. coli and enabled this bacterium to degrade BPA, too (Sasaki et al., 2008). Furthermore, the BisdAB proteins from S. bisphenolicum AO1 were able to degrade BPA in a cell free system in which spinach reductase was added (Sasaki et al., 2005b). Therefore, we assume that the BisdAB proteins also work in a cell free system together with the ferredoxin-NAD(P)+ oxidoreductase from E. coli. The suggested reaction mechanism of the first BPA degradation step is shown in the project overview image above and in the animated picture of Figure 2.
In 2008, the iGEM team from the University of Alberta submitted the codon usage optimized bisdAB genes from S. bisphenolicum AO1 to the registry of standard biological parts in the so-called Freiburg BioBrick assembly standard (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>). Via this assembly standard it is very easy to build fusion proteins. These already existing protein domains will be fused together with the NAD(P)+ oxidoreductase gene from E. coli to the gene encoding the fusion protein FNR:Fdbisd:P450bisd which subsequently will be fused to an S-layer gene. We have already shown that the Fdbisd:P450bisd fusion protein is degrading BPA more effectively in E. coli than the polycistronic bisdAB gene (data here).
NAD+ detection
Our selected NAD+ detection method utilizes a molecular beacon based approach. These have been initially described in 1996 as nucleic acid probes that fluoresce upon hybridization (Tyagi et al., 1996). For this effect the ends of a single-stranded DNA molecule are labeled with a fluorophore as well as with an appropriate quencher. Due to a formed stem-loop both are in such a close proximity to each other that no fluorescence signal is detected.
The molecular beacons closed state can be applied to a bioassay detecting NAD+ even in very low concentrations (Tang et al., 2011). Using two complementary targets hybridizing side-by-side with the hairpin enables NAD+-dependent DNA ligation (nick-closing) by E. coli DNA ligase. Only after closing the gap between both hybridized targets the stem opens and the secondary structure gets broken down to a linearized probe-target hybrid. The immediate consequence is a disruption of the close proximity of the fluorophore and the quencher, so that an excitation with light is converted into a visible fluorescence signal. Since NAD+ concentration determines DNA ligase activity, which is responsible for the formation of the molecular beacons open state, the emerging fluorescence directly correlates with the NAD+ concentration. Additionally, the highly selective NAD+ bioassay has a low limit of detection compared to other methods.
Because of the signal stability and the suitability for daily use the NAD+ bioassay can be coupled to NADH-dependent BPA degradation in the context of biosensing.
References
Kainz B, Steiner K, Möller M, Pum D, Schäffer C, Sleytr UB, Toca-Herrera JL (2010) Absorption, Steady-State Fluorescence, Fluorescence Lifetime, and 2D Self-Assembly Properties of Engineered Fluorescent S-Layer Fusion Proteins of Geobacillus stearothermophilus NRS 2004/3a, Biomacromolecules 11(1):207-214.
Sasaki M, Maki J, Oshiman K, Matsumura Y, Tsuchido T (2005a) Biodegradation of bisphenol A by cells and cell lysate from Sphingomonas sp. strain AO1, Biodegradation 16(5):449-459.
Sasaki M, Akahira A, Oshiman K, Tsuchido T, Matsumura Y (2005b) Purification of Cytochrome P450 and Ferredoxin, Involved in Bisphenol A Degradation, from Sphingomonas sp. Strain AO1, Appl Environ Microbiol 71(12):8024-8030.
Sasaki M, Tsuchido T, Matsumura Y (2008) Molecular cloning and characterization of cytochrome P450 and ferredoxin genes involved in bisphenol A degradation in Sphingomonas bisphenolicum strain AO1, J Appl Microbiol 105(4):1158-1169.
Schäffer C, Novotny R, Küpcü R, Zayni S, Scheberl A, Friedmann J, Sleytr UB, Messner P (2007) Novel Biocatalysts Based on S-Layer Self-Assembly of Geobacillus Stearothermophilus NRS 2004/3a: A Nanobiotechnological Approach, Small 3(9):1549-1559.
Sleytr UB, Huber C, Ilk N, Pum D, Schuster B, Egelseer EM (2007) S-layers as a tool kit for nanobiotechnological applications, FEMS Microbiol Lett 267(2):131-144.
Tang Z, Liu P, Ma C, Yang X, Wang K, Tan W, Lv X (2011) Molecular Beacon Based Bioassay for Highly Sensitive and Selective Detection of Nicotinamide Adenine Dinucleotide and the Activity of Alanine Aminotransferase, Anal Chem 83(7):2505-2510.
Tschiggerl H, Breitwieser A, de Roo G, Verwoerd T, Schäffer C, Sleytr U B (2008) Exploitation of the S-layer self-assembly system for site directed immobilization of enzymes demonstrated for an extremophilic laminarinase from Pyrococcus furiosus, Journal of Biotechnology 133:403-411
Tyagi S, Kramer FR (1996) Molecular beacons: probes hat fluoresce upon hybridization, Nature Biotechnology 14:303-308.
 "
"
