Team:Bielefeld-Germany/Labjournal
From 2011.igem.org


Labjournal: On this page we summarize the (successful) results and achievements of our teamwork.
Week 1: 2nd - 8th May
Bisphenol A:
- cloning of <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> behind weak (<partinfo>J23103</partinfo>) and medium strong (<partinfo>J23110</partinfo>) constitutive promoter (each part and both parts polycistronic)
- cloning of fusion protein between <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>, also assembly behind weak and medium strong constitutive promoter
- testing and establishing of HPLC method for BPA detection
- expression of the successfully assembled BPA degrading BioBricks in E. coli TOP10
S-layer:
- successful PCR on the S-layer genes of Corynebacterium glutamicum and Corynebacterium crenatum
- successful cloning of the S-layer gene of Brevibacterium flavum without TAT-sequence
Organizational:
- meeting with Bielefeld Marketing to discuss our participation at the science festival GENIALE in Bielefeld
Week 2: 9th - 15th May
Bisphenol A:
- assembly of <partinfo>K157011</partinfo> behind existing BPA degrading parts (for purification and testing of <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> in a cell free system)
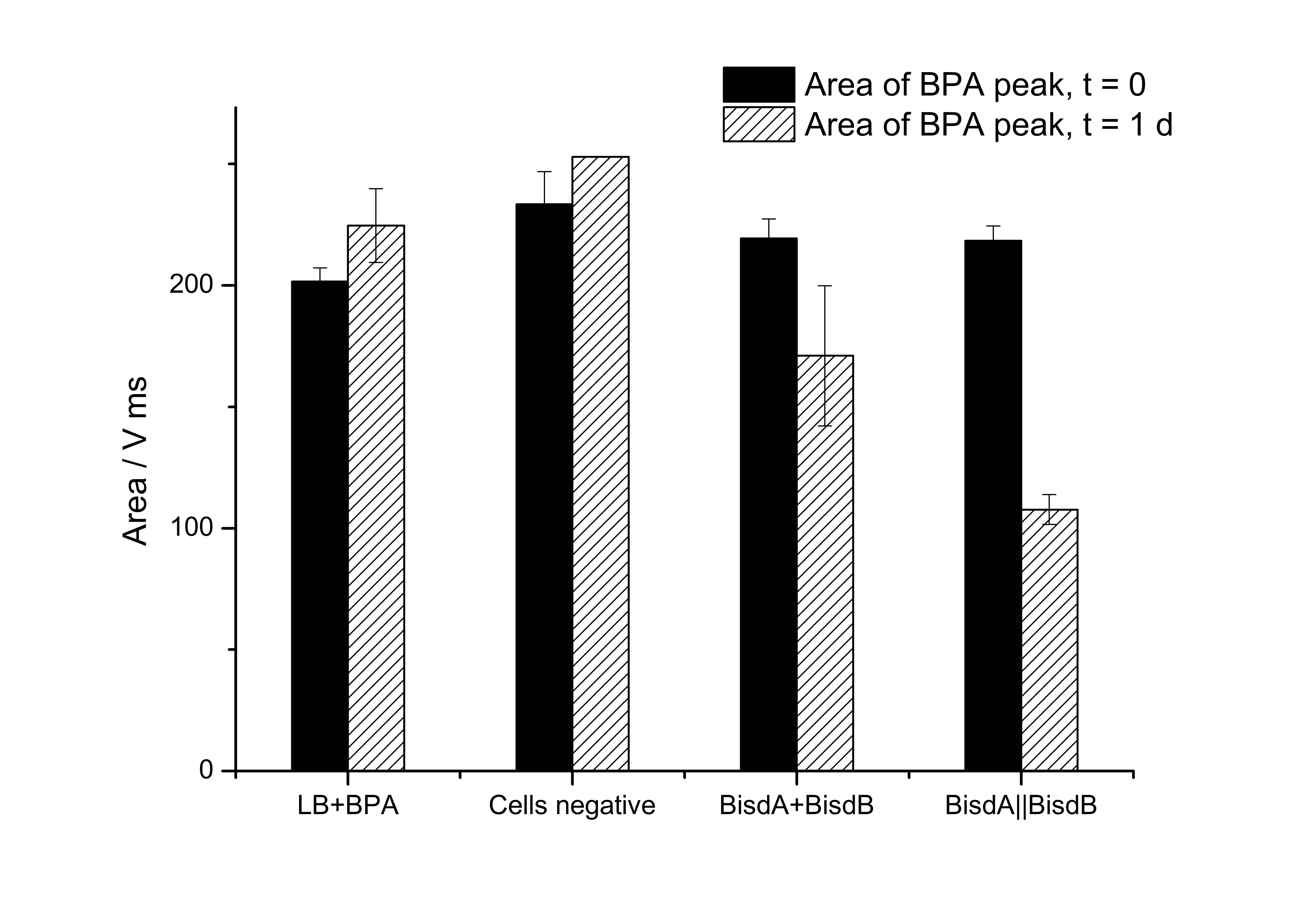
- HPLC results: Fusion protein between <partinfo>K123000</partinfo> and <partinfo>K123001</partinfo> can degrade BPA and seems to work better than the polycistronic version (compare Figure 1)
S-layer:
- expression of the S-layer gene without TAT-sequence of C. glutamicum in E. coli KRX and BL21-Gold(DE3) after sequencing gave correct results
- successful PCR on the S-layer gene of Corynebacterium halotolerans
Organizational:
- moving to our own room in the CeBiTec
Week 3: 16th - 22nd May
Bisphenol A:
- successful PCR on the NADP+ oxidoreductase gene from E. coli TOP10
S-layer:
- successful cloning of the complete S-layer gene cspB of C. glutamicum, C. crenatum and C. halotolerans
- successful cloning of the S-layer genes of B. flavum and C. halotolerans without TAT-sequence, without lipid anchor and without both (only self-assembly domain)
Week 4: 23rd - 29th May
Bisphenol A:
- establishing a new method for analysis of BPA concentrations (extraction + LC-ESI-QTOF-MS)
Organizational:
- arrange a BBQ for our workgroup in the CeBiTec to get to know our co-workers
- substantiating our contribution to the GENIALE
Week 5: 30th May - 5th June
Bisphenol A:
- beginning of first characterization experiments for BPA degrading BioBricks (<partinfo>K123000</partinfo> and <partinfo>K123001</partinfo>)
Organizational:
- meeting with Prof. Alfred Pühler to plan our contribution to the CeBiTec symposium in July
Week 6: 6th - 12th June
Organizational:
- presentation of the iGEM competition at the 2nd BIO.NRW (PhD) Student Convention
Week 7: 13th - 19th June
S-layer:
- successful fusion of modified cspB genes of C. glutamicum and C. halotolerans with a monomeric RFP (BBa_E1010) using Gibson assembly.
- C.glutamicum:
- C. halotolerans:
- First expression of K525133 in E. coli KRX to test different induction time points und L-rhamnose concentrations. A higher inducer concentration results in a decreasing maximum and final optical density (OD600) but in a higher fluorescence level. The variations of the induction time point has no significant influence on growth and the progression of fluorescence.
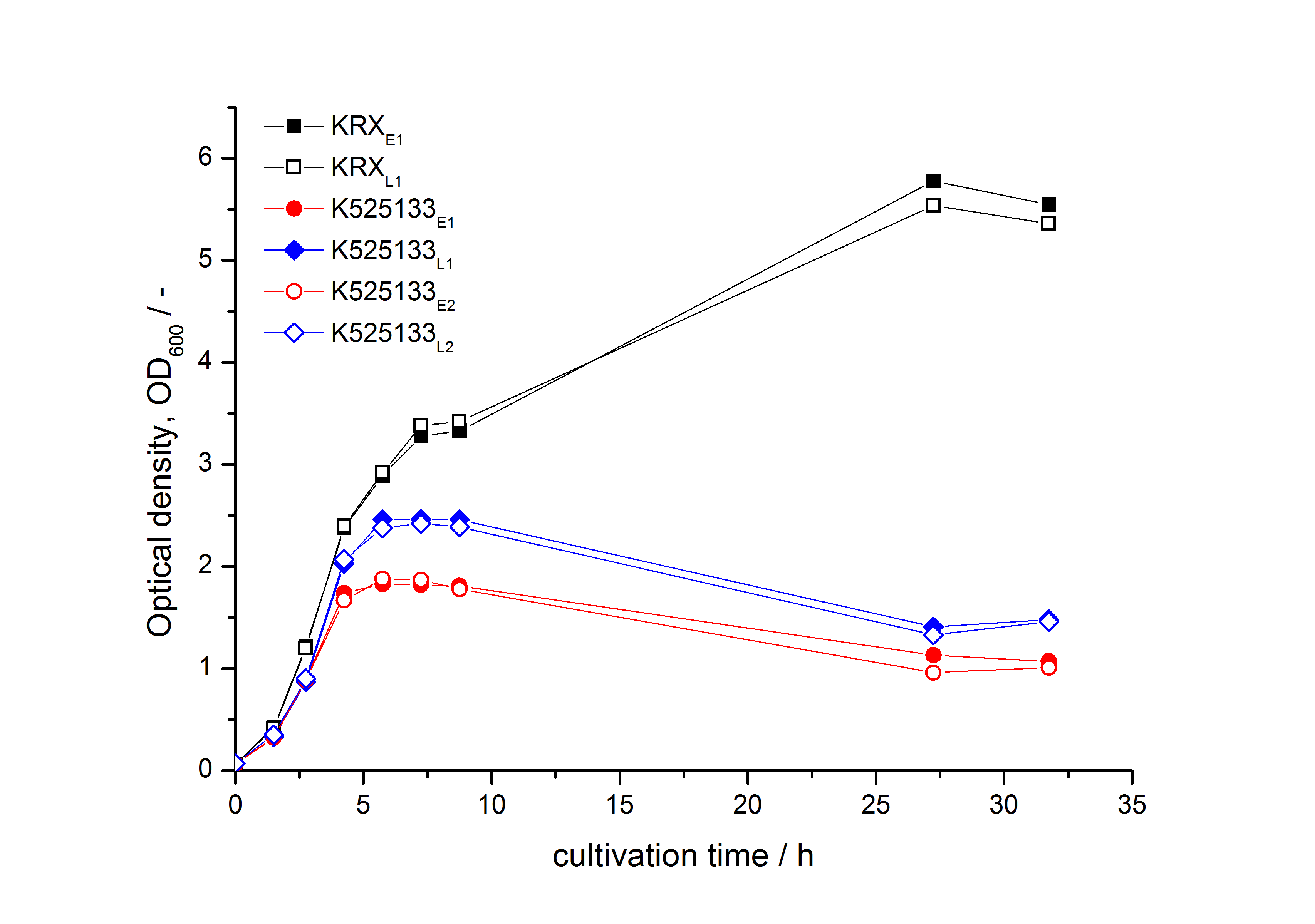
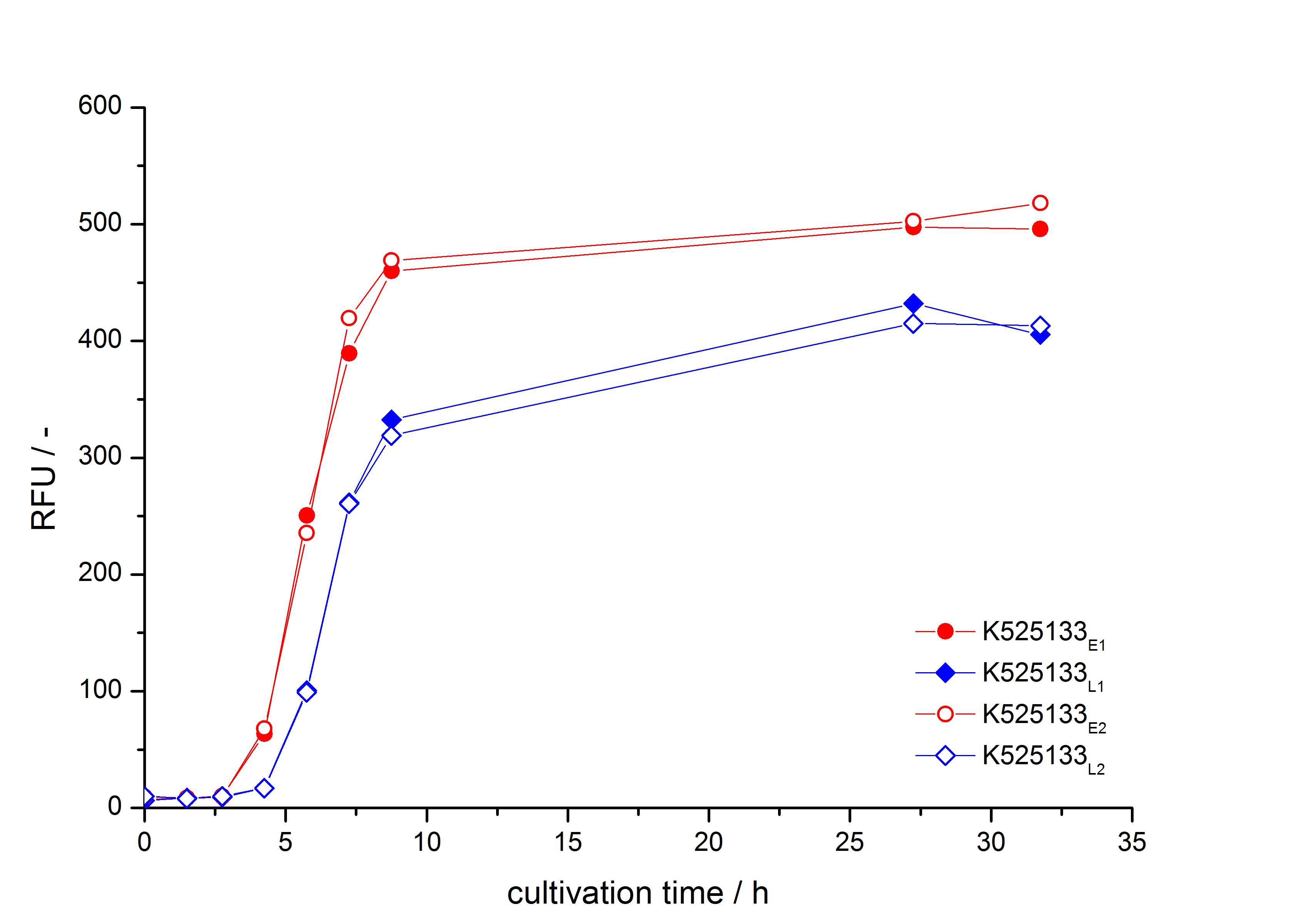
Bisphenol A:
- cloning of NADP oxidoreductase in pJET1.2 finally successful -> waiting for sequencing results to remove illegal restriction sites
Week 8: 20th - 26th June
S-layer:
- successful cloning of K525232 using Gibson assembly.
- K525232: fusion of modified cspB of C. halotolerans without lipid anchor and TAT-sequence <partinfo>K525222</partinfo> with BBa_E1010.
Organizational:
- finishing our first press release
- all devices (thermocycler etc.) and materials (competent cells, polymerase, kits) from our sponsors arrived
Week 9: 27th June - 3rd July
EXAMS !
Week 10: 4th July - 10th July
Bisphenol A:
- experiments on the influence of temperature, promoter strength and the characteristics of the fusion protein <partinfo>K123000</partinfo> || <partinfo>K123001</partinfo> on BPA degradation
S-layer:
- Expression of K525131 (with TAT-sequence and lipid anchor), K5252133 (with lipid anchor), K525232 (nothing); K525233 (with lipid anchor) and K525234 (with TAT-sequence) in E. coli KRX to test the functional efficiency of Corynebacterium TAT-sequence in E.coli and the effect of the autoinduction protocol. After one day of cultivation, an increased fluorescence in all periplasm fractions of E. coli expressing S-layer constructs with TAT-sequence could be recognized. This is an indication that the Corynebacterium TAT-signal sequence is functional in E. coli. Comparing the manual and the autoinduction protocol, no significant changes in fluorescence (after 14 h) are identifiable.
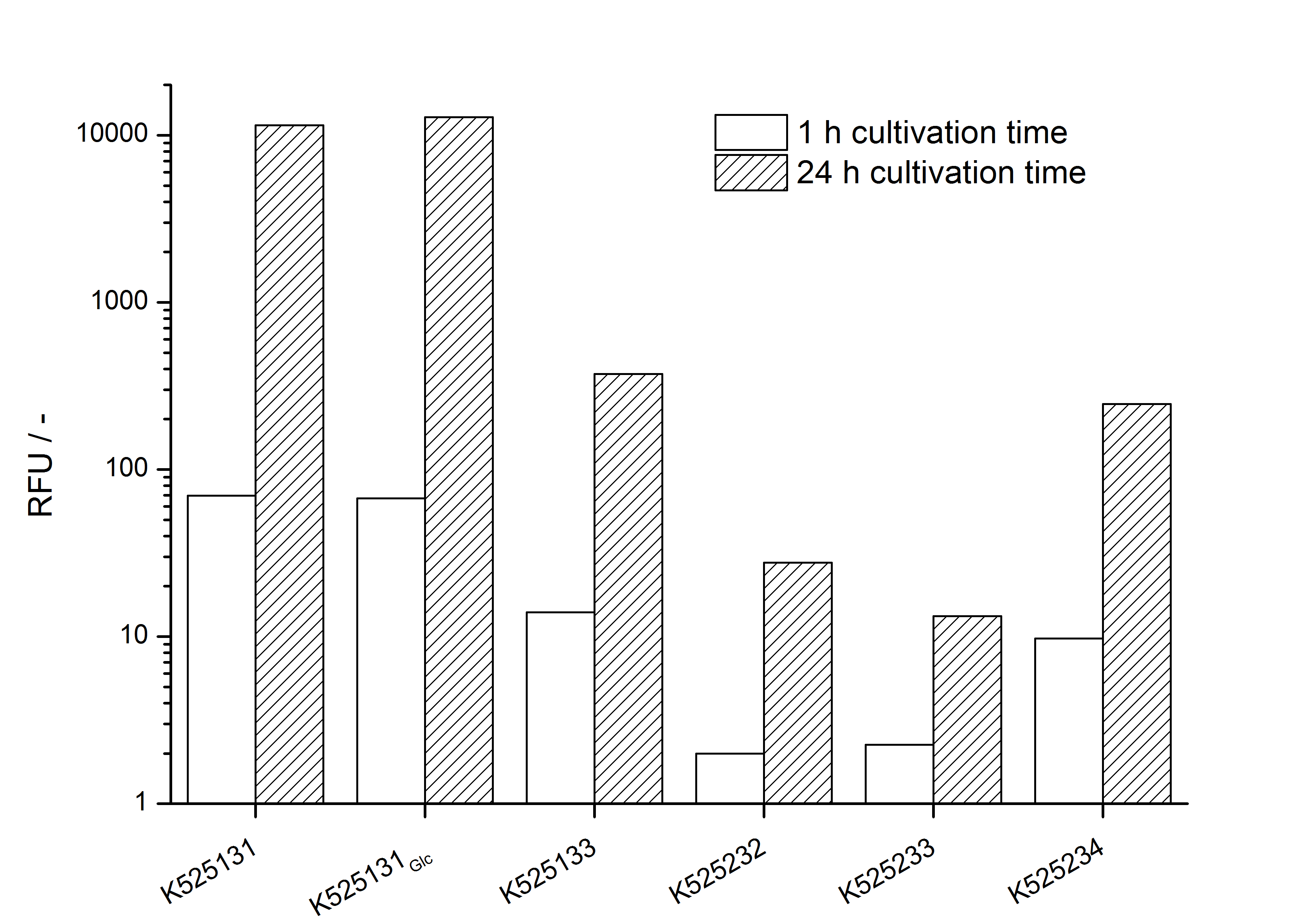

Organizational:
- presenting our posters from the teams of 2010 and 2011 at the congress Biotechnologie2020+ in Berlin hosted by the "Bundesministerium für Bildung und Forschung" (Federal Ministry of Education and Research).
Week 11: 11th July - 17th July
Bisphenol A / S-layer:
- our BioBrick order (some fluorescent proteins and cleavage sites) from iGEM HQ arrived
S-layer:
- our synthesized S-layers SgsE and SbpA finally arrived
Organizational:
- presenting the iGEM competition and our team project at the secondary school Ravensberger Gynmasium in Herford (Photos)
Week 12: 18th July - 24th July
Organizational:
- presenting our projects from 2010 and 2011 at the CeBiTec Symposium 2011 in Bielefeld
- meeting with the iGEM teams from Delft/NL, Edinburgh/UK, Odense/DK. Freiburg/DE and Ljubljana/SL at the Symposium (Photos) was fun!
Bisphenol A / S-Layer:
- removing illegal restriction sites to get valid BioBricks
NAD+ detection:
- cloning the NAD+-dependent DNA ligase from E. coli into BioBrick backbones
Week 13: 25th July - 31th July
S-layer:
- fusing the synthesized S-Layers to a bunch of fluorescent proteins
- successful cloning of K525231 using Gibson assembly.
- K525231: fusion of modified cspB of C. halotolerans including lipid anchor and TAT-sequence <partinfo>K525221</partinfo> with BBa_E1010.
- testing the basal transcription of E. coli KRX by adding inducer (L-rhamnose) or not. The increasing RFU per OD600 reveals the basal transcription of E. coli KRX. L-rhamnose in the medium leads to a ten times higher RFU per OD600.
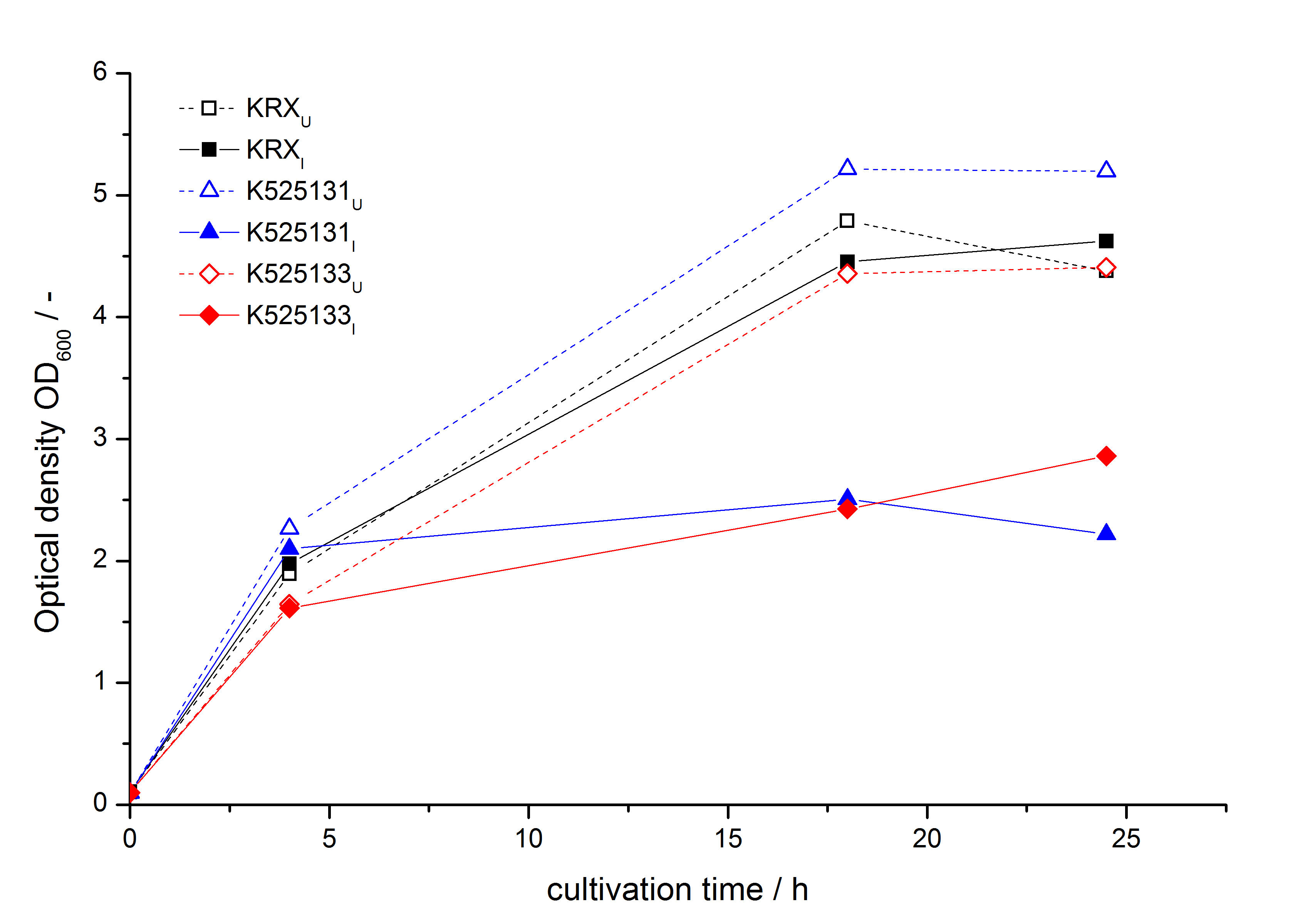
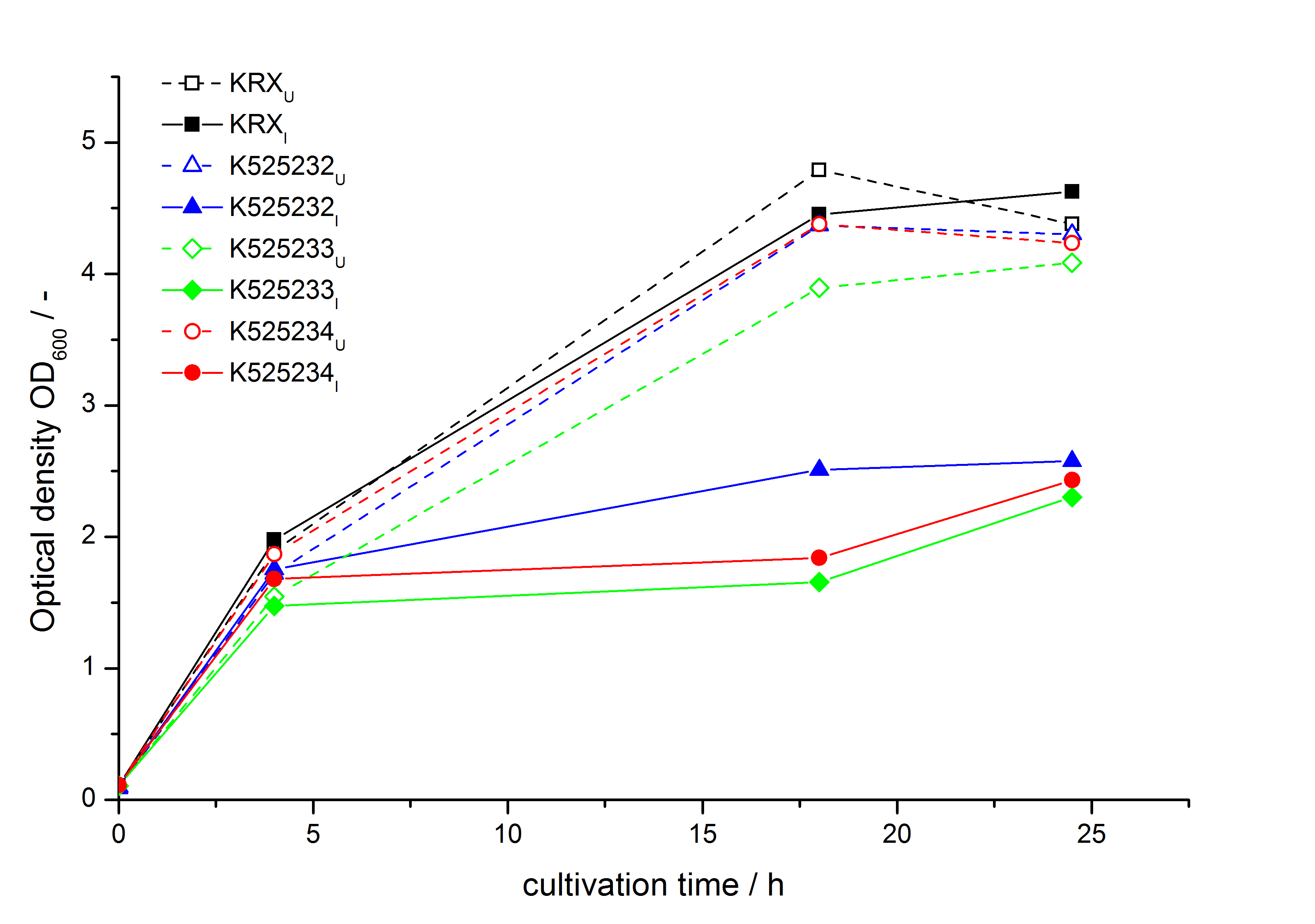
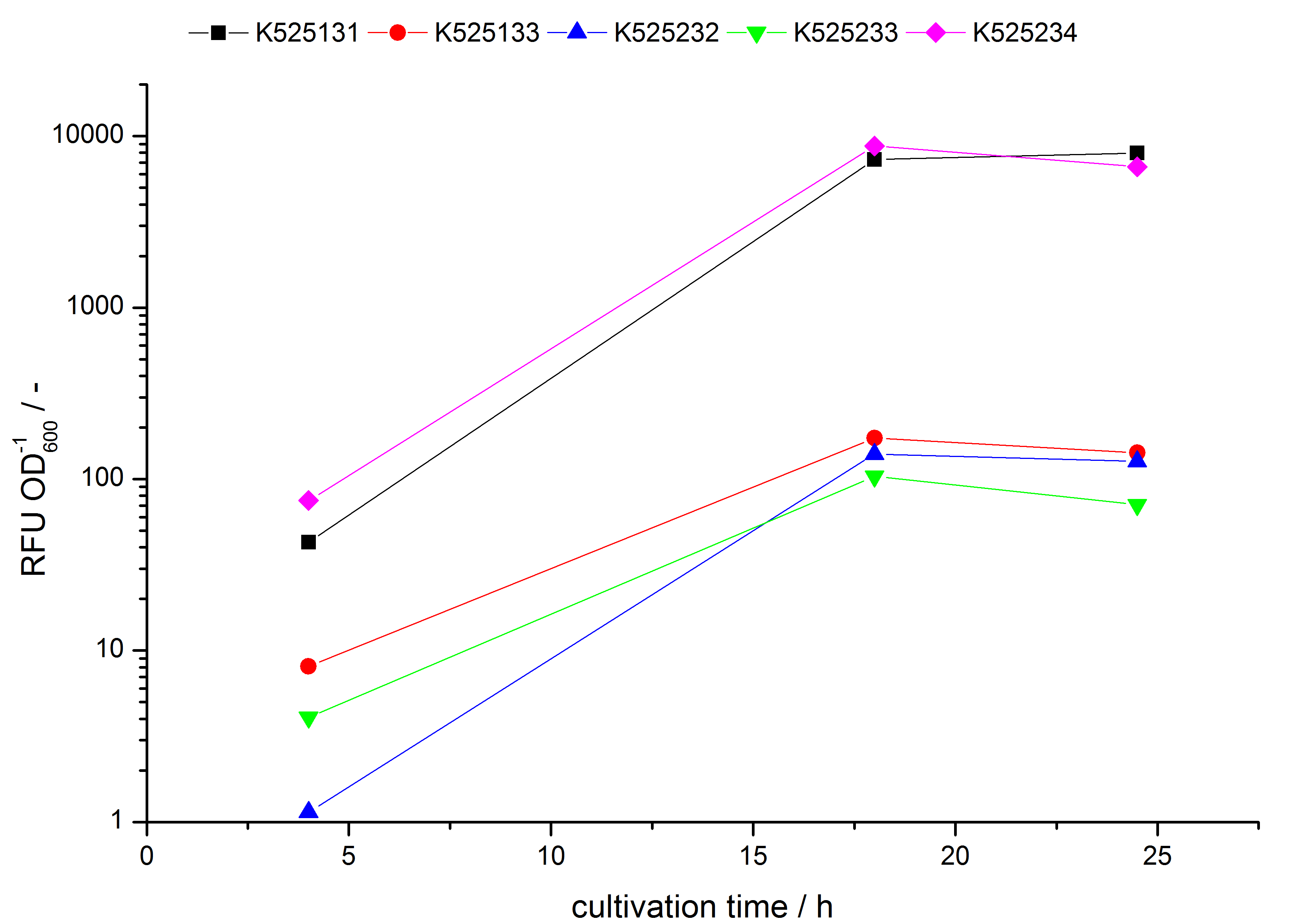
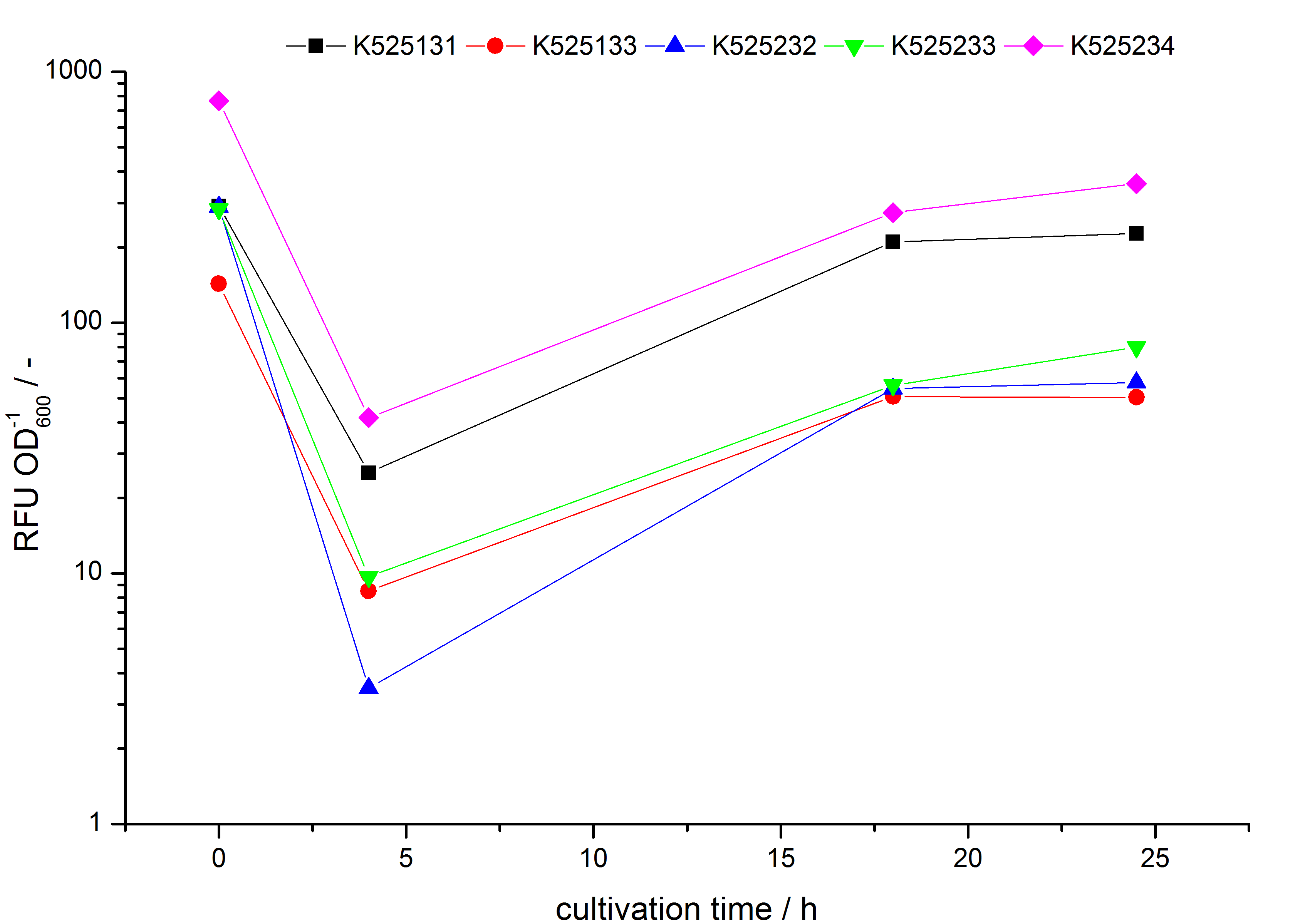
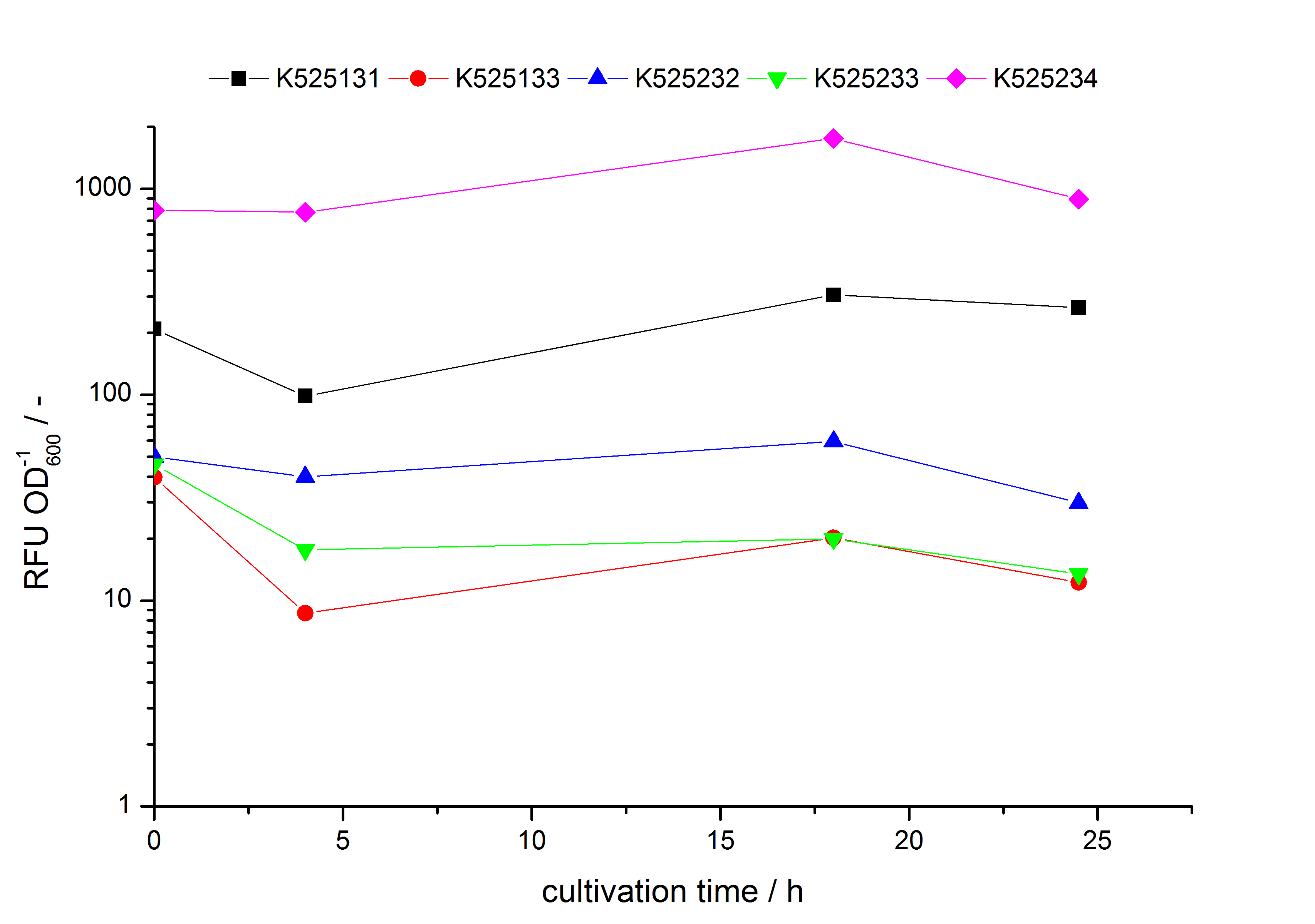
Bisphenol A:
- testing new BPA extraction protocols for LC-MS including an internal standard (bisphenol F)
Week 14: 1st August - 7th August
Bisphenol A:
- BPA analysis with extraction and LC-MS finally works and is very accurate
- Better results for BPA degradation in E. coli -> our fusion protein (<partinfo>K123000</partinfo> to <partinfo>K123001</partinfo>) can completely degrade BPA
- Measuring characterization results for different BPA degrading BioBricks
NAD+ detection:
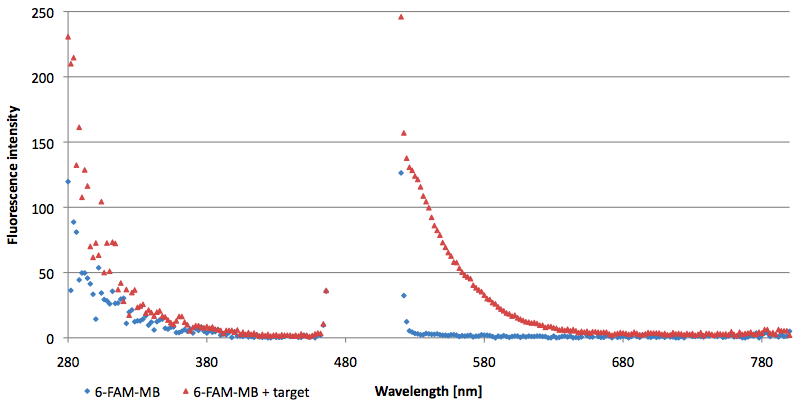
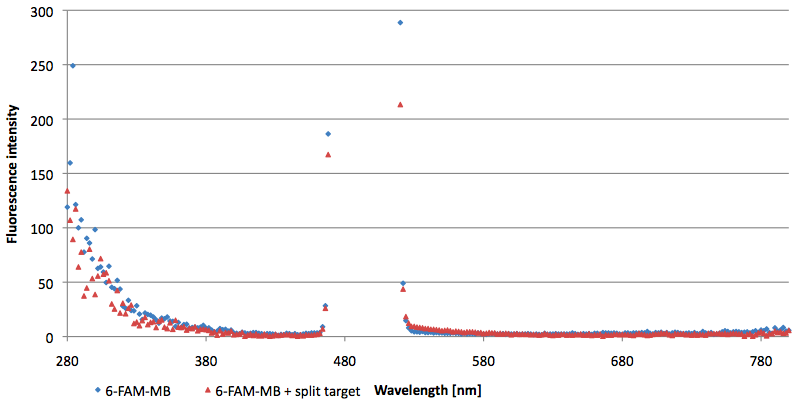
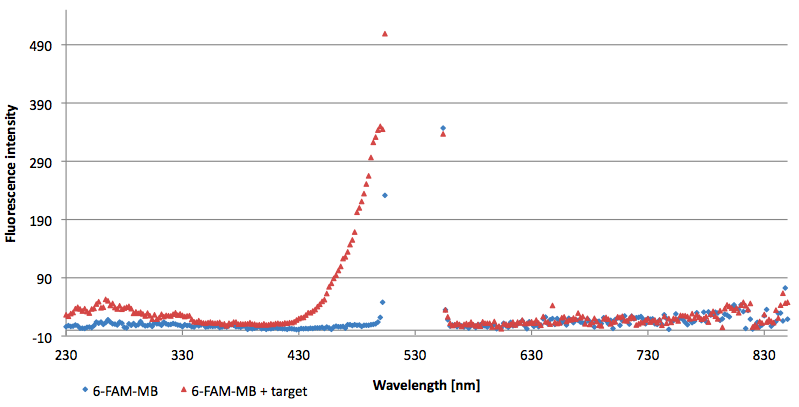
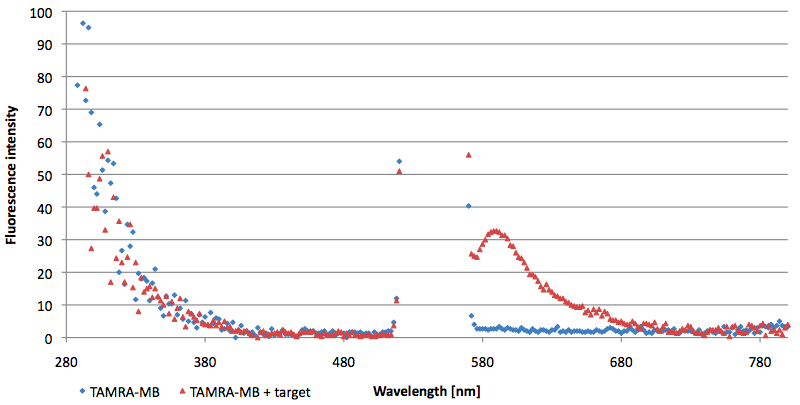
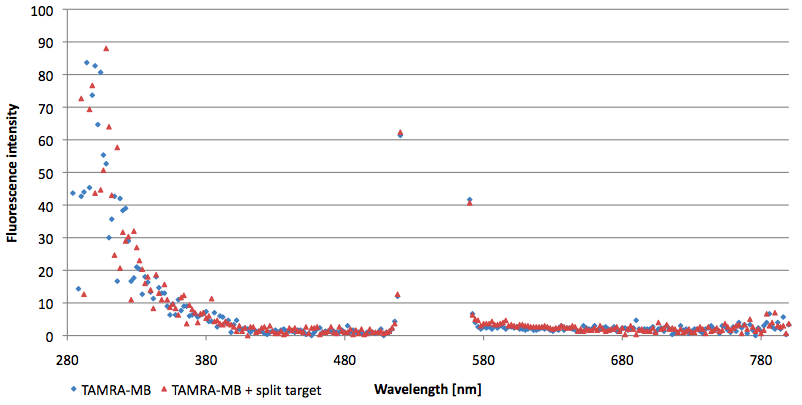
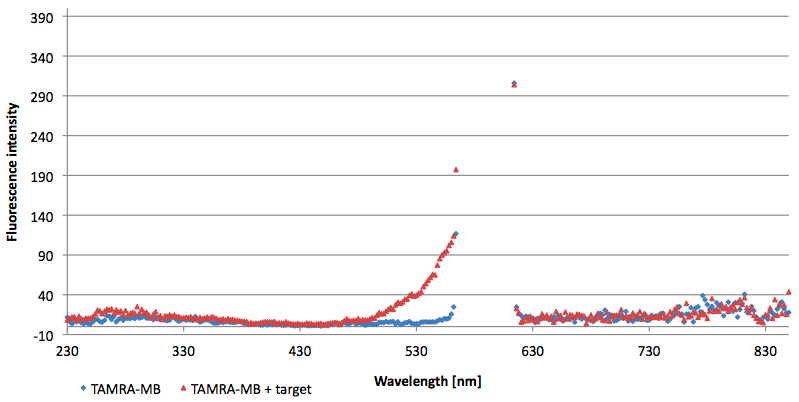
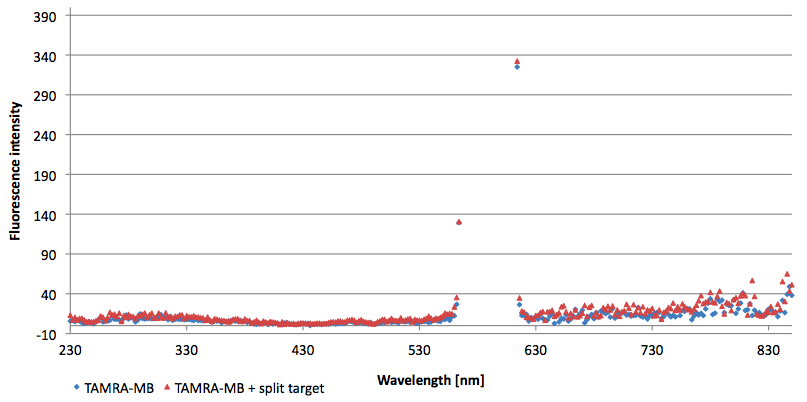
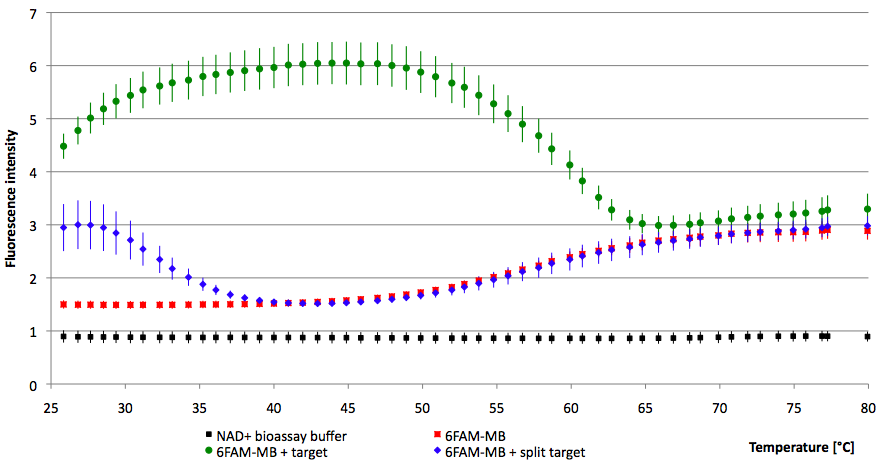
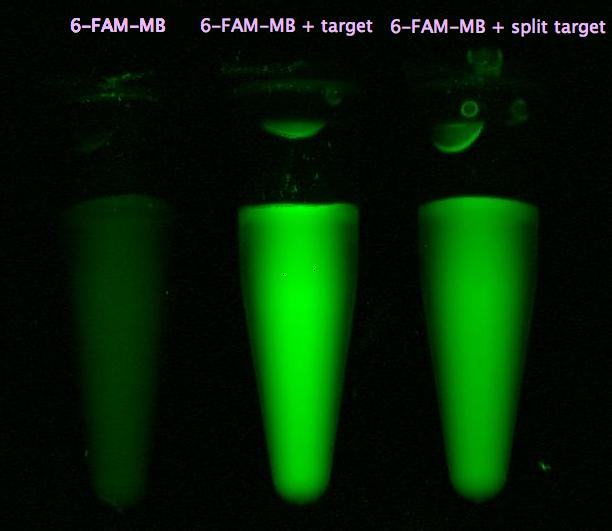
- Successful characterization of two differently labeled (6-FAM or TAMRA with Dabcyl) molecular beacons as a preparation for the NAD+ bioassay including autonomously produced NAD+-dependent DNA ligase (<partinfo>K525710</partinfo>) from E. coli (results shown below)
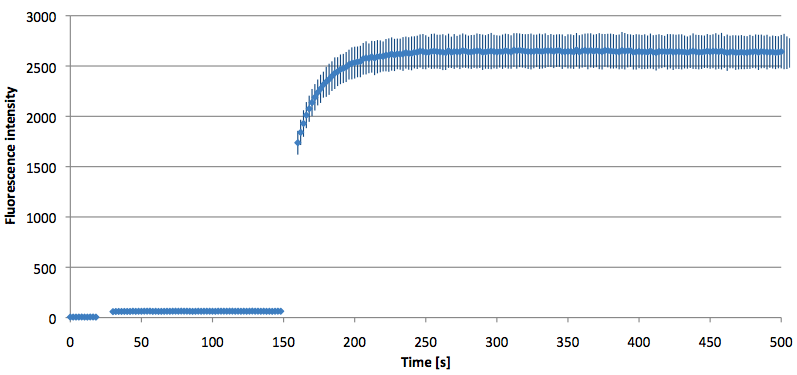
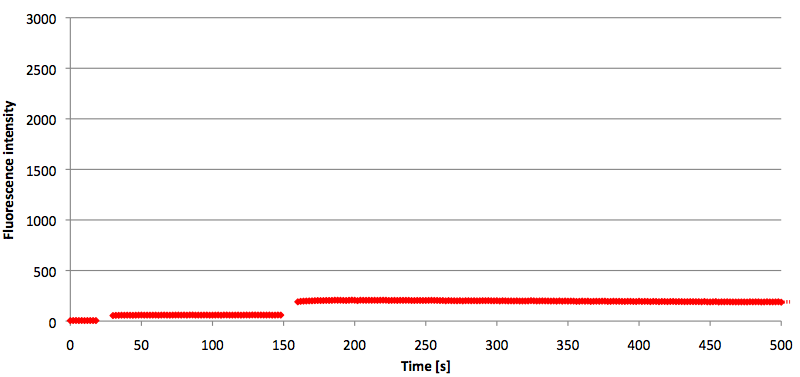
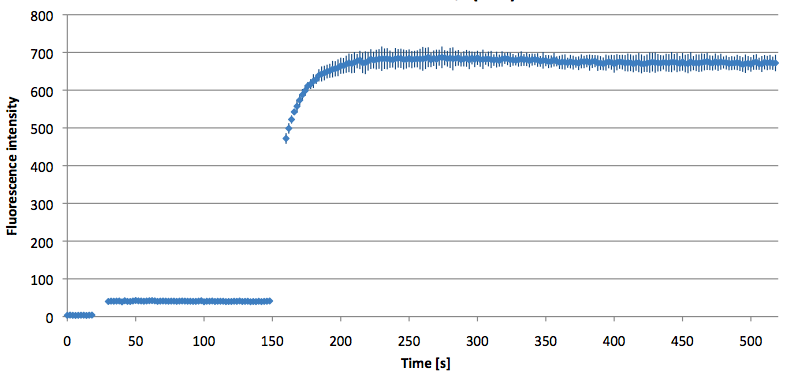
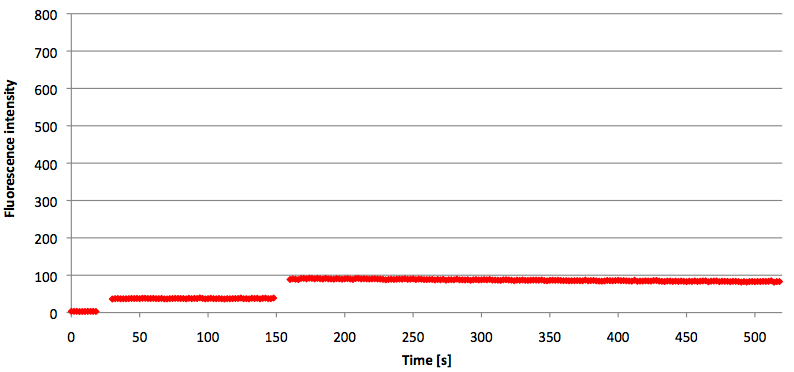
Week 15: 8th August - 14th August
Bisphenol A:
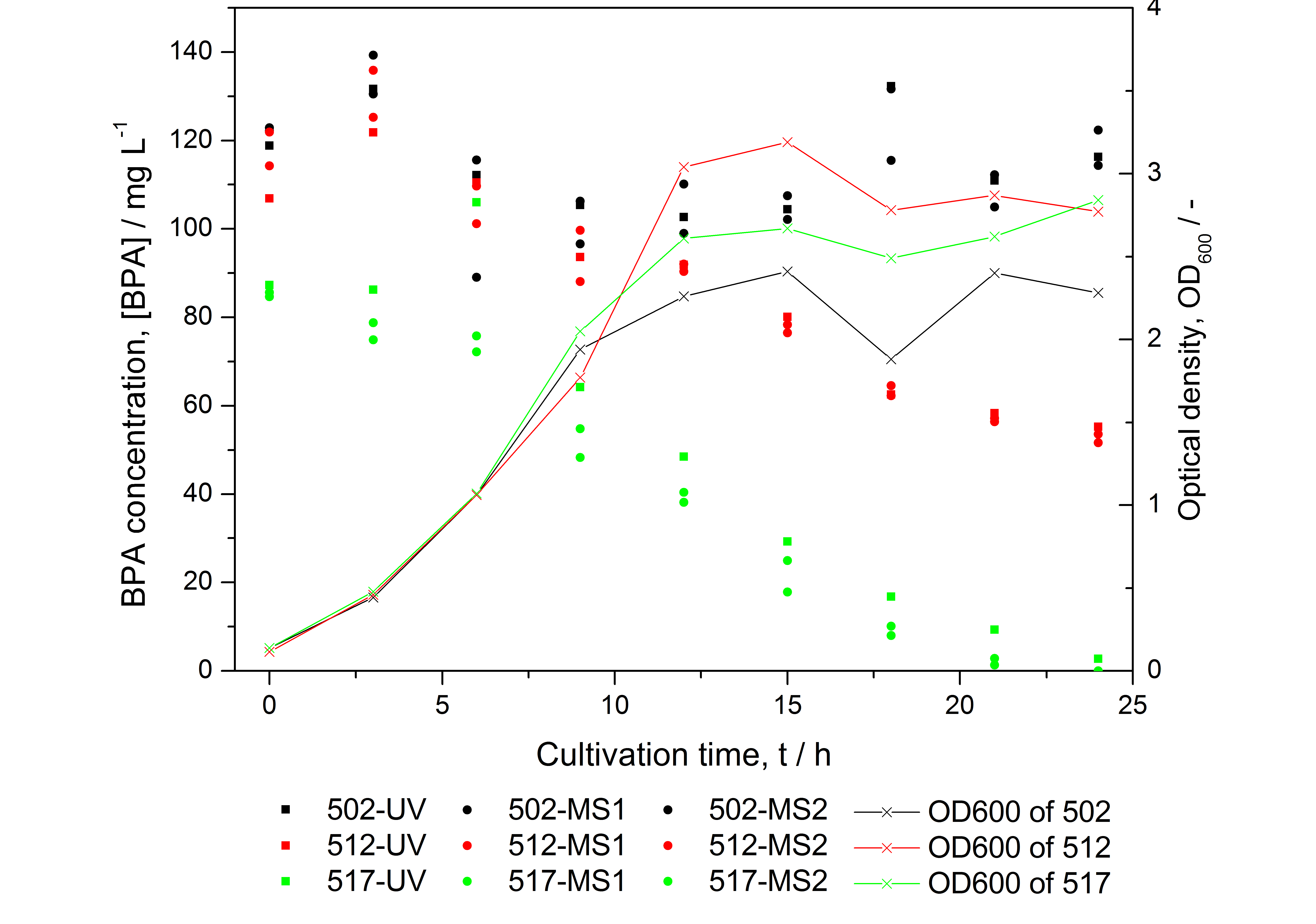
- BPA analysis with extraction and HPLC with UV detector leads to very similar results as the analysis with LC-MS (except for low BPA concentrations -> LOD / LOQ of LC-MS is lower than that of "normal" HPLC, compare Figure 25)
- measuring of more samples from cultivations with BPA degrading BioBricks for further characterization
- we discovered some interesting results in our MS data - soon more
- testing methods to purify his-tagged BisdA and BisdB for cell free BPA degradation and further characterization of these proteins
- testing the influence of BPA on the growth of E. coli
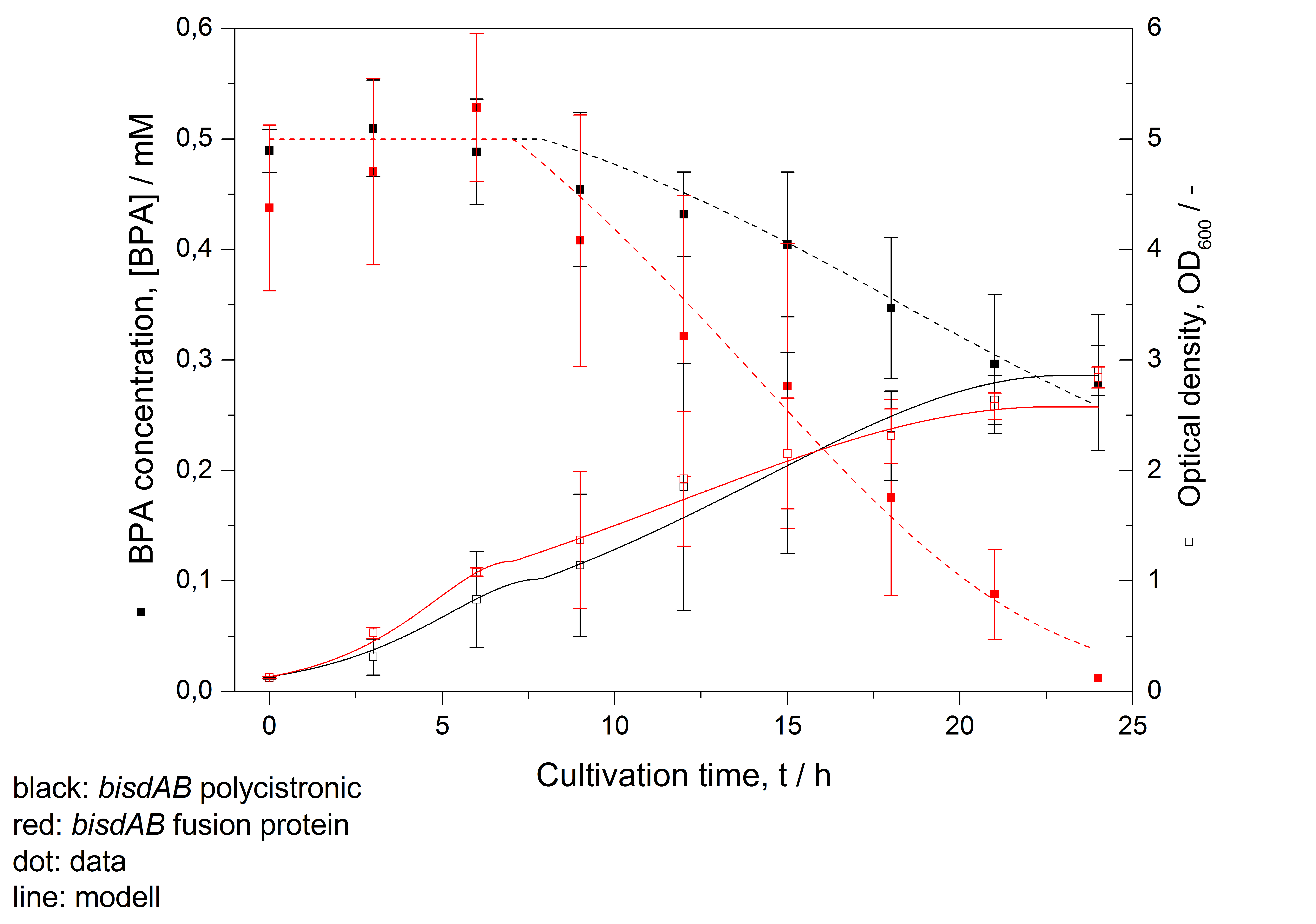
- developing a model for BPA degradation by E. coli (compare Figure 26)
S-layer:
- MALDI-TOF analysis of SDS-PAGEs to characterize the function of the TAT-sequence and the lipid anchor of PS2 (encoded by cspB gene) in E. coli KRX.
Week 16: 15th August - 21st August
S-layer:
- finally found our S-layer proteins in E. coli by analyzing the range of sizes in the polyacrylamid with a MALDI-TOF method-> now we can plan purification strategy
- fusion of FPs to sgsE successful
Bisphenol A:
- all metabolites of natural BPA degradation pathway found during degradation of BPA in E. coli by LC-MS -> now MS/MS to check structure of these metabolites to be sure
- some "metabolites" could also be found due to fragmentation during ionization of the product of BPA degradation
- testing whether E. coli can grow on BPA as the only carbon source (on M9 plates)
- seems like they can't
Week 17: 22nd August - 28th August
S-layer:
- cultivation and test purification of SgsE fusion proteins (<partinfo>K525304</partinfo>, <partinfo>K525305</partinfo> and <partinfo>K525306</partinfo>)
- SDS-PAGEs showed inclusion bodies formation in E coli KRX expressing sgsE.
Bisphenol A:
- purification of his-tagged BisdA and BisdB
- first characterization results entered into partsregistry
- cloning of fusion protein FNR:BisdA:BisdB successful
Week 18: 29th August - 4th September
S-layer:
- developing IEX clean-up for S-layers from Corynebacterium halotolerans
- Searching new purification methods for SgsE fusion proteins.
Bisphenol A:
- successful cloning of polycistronic FNR + BisdA + BisdB
NAD+ detection:
- Successful overexpression of NAD+-dependent DNA ligase (<partinfo>K525710</partinfo>) in E. coli KRX and purification with Ni-NTA columns utilizing the protein`s C-terminal His-tag. Trying out utility for the NAD+ bioassay now.
Organizational:
- participating at the science festival GENIALE in Bielefeld with a lab tour and a presentation with public discussion afterwards (science café). Our participation is also mentioned in the regional newspaper Westfalen Blatt and in the internet on Bielefeld Direkt.
Week 19: 5th September - 11th September
S-layer:
- coating silica beads with fluorescent S-layer fusion protein SgsE | mCitrine, remove these proteins again and find them in SDS-PAGE
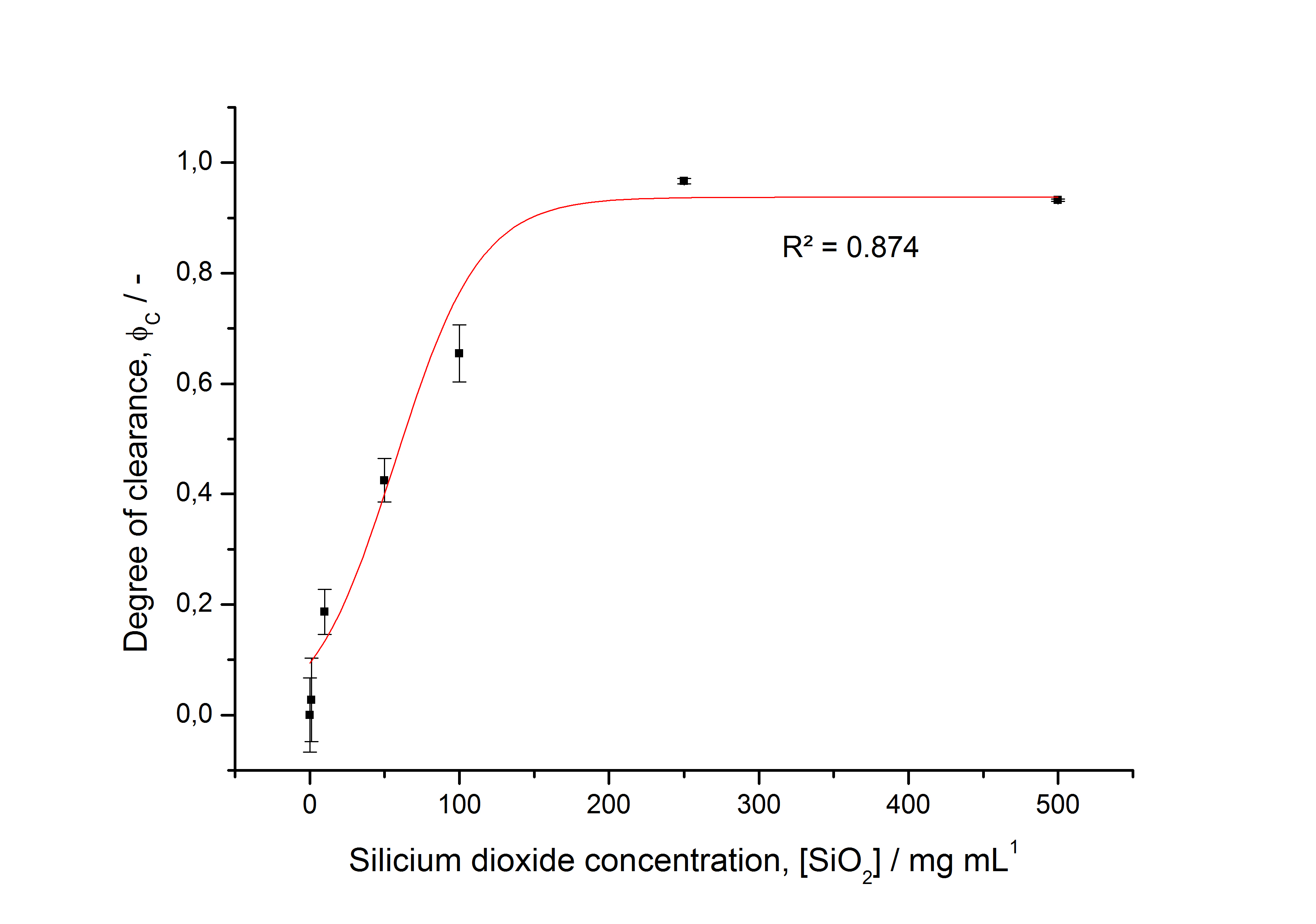
- characterizing immobilization behaviour of S-layers on silica beads -> seems to work (Figure 27)
- cultivate bigger amounts of and develop new purification strategy for S-layer fusion proteins
- testing purification of S-layer fusion proteins with his-tag
- cultivations of fusion protein between NADP+-ferredoxin oxidoreductase and SgsE / SbpA
- also SbpA | mCitrine fusion protein
NAD+ detection:
- Suitability of autonomously produced DNA ligase (<partinfo>K525710</partinfo>) from E. coli for the NAD+ bioassay could be demonstrated after treatment with NMN for deadenylation. Checking out the limit of detection right now.
Week 20: 12th September - 18th September
S-layer:
- testing a purification and coating protocol for S-layer fusion protein mCitrine|SbpA.
- characterizing immobilization behaviour of mCitrine|SbpA (K525405) on silica beads
- optimizing purification and coating protocol for SgsE fusion proteins.
- last characterization cultivations of K525304, K525305, K525306, K525405 and K525406.
Bisphenol A:
- successful cloning of fusion protein FNR + BisdA | BisdB and polycistronic FNR + BisdA + BisdB behind constitutive promoter -> characterizing these last constructs
NAD+ detection:
- NAD+ assay works :)
Week 21: 19th September - 25th September
Organizational:
- lab work is finished, the last measurements and cloning experiments into pSB1C3 are done
- now it is time to fill the Wiki and Partsregistry with brilliant data until European Wikifreeze @ September 21st 11:59 pm EDT!
Week 22: 26th September - 2nd October
Preparation of the presentation and the poster for the European Regional Jamboree.
- practicing the presentation with the Coryne and fermentation engineering working group as audience.
We are qualified for the final in Boston!!! Now labwork can continue.
Week 23: 3rd October - 9th October
S-layer:
- scale-up cultivation of E. coli expressing mCitrine|SgsE (K525305) and mCitrine|SbpA (K525405) fusion protein.
- inclusion body purification and testing further purification strategies.
- assembly of K525422 containing mCitrine|SbpA fused with a His-tag (K157011) for easer purification.
- assembly of K525311 consisting of the SbpA S-layer fused with firefly-luciferase (K525999) for testing an enzyme reaction on immobilized S-layer proteins.
- ordering silicon wafer as a new surface for immobilization.
- successful cloning of fusion protein of K525303 and K525560 (K525001),(K525311), SbpA|LigA (K525006), (K525422),(K525322)
NAD+ detection:
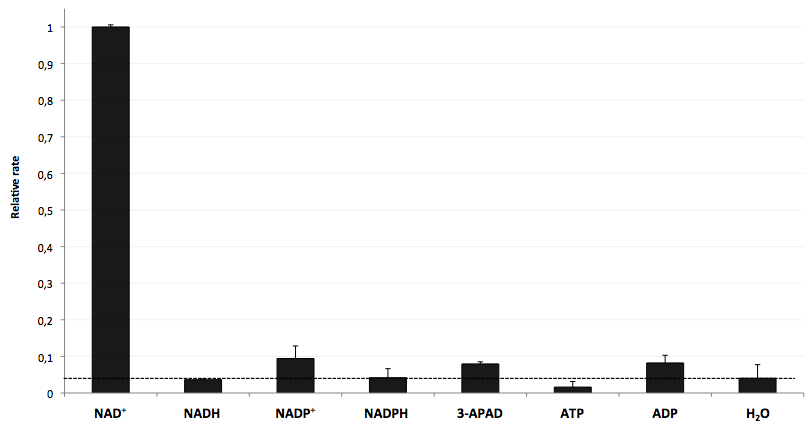
- testing selectivity of LigA (<partinfo>BBa_K525710</partinfo>) for its substrate NAD+ to verify further applications of the molecular beacon based NAD+ bioassay dealing with NAD+ detection in analyte mixtures such as cell lysates.
Week 24: 10th October - 16th October
S-layer:
- first expression of K525422 in E. coli KRX and His-tag affinity chromatography based purification using denaturing conditions.
- Test purifications were carried out using binding buffer with and without imidazole.
- new inclusion body purification of K525305 and K525405 to get new biological material to test the further purification methods.
- establishment of ion exchange chromatography (IEX) and hydrophobic interaction chromatography (HIC) as futher purification steps for S-layer proteins from Lysinibacillus sphaericus and Geobacillus stearothermophilus.
- diethylaminoethyl cellulose (DEAE) as IEX and butyl sepharose™ 4 Fast Flow as HIC purification media.
- test and scale-up expression of mCitrine|SbpA|His-tag (K525322) in E. coli KRX.
- after enzymatic cell lysis with lysozyme and inclusion body purification highest luminescence was measured in the lysis supernatant. This implicates that the fusion protein does not form inclusion bodies.
- successful cloning of fusion protein of CspB and luciferase with Gibson assembly
NAD+ detection:
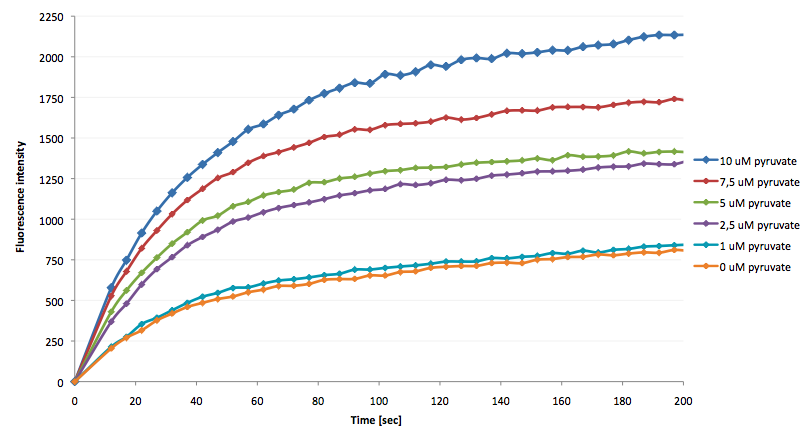
- the NAD+ bioassay was successfully coupled to the NADH-dependent reaction of lactic acid dehydrogenase and could therefore be used to quantify pyruvate
BPA-degradation:
- successful cloning of (K525562) ( Fusion protein of (K525499), (K123000)and (K5123001) with middle strong promoter) and (K525534)
Week 25: 17th October - 23th October
S-layer:
- new expression of <partinfo>K525311</partinfo> to develop an individual purification strategy without inclusion body purification.
- testing a new purification strategy for the fusion protein of the SgsE and the firefly-luciferase based on the IEX and HIC
- first expression of mCitrine|SbpA|His-tag (K525322) and successful His-tag affinity chromatography based purification using denaturing conditions.
NAD+ detection:
- testing LigA for molecular cloning
Week 26: 24th October - 30th October
S-layer:
- test expression of reductase|His-tag (K525534) in E. coli KRX. and testing non denaturating His-tag affinity chromatography.
- recrystallization of mCitrine|SgsE (K525305) and mCitrine|SbpA (K525405) on silicon wafer using two different immobilization strategies.
- trying to get pictures of the S-layer nanostructures on coated silicon wafers using reflection electron microscope (REM) and atomic force microscopy (AFM)
- not enough time left to find a successful method
 "
"